Abstract
Human Leukocyte Antigens (HLA) are cell surface molecules, central in coordinating innate and adaptive immune responses, that are targets of strong diversifying natural selection by pathogens. Of these pathogens, human herpesviruses have a uniquely ancient relationship with our species, where coevolution likely has reciprocating impact on HLA and viral genomic diversity. Consistent with this notion, genetic variation at multiple HLA loci is strongly associated with modulating immunity to herpesvirus infection. Here, we synthesize published genetic associations of HLA with herpesvirus infection and disease, both from case/control and genome-wide association studies. We analyze genetic associations across the eight human herpesviruses and identify HLA alleles that are associated with diverse herpesvirus-related phenotypes. We find that whereas most HLA genetic associations are virus- or disease-specific, HLA-A*01 and HLA-A*02 allotypes may be more generally associated with immune susceptibility and control, respectively, across multiple herpesviruses. Connecting genetic association data with functional corroboration, we discuss mechanisms by which diverse HLA and cognate receptor allotypes direct variable immune responses during herpesvirus infection and pathogenesis. Together, this review examines the complexity of HLA-herpesvirus interactions driven by differential T cell and Natural Killer cell immune responses.
Similar content being viewed by others
Human herpesviruses: infection and disease
Herpesviruses are large double-stranded DNA viruses, having a replication cycle characterized by an initial lytic stage, followed by lifelong latent infection that is largely asymptomatic (Adler et al. 2017; White et al. 2012) (Table 1). There are eight human herpesviruses (HHVs), which span three subfamilies: α, β, and γ. These include three α-herpesviruses: varicella-zoster virus (VZV), herpes simplex virus 1 (HSV-1), and herpes simplex virus 2 (HSV-2); three β-herpesviruses: human cytomegalovirus (HCMV), human herpesvirus 6 (HHV-6, often split into HHV-6A and HHV-6B), and HHV-7; and finally two γ-herpesviruses: Epstein-Barr virus (EBV) and Kaposi’s sarcoma-associated herpesvirus (KSHV). HHVs are successful pathogens—with high seroprevalences of most HHVs in most human populations. This success can be partly attributed to intricate mechanisms of immune evasion, host mimicry, and coercion of host signaling pathways that likely underlie their reduced host damage. For example, EBV infects naïve B cells and expresses a B cell receptor homolog that promotes differentiation into the memory B pool, the preferred site of EBV latency (Mancao and Hammerschmidt 2007). HSV-1 induces skin-homing factors in infected T cells, for eventual delivery of virions to peripheral nerves to establish latent infection (Arvin et al. 2010). HCMV encodes multiple Human Leukocyte Antigen (HLA) homologs, including UL18 and UL142, such that downregulation of host HLA cannot be detected by Natural Killer (NK) cells (Beck and Barrell 1988; Wills et al. 2005). While HHVs are considered border-line commensals and can even provide protection during some coinfections (White et al. 2012), they cause or are associated with a multitude of diseases (Table 1) (Goncalves et al. 2017; Griffiths et al. 2015; Mori and Yamanishi 2007; Rabinstein 2017; Taylor et al. 2015; Zerboni et al. 2014) and are likely, on average, detrimental to fitness. Additionally, primate herpesviruses have rarely switched hosts, and the majority of HHVs likely codiverged with humans from great apes (Azab et al. 2018). This antagonistic yet sustained relationship is expected to result in human-HHV coevolution. Indeed, long-term antagonistic coevolution is consistent with the high degree of HHV host specificity, the fine-tuned control of and evasion from the human immune system by HHVs, and the observed genetic variation in susceptibility to HHV infection and disease. Here, we describe the immunogenetic diversity that underlies HLA and HHV coevolution, particularly in relation to the functional consequences of HLA polymorphism in controlling HHV infection.
The arms of HLA immunity
HLA are cell surface proteins integral in the recognition of pathogens and the subsequent initiation of both cellular innate and adaptive immune responses (Rock et al. 2016). The primary role of HLA molecules is to present small peptide fragments for recognition by the T cell receptor (TCR) on T cells (Boegel et al. 2018; Rock et al. 2016). Polymorphic HLA class I are expressed by most nucleated cells and present intracellular peptides to cytotoxic (CD8+) T cells (Neefjes et al. 2011) and therefore are particularly important in the defense against intracellular pathogens, such as viruses. Additionally, some polymorphic HLA class I serve as ligands for NK cell receptors, most notably the killer cell immunoglobulin-like receptors (KIR) (Pollock et al. 2022) and leukocyte immunoglobulin-like receptors (LILR) (Brown et al. 2004). HLA class I on healthy cells signals “self” through inhibitory NK cell receptors to block NK cell cytotoxicity, but virus-driven HLA downregulation can remove this inhibition, leaving diseased cells vulnerable to NK cell attack (Braud et al. 1998; Hansen and Bouvier 2009; Hilton and Parham, 2017; Pazmany et al. 1996; Shukla et al. 2015). Conversely, activating NK cell receptors complement the function of inhibitory KIR, recognizing HLA class I in a peptide-dependent manner. Thus, activating KIR may promote NK cell cytotoxic responses only during certain infections or alongside loss of other inhibitory signals (Das and Khakoo 2015; Naiyer et al. 2017; Sim et al. 2019). Recent studies have also found that a subset of HLA-DP allotypes (i.e., alleles at the protein level) can interact with the NKp44 NK cell receptor (Niehrs et al. 2019).
HLA class II are canonically expressed by professional antigen-presenting cells, such as dendritic cells, macrophages, and B cells. These molecules can also be upregulated on activated CD4+ and CD8+ T cells. HLA class II proteins bind peptide fragments derived from extracellular pathogens, which have been phagocytosed and subsequently proteolyzed in the endo-lysosomal compartment. Class II presented peptides can be recognized by helper T cells (CD4+), which trigger a wider immune response, most notably including B cell activation (Neefjes et al. 2011).
Through interactions with T cells and NK cells, HLA thus serves as an intersection between innate and adaptive cellular immunity. The molecular pathways governing antigen processing, peptide loading, presentation, and immune responses downstream of HLA have been reviewed in depth elsewhere (Anczurowski and Hirano 2018; Elliott and Williams 2005; Hansen and Bouvier 2009; Kelly and Trowsdale 2019; Neefjes et al. 2011; Rock et al. 2016; Unanue et al. 2016).
HLA-HHV coevolution
HLA loci have long been known to be hyper-polymorphic (Thorsby 2009). For example, the polymorphic HLA classes I and II have approximately 100 times more alleles than the less-polymorphic HLA loci and the presence of HLA-DRB3, HLA-DRB4, and HLA-DRB5 varies across individuals (Robinson et al. 2020). These striking patterns have been attributed to host–pathogen coevolution, which can lead to the maintenance of polymorphism in populations (Anderson and May 1982; Lighten et al. 2017; Radwan et al. 2020). Biologically, this argument is supported by observations that distinct HLA allotypes bind an allotype-specific repertoire of pathogen peptides and that peptide-binding sites of HLA harbor the highest levels of polymorphism (Hughes and Nei 1988; Meyer et al. 2018; Parham et al. 1988; Reche and Reinherz 2003). Further, only a subset of HLA allotypes can direct NK cell responses through KIR. Therefore, individuals heterozygous at a given HLA locus may be able to defend against a greater pathogen diversity than homozygous individuals, or the benefit of specific alleles may fluctuate alongside pathogen incidence or load (Hughes and Nei 1988; Manczinger et al. 2019; Pierini and Lenz 2018; Takahata and Nei 1990). Throughout human evolution, these natural selective forces have led to the differentiation of HLA alleles among populations (Deng et al. 2021), the adaptive introgression of HLA alleles from ancient hominins (Abi-Rached et al. 2011; Greenbaum et al. 2019; Racimo et al. 2015), and ultimately the maintenance of diverse HLA alleles.
In the face of a diverse HLA landscape, HHVs have evolved to evade peptide presentation. An early example of this observation was A*11 epitope loss in populations having high A*11 allotype frequencies, consistent with population-specific evolution to evade local T cell responses (de Campos-Lima et al. 1993). More recent population genetic analyses have also found altered patterns of diversity in HLA-binding peptides and T cell epitopes, together suggesting natural selection could favor protein sequences that are less visible to T cells (Palmer et al. 2022; Vider-Shalit et al. 2007, 2009; Wegner et al. 2019). This is particularly evident for proteins involved in the establishment of latency and early lytic infection, which have fewer HLA-binding peptides (Vider-Shalit et al. 2007, 2009), and the remaining HLA-recognized peptides and T cell epitopes have greater protein diversity (Palmer et al. 2022; Santpere et al. 2014) and altered population structure (Palmer et al. 2022; Wegner et al. 2019). Across human populations, EBV latent, but not lytic, epitope frequencies are inversely correlated with frequency of the HLA class I allotype that presents them to CD8+ T cells (Palmer et al. 2022). These patterns highlight the crucial role of T cell responses during the establishment of latency. Evasion of HLA recognition and downstream T cell responses may be required to provide enough time to establish latency, from which a lifetime of reactivation could realize the full transmission potential of infection.
Alongside the many HHV-encoded inhibitors of the peptide presentation pathways (Griffin et al. 2010), these patterns support HHV adaptation to evade HLA peptide presentation. Support for the inverse, whether specific HLA allotypes are specialized in the control of herpesvirus infections, may be gleaned from genetic association studies.
GWAS highlight a central role for HLA in genetic susceptibility to herpesvirus disease
Genome-wide association studies (GWAS) can provide largely unbiased identification of disease-associated loci. GWAS and case–control studies for herpesviruses have often focused on herpesvirus-associated disease or titers of antibodies targeting herpesvirus genes (Bei et al. 2010; Crosslin et al. 2015; Engdahl et al. 2019; Hammer et al. 2015; Kleinstein et al. 2019; Kuparinen et al. 2012; Rubicz et al. 2013; Sallah et al. 2020; Tang et al. 2012; Tian et al. 2017; Tse et al. 2009). These herpesvirus-focused GWAS reflect three points. First, there are large disparities in research focus across the viruses, with EBV studies representing approximately half of the GWAS and often discriminating among disease phenotypes, while genetic control of beta-herpesviruses and KSHV has only been tested through IgG phenotypes. Second, the genetic variants underlying differential susceptibility to herpesvirus disease are often disease or virus specific. This pattern is formally exemplified in a study that performed 23 GWAS, including those for chickenpox, shingles, HSV-1 cold sore risk, and infectious mononucleosis, where little genetic correlation was observed between herpetic diseases (Tian et al. 2017). Third, and finally, HLA is consistently a component of the genetic architecture of herpesvirus complications and immune responses. In all herpesvirus GWAS that have found significant genetic associations (14/17), variants in HLA loci have been found significant, and in 12/14 of these, variants at HLA were the most significantly associated (Bei et al. 2010; Crosslin et al. 2015; Engdahl et al. 2019; Rubicz et al. 2013; Sallah et al. 2017, 2020; Tang et al. 2012; Tian et al. 2017; Tse et al. 2009). Both HLA class I and class II are represented, with class I associated with viral infection or disease phenotypes and class II associated with circulating IgG levels, consistent with the expected functions of each. For example, EBV IgG levels against EBNA1 are most highly associated with specific variants of HLA-DRB1 and HLA-DQA1 (Hammer et al. 2015; Mentzer et al. 2022; Rubicz et al. 2013; Sallah et al. 2017, 2020), while incidence of NPC and IM is associated with certain HLA-A and HLA-B alleles (Bei et al. 2010; Tang et al. 2012; Tian et al. 2017; Tse et al. 2009). An exception is the antibody response to KSHV, which is associated with a variant located in the genomic interval between HLA-B and HLA-C (Sallah et al. 2020). Interestingly, the enrichment of HLA class I alleles in these associations with disease may reflect, at the population genetic level, the importance of CD8+ T and NK cell responses to herpesvirus infections observed in immunological studies.
Overall, the consistent identification of HLA by independent GWAS across herpesviruses is important, as it establishes precedent for HLA in the genetic control of herpesviruses and warrants consideration of the many case–control studies that have been performed with the a priori hypothesis of an HLA-herpesvirus association.
HLA alleles associated with herpesvirus infection and disease
Associations between specific HLA alleles and herpesvirus diseases were described both before and alongside the discovery of HLA molecular function (Lipinski et al. 1979; Lu et al. 1990; Russell and Schlaut 1975; Simons et al. 1975; Stern 1978). Since then, a multitude of case–control studies have been performed, often focusing on European populations or populations with an unusually high rate of disease. For examples of the latter, EBV-associated nasopharyngeal carcinoma (NPC) is prevalent in Southern Chinese, Taiwanese, and North Africans, Kaposi’s Sarcoma (KS) has a high incidence in Sardinia, and HSV shows high prevalence in sub-Saharan African and South/Central America (Bei et al. 2012; Cottoni et al. 2004; Samandary et al. 2014). These studies have identified specific HLA class I and class II alleles associated with herpesvirus infection and disease severity and generally mirror the patterns identified in GWAS. Of particular interest are those alleles that have been found associated with herpesvirus infection or disease by multiple independent studies, across multiple populations, or across multiple herpesviruses (Fig. 1A, B).
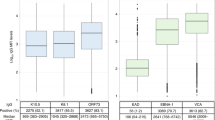
Shared and distinct HLA alleles are associated with herpetic risk. Odds ratios from 38 case/control and GWAS studies were used for a meta-analysis of HLA associations with herpesvirus infection or disease. A, B Odds ratios (ORs) and 95% confidence intervals are plotted from any HLA class I (A) or class II (B) allele with at least two significant associations. Non-significant associations are faded, while significant associations are bold (as determined from p values reported in source publication). Arrows indicate the confidence intervals extend through the bounds of the plot. Points are sized according to the number of participants in each study. Two analyses were performed on published ORs: using only two-field allele data or using both one and two-field data. Significant HLA:virus phenotype associations from two-field only data are provided in the margin in A, B. C Model output from the meta-analysis of herpesvirus-HLA associations using one and two-field data. For each allele having a significant association with at least one virus, the posterior probability distribution of the log odds ratio is plotted, estimated from association with all measured herpesvirus phenotypes. Colored lines and points show the estimated effect size of the individually significant virus-specific genetic associations. Associations specific to an allele at two-field resolution are indicated. All virus-specific effects are shown for A*01 and A*02, each of which had significant general associations across herpesviruses. D A subset of alleles differentially impacts herpesvirus infection or disease. Shown are specific genotypes (right) that have significant, but opposite, impacts on herpesvirus infection and subsequent disease. Measured infection (i.e., IgA seropositivity, shedding, and herpes zoster (HZ)) and disease (NPC, KS, and PHN) phenotypes are shown below the plot and colored by virus. Genotypes with a semicolon are joint KIR/HLA genotypes. C1 and Bw4 are HLA ligands that interact with KIR
To focus our review on the most reproducible evidence, we compiled 38 datasets that tested for associations between HLA alleles and herpesvirus-related phenotypes and performed a pan-herpesvirus meta-analysis of HLA associations. GWAS or HLA-genotyped case–control studies were identified through a literature search for the name of each virus species and either “HLA” or “Human Leukocyte Antigen.” Studies were included in the analysis if they provided either odds ratios (OR) with confidence intervals (CIs) or the appropriate raw frequency data that we could use to calculate OR and CIs. The final dataset included four genome-wide association studies (GWAS) that genotyped HLA to at least first-field resolution (three by imputation and one by PCR-SSOP) (Bei et al. 2010; Tang et al. 2012; Tian et al. 2017; Tse et al. 2009) and 34 candidate gene case/control studies. We recorded ORs and their 95% CIs from all alleles genotyped in each study, totaling 1358 published measurements. A small subset of studies measured two phenotypes (Gayà et al. 2004; Goedert et al. 2016; Lekstrom-Himes et al. 1999; Sato-Takeda et al. 2004), for example, Sato-Takeda et al. (2004) measured both herpes zoster (HZ) and postherpetic neuralgia (PHN) for the same individuals (Sato-Takeda et al. 2004). To avoid pseudoreplication, we used only one phenotype and their respective associations, chosen based on the strongest HLA association.
Of the 38 studies, 26 genotyped HLA at two-field resolution. We analyzed these two-field data in two ways. First, we analyzed only the studies that genotyped HLA at two-field resolution, removing lower resolution associations (Fig. 1A, B “2-field analysis”). Second, we analyzed all one and two-field associations together (Fig. 1C “1 and 2-field analysis”). To reduce the final model complexity, we performed a preliminary analysis to identify the two-field resolution alleles that have significantly different odds ratios than expected from first-field resolution typing. These analyses identified HLA-A*02, B*44, DPB1*04, DQA1*03, DRB1*03, DQB1*03, DRB1*13, DRB1*04, and DRB1*11 as having significant differences in associations at the two-field resolution, and their respective two-field alleles were included in the final analysis. We used MCMCglmm (Hadfield 2010) to model the log odds ratio of HLA-HHV associations using a random effects model with predictors associated with one-field genotypes, two-field genotypes, and the interactions of each with different HHVs. The standard error for each odds ratio was also included as a random effect to model the sampling variance. We assess significance of each allele, for each virus and across viruses, by assessment of the proportion of iterations the Markov chain is positive or negative (Palmer et al. 2018). We include all alleles identified as significant by two or more studies, independent of our model, in Fig. 1A, B, and discuss those results significant from the 1 and 2-field analysis in the following sections. All published associations can be found in Table S1.
First, we tested whether any HLA allele defined at one-field is broadly associated with herpetic infection and disease across all HHVs. We found evidence that A*01 and A*02 allotypes are generally associated, respectively, with susceptibility (random effects model; MCMCp = 0.016) and resistance (random effects model; MCMCp = 0.03) to HHV infection and disease (Fig. 1C). In European populations, A*01 is consistently the most significant HLA susceptibility factor for EBV + HL and IM (Hjalgrim et al. 2010; Huang et al. 2012b; Johnson et al. 2015; Niens et al. 2007) is significantly associated with HSV-1 shedding and lesion rate (Lekstrom-Himes et al. 1999) and with HSV-2 lesion rate (Magaret et al. 2016). Conversely, A*01 is found to be associated with lower risk of HCMV reactivation in solid organ transplant donors in an Irish cohort (Hassan et al. 2016), although this is not the case in an Ashkenazi Jewish cohort (Parks et al. 2018). A*02 is well replicated to be protective against infectious mononucleosis (IM) and Hodgkin’s lymphoma (HL) (Hjalgrim et al. 2010; Huang, Kushekhar et al. 2012; Niens et al. 2007) and also significantly associated with a lower risk of chickenpox (Tian et al. 2017), shingles (Tian et al. 2017), PHN (Meysman et al. 2015), and HCMV reactivation in glioma patients (Han et al. 2017). A*02 carriers also have lower titers of antibodies that recognize HHV6A, which could be caused by more efficient clearance of infected cells by CD8 + T cells (Engdahl et al. 2019; Hammer et al. 2015). Notably, A*02 is present at moderate frequency in most populations (Gonzalez-Galarza et al. 2020). Specifically, protection is more often associated with the A*02:01 allele (Jones et al. 2016; Tian et al. 2017), whereas the A*02:07 allele is a risk factor for EBV-mediated NPC and HL in Chinese and Taiwanese populations (Hildesheim et al. 2002; Huang et al. 2012a; Su et al. 2013; Tse et al. 2009). The single amino acid difference (Y99C) between A*02:01 and A*02:07 occurs in the peptide binding pocket and has been shown to impact the efficiency of peptide presentation from the EBV latent protein, LMP2, suggesting differential coordination of the CD8 T cell response could underlie NPC protection (Lee et al. 1997). Other explanations include strong interactions between A*02 and an unknown environmental variable or functional epistasis with other population-specific variants.
Next, we highlight HLA alleles with supported associations that are individual to or variable across HHVs (Fig. 1C). B*08 mirrors A*01 associations. Although both allotypes are part of the long-range A1-B8-DR3-DQ2 haplotype found at high frequency in Europeans, the B*08 association with HL risk remains after accounting for A*01 (Johnson et al. 2015). The presence of B*44 is associated with either greater or lesser disease risk, depending on the HHV. For example, it is a risk factor for PHN (VZV) in Japanese populations (Meysman et al. 2015), protective against NPC (EBV) in Taiwanese (Hildesheim et al. 2002), protective against shingles (VZV) and HL (EBV) in Europeans (Johnson et al. 2015; Tian et al. 2017), associated with increased risk of KS (KSHV) in Cameroon (Cornejo Castro et al. 2019; Goedert et al. 2016; Guech-Ongey et al. 2010) and with HCMV reactivation in Iran (Futohi et al. 2015). The two most frequent B*44 alleles are B*44:02 and B*44:03, which differ by a single amino acid and can direct differential CD8 T cell responses (Herman et al. 1999). Notably, B*44:03 is the dominant allele in Japanese and Cameroonian populations where B*44 is associated with HHV risk, with B*44:02 close to absent in these populations. Analyses with two-field genotyping have identified B*44:02 as significantly associated with a decreased risk in EBV infection and B*44:03 with an increased risk during VZV infection (Fig. 1C). For class II genes, a haplotype with DRB1*13:02 and DQB1*06:04 is associated with susceptibility to KS in AIDS patients in the US and Italy (Dorak et al. 2005; Guerini et al. 2006; Masala et al. 2005), with PHN in Japan (Meysman et al. 2015), and a similar haplotype (DRB1*13:01 and DQB1*06:03) associated with higher IgG levels to HHV-6A (Engdahl et al. 2019). DRB1*13 alleles are also associated with higher risk of HSV-2 lesions (Lekstrom-Himes et al. 1999), while DRB1*13:02 is associated with lower risk of HSV-1 lesions (Malo and Wank 1998), highlighting the impact of a few residue differences on disease risk. Finally of note, while A*11 is only identified as significant in the meta-analysis in its well-defined association with decreased NPC (EBV) risk in South Asian populations (Bei et al. 2010; Hildesheim et al. 2002; Tang et al. 2012; Tse et al. 2009), it has also been found associated with KS in Italians (Goedert et al. 2016) and HCMV reactivation in transplant patients (Chen et al. 2001).
Intriguingly, a subset of genotypes was observed to have significant but opposite impact on HHV infection versus disease phenotypes (Fig. 1D). A compound genotype encoding KIR2DS4 and its ligand A*11:02 is observed to be associated with reduced EBV/IgA/VCA seroconversion, a risk factor for NPC, but with increased risk of developing NPC (Gao et al. 2017, 2015). A*24 and B*44 are associated with increased (Sato-Takeda et al. 2004) and decreased (Tian et al. 2017) risk of VZV reactivation, respectively, but when the VZV complication, PHN, is considered, the inverse is found (Ozawa et al. 1999; Sato et al. 2002; Sato-Takeda et al. 2004; Sumiyama et al. 2008). A joint genotype encoding KIR3DS1 and its putative Bw4 ligand are associated with reduced KSHV seropositivity, but increased risk of KS (Goedert et al. 2016). Inverse genetic associations with KSHV infection and development of KS were also observed in the context of A*30 (Goedert et al. 2016; Masala et al. 2005), DRB1*04 (Alkharsah et al. 2007; Dorak et al. 2005), and homozygosity of HLA-C1 allotypes (Caselli et al. 2014; Goedert et al. 2016; Guerini et al. 2012). Finally, A*01 is associated with a reduced HSV-2 (Lekstrom-Himes et al. 1999) seropositivity rate but increased occurrence of lesions (Magaret et al. 2016). An alluring hypothesis is that these observations could represent immunological trade-offs, whereby protection against infection has pathological consequences.
HLA alleles and their receptors coordinate variable immune responses
Variation in HLA-mediated immunity is largely described as differential CD8+ T and NK cell responses to infection, broadly aligning with the known importance of these cells in immune responses to herpesviruses (Dittmer and Damania 2016; Egan et al. 2013; Laing et al. 2018; Paludan et al. 2011; Picarda and Benedict 2018; Smith and Khanna 2013; Taylor et al. 2015). CD8+ T cells recognize peptides presented by HLA, and therefore, the distinct viral peptides bound by specific HLA allotypes directly alter T cell recognition (Jing et al. 2016; Sylwester et al. 2005; Taylor et al. 2015). NK cell receptors, some highly polymorphic, recognize infected cells through the combinatorial engagement of ligands, including HLA, on a target cell (Guethlein et al. 2015; Hilton and Parham 2017; Vivier et al. 2011). Allotype-specific peptide presentation by HLA can coordinate variable T cell responses, leading to variation in disease outcome following herpesvirus reactivation. Thus, variable NK cell responses can be directed by allotype-specific engagement of HLA by NK cell receptors. In some cases, there are described immunological mechanisms consistent with and possibly driving the observed HLA genetic associations with HHV phenotypes. Thus, for each HHV subfamily (α, β, and γ), we briefly describe the role of HLA in the immune response to HHVs and then focus on experimental data that lend support to genetic associations described in population studies (i.e., case/control and GWAS).
α-Herpesviruses. The genomes of HSV-1, HSV-2, and VZV share a high proportion of orthologues, likely explaining why the viruses induce cross-reactive antibody (Schmidt et al. 1969) and T cell responses (Jing et al. 2016; Laing et al. 2018; Ouwendijk et al. 2013). CD8 and CD4 T cells control HSV-1 and HSV-2 infection and can be observed surrounding latently infected ganglia (Ouwendijk et al. 2013; van Velzen et al. 2013; Zhu et al. 2007). However, the CD4 T cell response is considered most important during VZV infection (Haberthur et al. 2011; Laing et al. 2018) and its magnitude is associated with less severe HZ (Weinberg et al. 2009).
Proteome-wide CD4 and CD8 T cell screening for HSV-1, HSV-2, and VZV antigens has identified individual and sometimes HLA-restricted responses. While these studies have confirmed the relevance, breadth, and interindividual variation of CD4+ T cell responses to HSV-1 and VZV (Jing et al. 2012; van Velzen et al. 2013), we did not observe overlap between any HLA class II associations with α-herpesvirus disease (Fig. 1) with any obvious trends in class II-restricted T cell responses. However, CD8+ T cell screens have identified variation in HLA class I-restricted T cell responses that overlap with the class I associations found in case–control studies. In two genome-wide screens for HSV-1 and HSV-2 T cell antigens, B*44 allotypes tended towards weak CD8 T cell responses (Jing et al. 2012; Koelle et al. 2003; van Velzen et al. 2013). While no direct associations with HSV-1 and these alleles have been found, the A*33:03-B*44:03-DRB1*13:02 haplotype is associated with PHN (Sato et al. 2002; Sato-Takeda et al. 2004). Also, A*02, associated with lower risk of PHN, is predicted to bind approximately seven times more high-affinity peptides from VZV than B*44 (Meysman et al. 2015) and mediates protective CD8 T cell responses against ocular herpes (Dervillez et al. 2013; Srivastava et al. 2015). Similar to B*44, three HLA-C allotypes including C*04:02 were found to generate weaker CD8 T cell responses (Jing et al. 2012), and C*04 allotypes are associated with increased risk for symptomatic HSV-2 infection (Lekstrom-Himes et al. 1999). Taken together, subpar CD8 T cell responses generated by B*44 and C*04 allotypes could underlie genetic associations between these alleles and α-herpesvirus pathogenesis.
β-Herpesviruses. Interestingly, GWAS for HCMV IgG levels did not find significant genetic associations (Hammer et al. 2015; Kuparinen et al. 2012) and ex vivo proteome-wide epitope mapping did not observe differences in the CD4 or CD8 T cell response across diverse HLA haplotypes (Sylwester et al. 2005). However, solid organ transplant patients with A*01, A*02, or B*44 had higher numbers of HCMV-specific activated T cells (Fernández-Ruiz et al. 2015), and A*01 and A*02 are associated with reduced HCMV seropositivity and reactivation, respectively (Han et al. 2017; Hassan et al. 2016), suggesting differential CD8 T cell responses could drive transplant outcomes in an HLA-dependent manner.
HCMV case–control studies have primarily focused on epistatic genetic variation between NK cell receptors and their HLA ligands as underlying HCMV disease susceptibility, most often in the context of transplantation. This focus is supported by HCMV biology—HCMV infection induces a striking reorganization of the NK cell compartment, with NKG2C+, KIR+ NK cells proliferating following initial infection or HCMV reactivation, leaving stable imprints on the NK cell repertoire that can last over two years (Béziat et al. 2013; Gumá et al. 2004, 2006; Rölle et al. 2014). These NK cells are likely effective in recognizing HCMV-infected cells, as they readily produce IFNγ (Foley et al. 2012) and exhibit antibody-dependent cellular cytotoxic lysis of infected fibroblasts (Costa-Garcia et al. 2015).
NKG2C is an NK cell-activating receptor that recognizes HLA-E, which is a moderately polymorphic HLA class Ib. HLA-E presents peptides derived from HLA class I leader sequences but can also present peptides derived from the UL40 protein of HCMV (Jouand et al. 2018; Pietra et al. 2003; Tomasec et al. 2000). The UL40 peptide is hyper-variable as compared to the rest of the protein, with diversity peaking at a residue that binds to NKG2C, and some UL40 variants fail to provoke a strong adaptive NK cell response (Hammer et al. 2018; Heatley et al. 2013; Vietzen et al. 2021). HLA-E also interacts with the inhibitory NK cell receptor, NKG2A, suggesting HCMV-mediated stabilization of HLA-E and UL40 diversification may reflect a trade-off between evasion of and susceptibility to NK cell damage mediated by NKG2A and NKG2C, respectively (Vietzen et al. 2021). The importance of these differential NK cell responses is demonstrated by genetic associations between UL40 and HLA-E variants with transplant outcomes (Guberina et al. 2018; Vietzen et al. 2021).
Whereas interference with NKG2C or HLA-E on NK cells or infected fibroblasts, respectively, can reduce NK cell proliferation (Rölle et al. 2014), KIR+ NK cells from NKG2C-null individuals still proliferate in response to HCMV infection. This observation suggests additional involvement from KIR and their HLA ligands (Della Chiesa et al. 2014). Indeed, numerous case–control studies have examined the impact of KIR repertoire on HCMV infection following solid organ or hematopoietic cell transplants. Most of these studies find a protective effect of activating KIR on HCMV reactivation, from the donor in the case of hematopoietic transplant or in the recipient during solid organ transplant (Cook et al. 2006; Gonzalez et al. 2014; Hadaya et al. 2008; Stern et al. 2011; Zaia et al. 2009). This protection by activating KIR has been confirmed during primary HCMV infection in immunocompetent hosts, outside the context of immunosuppressed transplant patients (Di Bona et al. 2014). However, other case–control studies find specific combinations of KIR alongside the HLA-C1 ligand to be protective against primary HCMV infection following transplant from a seropositive donor (Jones et al. 2014; van Duin et al. 2014). These findings are broadly reflected in the immunological data: NK cells expressing both activating and inhibitory KIR proliferate in response to HCMV infection (Béziat et al. 2013; Della Chiesa et al. 2014). The activating KIR, KIR2DS1, can directly recognize HCMV infection and aids decidual NK cells in the clearance of HCMV-infected placental cells (Crespo et al. 2016; van der Ploeg et al. 2017). Proliferation of NK cells expressing inhibitory KIR is dependent on NK cell education and is observed only in individuals with allotypes of inhibitory KIR and HLA that can interact (Charoudeh et al. 2013; Djaoud et al. 2013; Manser et al. 2019). Thus, the variable associations in case–control studies may reflect specific gene content or gene-by-environment interactions, as individual KIR haplotypes can contain 0 to 5 activating KIR with variable presence of inhibitory KIR and individual HLA haplotypes can encode a variable number of KIR ligands.
LILRB1 is an inhibitory immune cell receptor that binds broadly to HLA class I ligands (Cosman et al. 1997), and specific variants have also been associated with HCMV infection following transplantation (Yu et al. 2018). To exploit LILRB1 and other inhibitory receptors that recognize class I ligands, HCMV encodes an MHC class I homolog, UL18 (Beck and Barrell 1988), which has been shown to inhibit LILRB1-expressing NK cells (Cosman et al. 1997; Prod’homme et al. 2007). LILRB1-binding domain polymorphism underlies functional variation in the ability to interact with classical HLA class I and UL18 (Yu et al. 2018), but not with HLA-G, where this interaction has a role in placentation (Apps et al. 2007). Murine cytomegalovirus also encodes an MHC-I mimic (Farrell et al. 1997), suggesting that the altering of NK cell responses is critical for infection by cytomegaloviruses.
Although other β-herpesviruses, HHV-6A, HHV-6B, and HHV-7, have not been studied as extensively as HCMV, NK cells seem similarly important in the immune response mounted during their infection (Atedzoe et al. 1997; Flamand et al. 1996), and HHV-6 is both T- and NK-tropic (Eliassen et al. 2017; Lusso et al. 1993). Possibly reflecting functional variation of HLA and KIR, NK cell clones within and across individuals have vastly different cytolytic ability against HHV-6-infected targets (Malnati et al. 1993). Supporting an important function for HLA/KIR interactions during HHV-6A infection, a genotype containing HLA-C1 and its cognate receptors KIR2DS2 and KIR2DL2 is associated with susceptibility to HHV-6A infection (Rizzo et al. 2019). In total, more case–control, genome-wide association, and functional immunity studies will be required to understand the role of HLA variants in infection by these β-herpesviruses.
γ-Herpesviruses. An important role for the cytotoxic T cell response during EBV infection is supported by the large expansion of EBV-reactive CD8 + T cells that defines IM, their sometimes diminished response in HL and NPC patients (Fogg et al. 2009; Gandhi et al. 2006; Li et al. 2007), and strong HLA class I associations localizing to the residues of the peptide-binding groove. During both primary infection and IM, CD8 + T cells against highly immunogenic lytic protein epitopes abound, followed by an immunodominant response towards EBNA3 proteins during the switch to latency. However, while EBNA3 expression is observed in EBV complications of the immunosuppressed (e.g., HIV-associated or posttransplant lymphoproliferative disorders), they are not often expressed in latently infected B cells, nor in HL or NPC malignancies (Price and Luftig 2015). Subdominant T cell responses towards latent proteins EBNA1, EBNA2, and LMP2 also occur, can be HLA-restricted, and may reflect HLA genetic associations with disease (Blake et al. 1997; Brooks et al. 2016; Khanna et al. 1998; Lee et al. 1993). For example, a GWAS identified the peptide-binding grooves of B*38:02 and B*55:02 associated with susceptibility and resistance to NPC, respectively (Tang et al. 2012), and B*38:01 and B*55:01 have been found unique in orchestrating immunodominant responses to EBNA2 (Brooks et al. 2016). Additionally, A*01, A*02, and A*11 have some of the most consistent associations with EBV-associated HL and NPC. Numerous A*02 and A*11 presented peptides from EBV latent proteins stimulate CD8+ T cells, whereas none have been described for the susceptibility allele, A*01 (Taylor et al. 2015). For example, cytotoxic T cell expansions directed against LMP2A are greater from A*02+ donors than A*02− donors in EBV+ HL, but not EBV − HL (Jones et al. 2016). Consistent with these findings, peptide prediction software finds approximately tenfold more high affinity EBNA1, LMP1, and LMP2 peptides presented by A*02 than A*01, whereas this was not observed for other sets of length-matched EBV lytic proteins. While there has been considerably less effort in discovering KSHV epitopes presented by different HLA allotypes (Fang et al. 2019), the current evidence suggests A*02 could be similarly protective (Ribechini et al. 2006; Robey et al. 2010).
NK cells also mediate protection against EBV disease (Chijioke et al. 2016). In response to lytic EBV infection, an early differentiated NK cell subset (CD56dim, NKG2A+, KIR−) proliferates in peripheral blood, protects against both infection and IM, and persists (Azzi et al. 2014; Chijioke et al. 2013; Hendricks et al. 2014; Pappworth et al. 2007; Williams et al. 2005). A similar subset of dendritic cell-activated IFNγhigh NK cells (CD56bright, NKG2A+) is found in tonsils and more potently restricts EBV transformation of B cells (Jud et al. 2017; Lünemann et al. 2013; Strowig et al. 2008). While the exact role for KIR during EBV infection and disease remains unclear, disconnected observations support the hypothesis of KIR involvement. A peptide presented by HLA-C2 allotypes during EBV infection is recognized by the activating KIR2DS1, tonsillar EBV-responsive NK cells express KIR, and acute EBV infection is associated with variable shifts in the percentage of mature NK cells that express KIR (Hendricks et al. 2014; Lünemann et al. 2013; Stewart et al. 2005). Further, multiple candidate gene case/control studies have found associations between KIR polymorphism and EBV infection and complications (Besson et al. 2007; Durovic et al. 2013; Gao et al. 2015; Huo et al. 2015; Kovacic et al. 2005; Qiang et al. 2012). Although not altogether consistent in associations, these studies tend towards the finding that activating KIR underlie susceptibility to NPC, BL, and EBV-HLH (Huo et al. 2015; Kovacic et al. 2005; Muriuki et al. 2021; Nowak et al. 2019; Qiang et al. 2012), although protective against HL (Besson et al. 2007; Jiang et al. 2022). Interestingly, while the A*11:01 allotype is protective from NPC, A*11:02, which differs by one amino acid that results in productive KIR binding, is not and is associated with NPC in a KIR-dependent manner (Gao et al. 2017, 2015; Graef et al. 2009). Similar to EBV, NK cells respond to KSHV and KS (Caselli et al. 2014; Goedert et al. 2016; Guerini et al. 2012), but the only suggestion of a role for KIR are genetic association studies (Beldi-Ferchiou et al. 2016; Dupuy et al. 2012; Sirianni et al. 2002). Together, these genetic associations indicate a function for KIR + NK cells during γ-herpesvirus infection and pathogenesis of malignant disease.
Finally, and unique to EBV, HLA class II serves as an EBV entry receptor into B cells (Li et al. 1997). HLA-DQ allotypes vary in their binding affinity for the EBV envelope glycoprotein, gp42, where glutamic acid at residue 46 of DQB1 is required for gp42 interaction and EBV entry (Haan and Longnecker 2000; McShane et al. 2003). As such, DQB1*03 homozygotes are resistant to EBV infection through the DQ heterodimer (Haan and Longnecker 2000; McShane et al. 2003). However, EBV infection of individuals with non-permissive HLA-DQ allotypes could conceivably proceed through other class II HLA molecules. Consistent with these molecular differences, HLA-DQB1*02 allotypes are associated with gp42 binding and increased EBV seropositivity whereas HLA-DQB1*03, *04, *05, and *06 alleles associate with lack of seroconversion and less strong binding affinity (Li et al. 2017). Following infection, gp42 is repurposed as an HLA class II evasion protein to dampen the CD4 + T cell response (Ressing et al. 2003). While the role of HLA-DQB1 allotypic diversity in immune evasion has not been determined, a failure of gp42 to bind certain HLA-DQ allotypes could promote CD4+ T cell activation, leading to the observed association of specific HLA-DQB1 variants with seropositivity.
Concluding remarks
HLA undoubtedly is under unique natural selective pressures, driven in part by the role in protection from diverse pathogens, resulting in the maintenance of genetic variation that impacts immune responses. The association of HLA variation with HHV pathology is an imperfect window into their sustained coevolutionary relationship. In this complex scenario, diverse HLA alleles are associated with HHV infection and disease differentially across populations. A*01 and A*02 may underlie more general genetic differences in HHV control, but these allotypes also among the most well studied, which could suggest hidden ascertainment biases. A better understanding of HLA-HHV coevolution and the impact of HLA on HHV disease will be gained from association studies from under-represented populations and the continued functional dissection of observed disease associations.
References
Abi-Rached L, Jobin MJ, Kulkarni S, McWhinnie A, Dalva K, Gragert L, Babrzadeh F, Gharizadeh B, Luo M, Plummer FA, Kimani J, Carrington M, Middleton D, Rajalingam R, Beksac M, Marsh SGE, Maiers M, Guethlein LA, Tavoularis S, Parham P (2011) The shaping of modern human immune systems by multiregional admixture with archaic humans. Science 334(6052):89–94. https://doi.org/10.1126/science.1209202
Adler B, Sattler C, Adler H (2017) Herpesviruses and their host cells: a successful liaison. Trends Microbiol 25(3):229–241. https://doi.org/10.1016/j.tim.2016.11.009
Alkharsah KR, Dedicoat M, Blasczyk R, Newton R, Schulz TF (2007) Influence of HLA alleles on shedding of Kaposi sarcoma–associated herpesvirus in saliva in an African population. J Infect Dis 195(6):809–816. https://doi.org/10.1086/511827
Anczurowski M, Hirano N (2018) Mechanisms of HLA-DP antigen processing and presentation revisited. In (Vol. 39, pp. 960–964): Elsevier Ltd
Anderson RM, May RM (1982) Coevolution of hosts and parasites. Parasitology 85(2):411–426. https://doi.org/10.1017/S0031182000055360
Apps R, Gardner L, Sharkey AM, Holmes N, Moffett A (2007) A homodimeric complex of HLA-G on normal trophoblast cells modulates antigen-presenting cells via LILRB1. Eur J Immunol 37(7):1924–1937. https://doi.org/10.1002/eji.200737089
Arvin AM, Moffat JF, Sommer M, Oliver S, Che X, Vleck S, Zerboni L, Ku CC (2010) Varicella-zoster virus T cell tropism and the pathogenesis of skin infection. Curr Top Microbiol Immunol 342(1):189–209. https://doi.org/10.1007/82-2010-29
Atedzoe BN, Ahmad A, Menezes J (1997) Enhancement of natural killer cell cytotoxicity by the human herpesvirus-7 via IL-15 induction. J Immun (Baltimore, Md. 1950), 159(10):4966–4972
Azab W, Dayaram A, Greenwood AD, Osterrieder N (2018) How host specific are herpesviruses? Lessons from herpesviruses infecting wild and endangered mammals. Annual Review of Virology 5(1):53–68. https://doi.org/10.1146/annurev-virology-092917-043227
Azzi T, Lünemann A, Murer A, Ueda S, Béziat V, Malmberg KJ, Staubli G, Gysin C, Berger C, Münz C, Chijioke O, Nadal D (2014) Role for early-differentiated natural killer cells in infectious mononucleosis. Blood 124(16):2533–2543. https://doi.org/10.1182/blood-2014-01-553024
Beck S, Barrell BG (1988) Human cytomegalovirus encodes a glycoprotein homologous to MHC class-I antigens. Nature 331(6153):269–272. https://doi.org/10.1038/331269a0
Bei JX, Jia WH, Zeng YX (2012) Familial and large-scale case-control studies identify genes associated with nasopharyngeal carcinoma. In (Vol. 22, pp. 96–106)
Bei JX, Li Y, Jia WH, Feng BJ, Zhou G, Chen LZ, Feng QS, Low HQ, Zhang H, He F, Tai ES, Kang T, Liu ET, Liu J, Zeng YX (2010) A genome-wide association study of nasopharyngeal carcinoma identifies three new susceptibility loci. Nat Genet 42(7):599–603. https://doi.org/10.1038/ng.601
Beldi-Ferchiou A, Lambert M, Dogniaux S, Vély F, Vivier E, Olive D, Dupuy S, Levasseur F, Zucman D, Lebbé C, Sène D, Hivroz C, Caillat-Zucman S (2016) PD-1 mediates functional exhaustion of activated NK cells in patients with Kaposi sarcoma. Oncotarget 7(45):72961–72977. https://doi.org/10.18632/oncotarget.12150
Besson C, Roetynck S, Williams F, Orsi L, Amiel C, Lependeven C, Antoni G, Hermine O, Brice P, Ferme C, Carde P, Canioni D, Brière J, Raphael M, Nicolas JC, Clavel J, Middleton D, Vivier E, Abel L (2007) Association of killer cell immunoglobulin-like receptor genes with Hogkin’s lymphoma in a familial study. PLoS ONE 2(5). https://doi.org/10.1371/journal.pone.0000406
Béziat V, Liu LL, Malmberg JA, Ivarsson MA, Sohlberg E, Björklund AT, Retière C, Sverremark-Ekström E, Traherne J, Ljungman P, Schaffer M, Price DA, Trowsdale J, Michaëlsson J, Ljunggren HG, Malmberg KJ (2013) NK cell responses to cytomegalovirus infection lead to stable imprints in the human KIR repertoire and involve activating KIRs. Blood 121(14):2678–2688. https://doi.org/10.1182/blood-2012-10-459545
Blake N, Lee S, Redchenko I, Thomas W, Steven N, Leese A, Steigerwald-Mullen P, Kurilla MG, Frappier L, Rickinson A (1997) Human CD8+ T cell responses to EBV EBNA1: HLA class I presentation of the (Gly-Ala)-containing protein requires exogenous processing. Immunity 7(6):791–802. https://doi.org/10.1016/S1074-7613(00)80397-0
Boegel S, Löwer M, Bukur T, Sorn P, Castle JC, Sahin U (2018) HLA and proteasome expression body map. BMC Med Genomics 11(1):36–36. https://doi.org/10.1186/s12920-018-0354-x
Braud VM, Allan DSJ, O’Callaghan CA, Soderstrom K, D’Andrea A, Ogg GS, Lazetic S, Young NT, Bell JI, Phillips JH, Lanler LL, McMichael AJ (1998) HLA-E binds to natural killer cell receptors CD94/NKG2A. B and c Nature 391(6669):795–799. https://doi.org/10.1038/35869
Brooks JM, Long HM, Tierney RJ, Shannon-Lowe C, Leese AM, Fitzpatrick M, Taylor GS, Rickinson AB (2016) Early T cell recognition of B cells following Epstein-Barr virus infection: identifying potential targets for prophylactic vaccination. PLoS Pathogens 12(4). https://doi.org/10.1371/journal.ppat.1005549
Brown D, Trowsdale J, Allen R (2004) The LILR family: modulators of innate and adaptive immune pathways in health and disease. In (Vol. 64, pp. 215–225): Tissue Antigens
Caselli E, Rizzo R, Ingianni A, Contini P, Pompei R, Di Luca D (2014) High prevalence of HHV8 infection and specific killer cell immunoglobulin-like receptors allotypes in Sardinian patients with type 2 diabetes mellitus. J Med Virol 86(10):1745–1751. https://doi.org/10.1002/jmv.23771
Charoudeh HN, Terszowski G, Czaja K, Gonzalez A, Schmitter K, Stern M (2013) Modulation of the natural killer cell KIR repertoire by cytomegalovirus infection. Eur J Immunol 43(2):480–487. https://doi.org/10.1002/eji.201242389
Chen Y, Rocha V, Bittencourt H, Scieux C, Loiseau P, Espérou H, Devergie A, Guardiola P, Socié G, Chevret S, Charron D, Gluckman E, Ribaud P (2001) Relationship between HLA alleles and cytomegalovirus infection after allogenic hematopoietic stem cell transplant. Blood 98(2):500–501. https://doi.org/10.1182/blood.v98.2.500
Chijioke O, Landtwing V, Münz C (2016) NK cell influence on the outcome of primary Epstein-Barr virus infection. In (Vol. 7, pp. 323–323): Frontiers Media S.A
Chijioke O, Müller A, Feederle R, Barros MHM, Krieg C, Emmel V, Marcenaro E, Leung CS, Antsiferova O, Landtwing V, Bossart W, Moretta A, Hassan R, Boyman O, Niedobitek G, Delecluse HJ, Capaul R, Münz C (2013) Human natural killer cells prevent infectious mononucleosis features by targeting lytic epstein-barr virus infection. Cell Rep 5(6):1489–1498. https://doi.org/10.1016/j.celrep.2013.11.041
Cook M, Briggs D, Craddock C, Mahendra P, Milligan D, Fegan C, Darbyshire P, Lawson S, Boxall E, Moss P (2006) Donor KIR genotype has a major influence on the rate of cytomegalovirus reactivation following T-cell replete stem cell transplantation. Blood 107(3):1230–1232. https://doi.org/10.1182/blood-2005-03-1039
Cornejo Castro EM, Morrison BJ, Marshall VA, Labo N, Miley WJ, Clements N, Nelson G, Ndom P, Stolka K, Hemingway-Foday JJ, Abassora M, Gao X, Smith JS, Carrington M, Whitby D (2019) Relationship between human leukocyte antigen alleles and risk of Kaposi’s sarcoma in Cameroon. Genes Immun 20(8):684–689. https://doi.org/10.1038/s41435-019-0077-9
Cosman D, Fanger N, Borges L, Kubin M, Chin W, Peterson L, Hsu ML (1997) A novel immunoglobulin superfamily receptor for cellular and viral MHC class I molecules. Immunity 7(2):273–282. https://doi.org/10.1016/S1074-7613(00)80529-4
Costa-Garcia M, Vera A, Moraru M, Vilches C, López-Botet M, Muntasell A (2015) Antibody-mediated response of NKG2C bright NK cells against human cytomegalovirus. J Immunol 194(6):2715–2724. https://doi.org/10.4049/jimmunol.1402281
Cottoni F, Masala MV, Santarelli R, Carcassi C, Uccini S, Montesu MA, Satta R, Angeloni A, Faggioni A, Cerimele D (2004) Susceptibility to human herpesvirus-8 infection in a healthy population from Sardinia is not directly correlated with the expression of HLA-DR alleles [12]. In (Vol. 151, pp. 247–249)
Crespo ÂC, Strominger JL, Tilburgs T (2016) Expression of KIR2DS1 by decidual natural killer cells increases their ability to control placental HCMV infection. Proc Natl Acad Sci USA 113(52):15072–15077. https://doi.org/10.1073/pnas.1617927114
Crosslin DR, Carrell DS, Burt A, Kim DS, Underwood JG, Hanna DS, Comstock BA, Baldwin E, De Andrade M, Kullo IJ, Tromp G, Kuivaniemi H, Borthwick KM, McCarty CA, Peissig PL, Doheny KF, Pugh E, Kho A, Pacheco J, Jarvik GP (2015) Genetic variation in the HLA region is associated with susceptibility to herpes zoster. Genes Immun 16(1):1–7. https://doi.org/10.1038/gene.2014.51
Das J, Khakoo SI (2015) NK cells: tuned by peptide? Immunol Rev 267(1):214–227. https://doi.org/10.1111/imr.12315
de Campos-Lima PO, Gavioli R, Zhang QJ, Wallace LE, Dolcetti R, Rowe M, Rickinson AB, Masucci MG (1993) HLA-A11 epitope loss isolates of Epstein-Barr virus from a highly A11+ population. Science 260(5104):98–100. https://doi.org/10.1126/science.7682013
Della Chiesa M, Falco M, Bertaina A, Muccio L, Alicata C, Frassoni F, Locatelli F, Moretta L, Moretta A (2014) Human cytomegalovirus infection promotes rapid maturation of NK cells expressing activating killer Ig–like receptor in patients transplanted with NKG2C −/− umbilical cord blood. J Immunol 192(4):1471–1479. https://doi.org/10.4049/jimmunol.1302053
Deng Z, Zhen J, Harrison GF, Zhang G, Chen R, Sun G, Yu Q, Nemat-Gorgani N, Guethlein LA, He L, Tang M, Gao X, Cai S, Palmer WH, Shortt JA, Gignoux CR, Carrington M, Zou H, Parham P, Norman PJ (2021) Adaptive admixture of HLA class I allotypes enhanced genetically determined strength of natural killer cells in East Asians. Mol Biol Evol 38(6):2582–2596. https://doi.org/10.1093/molbev/msab053
Dervillez X, Qureshi H, Chentoufi AA, Khan AA, Kritzer E, Yu DC, Diaz OR, Gottimukkala C, Kalantari M, Villacres MC, Scarfone VM, McKinney DM, Sidney J, Sette A, Nesburn AB, Wechsler SL, BenMohamed L (2013) Asymptomatic HLA-A*02:01–restricted epitopes from herpes simplex virus glycoprotein B preferentially recall polyfunctional CD8 + T cells from seropositive asymptomatic individuals and protect HLA transgenic mice against ocular herpes. J Immunol 191(10):5124–5138. https://doi.org/10.4049/jimmunol.1301415
Di Bona D, Scafidi V, Plaia A, Colomba C, Nuzzo D, Occhino C, Tuttolomondo A, Giammanco G, De Grazia S, Montalto G, Duro G, Cippitelli M, Caruso C (2014) HLA and killer cell immunoglobulin-like receptors influence the natural course of CMV infection. J Infect Dis 210(7):1083–1089. https://doi.org/10.1093/infdis/jiu226
Dittmer DP, Damania B (2016) Kaposi sarcoma-associated herpesvirus: immunobiology, oncogenesis, and therapy. In (Vol. 126, pp. 3165–3175): Am Soc Clin Invest
Djaoud Z, David G, Bressollette C, Willem C, Rettman P, Gagne K, Legrand N, Mehlal S, Cesbron A, Imbert-Marcille B-M, Retière C (2013) Amplified NKG2C + NK cells in cytomegalovirus (CMV) infection preferentially express killer cell Ig-like receptor 2DL: functional impact in controlling CMV-infected dendritic cells. J Immunol 191(5):2708–2716. https://doi.org/10.4049/jimmunol.1301138
Dorak MT, Yee LJ, Tang J, Shao W, Lobashevsky ES, Jacobson LP, Kaslow RA (2005) HLA-B, -DRB1/3/4/5, and -DQB1 gene polymorphisms in human immunodeficiency virus-related Kaposi’s sarcoma. J Med Virol 76(3):302–310. https://doi.org/10.1002/jmv.20361
Dupuy S, Lambert M, Zucman D, Choukem SP, Tognarelli S, Pages C, Lebbé C, Caillat-Zucman S (2012) Human herpesvirus 8 (HHV8) sequentially shapes the NK cell repertoire during the course of asymptomatic infection and Kaposi sarcoma. PLoS Pathogens 8(1). https://doi.org/10.1371/journal.ppat.1002486
Durovic B, Gasser O, Gubser P, Sigle J, Hirsch HH, Stern M, Buser A, Hess C (2013) Epstein-Barr virus negativity among individuals older than 60 years is associated with HLA-C and HLA-Bw4 variants and tonsillectomy. J Virol 87(11):6526–6529. https://doi.org/10.1128/jvi.00169-13
Egan KP, Wu S, Wigdahl B, Jennings SR (2013) Immunological control of herpes simplex virus infections. In (Vol. 19, pp. 328–345): J Neurovirol
Eliassen E, Di Luca D, Rizzo R, Barao I (2017) The interplay between natural killer cells and human herpesvirus-6. In (Vol. 9): MDPI AG
Elliott T, Williams A (2005) The optimization of peptide cargo bound to MHC class I molecules by the peptide-loading complex. In (Vol. 207, pp. 89–99)
Engdahl E, Gustafsson R, Huang J, Biström M, Lima Bomfim I, Stridh P, Khademi M, Brenner N, Butt J, Michel A, Jons D, Hortlund M, Alonso-Magdalena L, Hedström AK, Flamand L, Ihira M, Yoshikawa T, Andersen O, Hillert J, Fogdell-Hahn A (2019) Increased serological response against human herpesvirus 6A is associated with risk for multiple sclerosis. Front Immunol 10:2715–2715. https://doi.org/10.3389/fimmu.2019.02715
Fang Q, Liu Z, Zhang T (2019) Human leukocyte antigen polymorphisms and Kaposi's sarcoma-associated herpesvirus infection outcomes: a call for deeper exploration. In (Vol. 91, pp. 541–548): John Wiley and Sons Inc
Farrell HE, Vally H, Lynch DM, Fleming P, Shellam GR, Scalzo AA, Davis-Poynter NJ (1997) Inhibition of natural killer cells by a cytomegalovirus MHC class I homologue in vivo. Nature 386(6624):510–514. https://doi.org/10.1038/386510a0
Fernández-Ruiz M, Corrales I, Amat P, González E, Andrés A, Navarro D, Aguado JM (2015) Influence of age and HLA alleles on the CMV-specific cell-mediated immunity among CMV-seropositive kidney transplant candidates. In (Vol. 15, pp. 2525–2526): Blackwell Publishing Ltd
Flamand L, Stefanescu I, Menezes J (1996) Human herpesvirus-6 enhances natural killer cell cytotoxicity via IL-15. J Clin Investig 97(6):1373–1381. https://doi.org/10.1172/JCI118557
Fogg MH, Wirth LJ, Posner M, Wang F (2009) Decreased EBNA-1-specific CD8+ T cells in patients with Epstein-Barr virus-associated nasopharyngeal carcinoma. Proc Natl Acad Sci USA 106(9):3318–3323. https://doi.org/10.1073/pnas.0813320106
Foley B, Cooley S, Verneris MR, Pitt M, Curtsinger J, Luo X, Lopez-Vergès S, Lanier LL, Weisdorf D, Miller JS (2012) Cytomegalovirus reactivation after allogeneic transplantation promotes a lasting increase in educated NKG2C + natural killer cells with potent function. Blood 119(11):2665–2674. https://doi.org/10.1182/blood-2011-10-386995
Futohi F, Saber A, Nemati E, Einollahi B, Rostami Z (2015) Human leukocyte antigen alleles and cytomegalovirus infection after renal transplantation. Nephro-Urology Monthly 7(6). https://doi.org/10.5812/numonthly.31635
Gandhi MK, Lambley E, Duraiswamy J, Dua U, Smith C, Elliott S, Gill D, Marlton P, Seymour J, Khanna R (2006) Expression of LAG-3 by tumor-infiltrating lymphocytes is coincident with the suppression of latent membrane antigen-specific CD8+ T-cell function in Hodgkin lymphoma patients. Blood 108(7):2280–2289. https://doi.org/10.1182/blood-2006-04-015164
Gao X, Martin P, Tang M, Hildesheim A, Zeng Y, Naranbhai V, Carrington M (2017) P032 Compound HLA/KIR genotypes influence risk of nasopharyngeal carcinoma in a Southern Chinese cohort. Hum Immunol 78:76–76. https://doi.org/10.1016/j.humimm.2017.06.092
Gao X, Tang M, Martin P, Zeng Y, Carrington M (2015) Compound HLA/KIR genotypes influence risk of nasopharyngeal carcinoma (NPC) in a southern chinese cohort. Hum Immunol 76:52–52. https://doi.org/10.1016/j.humimm.2015.07.071
Gayà A, Esteve A, Casabona J, McCarthy JJ, Martorell J, Schulz TF, Whitby D (2004) Amino acid residue at position 13 in HLA-DR beta chain plays a critical role in the development of Kaposi’s sarcoma in AIDS patients. AIDS 18(2):199–204. https://doi.org/10.1097/00002030-200401230-00008
Goedert JJ, Martin MP, Vitale F, Lauria C, Whitby D, Qi Y, Gao X, Carrington M (2016) Risk of classic Kaposi sarcoma with combinations of killer immunoglobulin-like receptor and human leukocyte antigen loci: a population-based case-control study. J Infect Dis 213(3):432–438. https://doi.org/10.1093/infdis/jiv413
Goncalves PH, Ziegelbauer J, Uldrick TS, Yarchoan R (2017) Kaposi sarcoma herpesvirus-associated cancers and related diseases. Curr Opin HIV AIDS 12(1):47–56. https://doi.org/10.1097/COH.0000000000000330
Gonzalez A, Schmitter K, Hirsch HH, Garzoni C, Van Delden C, Boggian K, Mueller NJ, Berger C, Villard J, Manuel O, Meylan P, Stern M, Hess C (2014) KIR-associated protection from CMV replication requires pre-existing immunity: a prospective study in solid organ transplant recipients. Genes Immun 15(7):495–499. https://doi.org/10.1038/gene.2014.39
Gonzalez-Galarza FF, McCabe A, Santos EJMD, Jones J, Takeshita L, Ortega-Rivera ND, Cid-Pavon GMD, Ramsbottom K, Ghattaoraya G, Alfirevic A, Middleton D, Jones AR (2020) Allele frequency net database (AFND) 2020 update: gold-standard data classification, open access genotype data and new query tools. Nucleic Acids Res 48(D1):D783–D788. https://doi.org/10.1093/nar/gkz1029
Graef T, Moesta AK, Norman PJ, Abi-Rached L, Vago L, Older Aguilar AM, Gleimer M, Hammond JA, Guethlein LA, Bushnell DA, Robinson PJ, Parham P (2009) KIR2DS4 is a product of gene conversion with KIR3DL2 that introduced specificity for HLA-A*11 while diminishing avidity for HLA-C. J Exp Med 206(11):2557–2572. https://doi.org/10.1084/jem.20091010
Greenbaum G, Getz WM, Rosenberg NA, Feldman MW, Hovers E, Kolodny O (2019) Disease transmission and introgression can explain the long-lasting contact zone of modern humans and Neanderthals. Nat Commun 10(1). https://doi.org/10.1038/s41467-019-12862-7
Griffin BD, Verweij MC, Wiertz EJ (2010) Herpesviruses and immunity: the art of evasion. Vet Microbiol 143(1):89–100. https://doi.org/10.1016/j.vetmic.2010.02.017
Griffiths P, Baraniak I, Reeves M (2015) The pathogenesis of human cytomegalovirus. J Pathol 235(2):288–297. https://doi.org/10.1002/path.4437
Guberina H, da Silva Nardi F, Michita RT, Dolff S, Bienholz A, Heinemann FM, Wilde B, Trilling M, Horn PA, Kribben A, Witzke O, Rebmann V (2018) Susceptibility of HLA-E*01:03 allele carriers to develop cytomegalovirus replication after living-donor kidney transplantation. J Infect Dis 217(12):1918–1922. https://doi.org/10.1093/infdis/jix638
Guech-Ongey M, Verboom M, Pfeiffer RM, Schulz TF, Ndugwa CM, Owor AM, Bakaki PM, Bhatia K, Figueiredo C, Eiz-Vesper B, Blasczyk R, Mbulaiteye SM (2010) HLA polymorphisms and detection of kaposi sarcoma-associated herpesvirus DNA in saliva and peripheral blood among children and their mothers in the Uganda sickle cell anemia KSHV study. Infectious Agents and Cancer 5(1):21–21. https://doi.org/10.1186/1750-9378-5-21
Guerini FR, Agliardi C, Mancuso R, Brambilla L, Biffi R, Ferrucci S, Zanetta L, Zanzottera M, Brambati M, Boneschi V, Ferrante P (2006) Association of HLA-DRB1 and -DQB1 with classic Kaposi’s sarcoma in mainland Italy. Cancer Genomics Proteomics 3(3–4):191–196
Guerini FR, Mancuso R, Agostini S, Agliardi C, Zanzottera M, Hernis A, Tourlaki A, Calvo MG, Bellinvia M, Brambilla L, Clerici M (2012) Activating KIR/HLA complexes in classic Kaposi’s sarcoma. Infectious Agents and Cancer 7(1):9–9. https://doi.org/10.1186/1750-9378-7-9
Guethlein LA, Norman PJ, Hilton HHG, Parham P (2015) Co-evolution of MHC class I and variable NK cell receptors in placental mammals. Immunol Rev 267(1):259–282. https://doi.org/10.1111/imr.12326
Gumá M, Angulo A, Vilches C, Gómez-Lozano N, Malats N, López-Botet M (2004) Imprint of human cytomegalovirus infection on the NK cell receptor repertoire. Blood 104(12):3664–3671. https://doi.org/10.1182/blood-2004-05-2058
Gumá M, Budt M, Sáez A, Brckalo T, Hengel H, Angulo A, López-Botet M (2006) Expansion of CD94/NKG2C+ NK cells in response to human cytomegalovirus-infected fibroblasts. Blood 107(9):3624–3631. https://doi.org/10.1182/blood-2005-09-3682
Haan KM, Longnecker R (2000) Coreceptor restriction within the HLA-DQ locus for Epstein-Barr virus infection. Proc Natl Acad Sci USA 97(16):9252–9257. https://doi.org/10.1073/pnas.160171697
Haberthur K, Engelmann F, Park B, Barron A, Legasse A, Dewane J, Fischer M, Kerns A, Brown M, Messaoudi I (2011) CD4 T cell immunity is critical for the control of Simian varicella virus infection in a nonhuman primate model of VZV infection. PLoS Pathog 7(11):e1002367–e1002367. https://doi.org/10.1371/journal.ppat.1002367
Hadaya K, de Rham C, Bandelier C, Bandelier C, Ferrari-Lacraz S, Jendly S, Berney T, Buhler L, Kaiser L, Seebach JD, Tiercy JM, Martin PY, Villard J (2008) Natural killer cell receptor repertoire and their ligands, and the risk of CMV infection after kidney transplantation. Am J Transplant 8(12):2674–2683. https://doi.org/10.1111/j.1600-6143.2008.02431.x
Hadfield JD (2010) MCMC methods for multi-response generalized linear mixed models: the MCMCglmm R package. J Statis Soft 33(2):1–22. https://doi.org/10.18637/jss.v033.i02
Hammer C, Begemann M, McLaren PJ, Bartha I, Michel A, Klose B, Schmitt C, Waterboer T, Pawlita M, Schulz TF, Ehrenreich H, Fellay J (2015) Amino acid variation in HLA class II proteins is a major determinant of humoral response to common viruses. Am J Hum Genet 97(5):738–743. https://doi.org/10.1016/j.ajhg.2015.09.008
Hammer Q, Rückert T, Borst EM, Dunst J, Haubner A, Durek P, Heinrich F, Gasparoni G, Babic M, Tomic A, Pietra G, Nienen M, Blau IW, Hofmann J, Na I-K, Prinz I, Koenecke C, Hemmati P, Babel N, Romagnani C (2018) Peptide-specific recognition of human cytomegalovirus strains controls adaptive natural killer cells. Nat Immunol 19(5):453–463. https://doi.org/10.1038/s41590-018-0082-6
Han S, Deng J, Wang Z, Liu H, Cheng W, Wu A (2017) Decreased human leukocyte antigen A*02:01 frequency is associated with risk of glioma and existence of human cytomegalovirus: a case–control study in Northern China. Cancer Immunol Immunother 66(10):1265–1273. https://doi.org/10.1007/s00262-017-2018-7
Hansen TH, Bouvier M (2009) MHC class i antigen presentation: learning from viral evasion strategies. In (Vol. 9, pp. 503–513)
Hassan J, O'Neill D, Honari B, De Gascun C, Connell J, Keogan M, Hickey D (2016) Cytomegalovirus infection in Ireland. Medicine (United States) 95(6). https://doi.org/10.1097/MD.0000000000002735
Heatley SL, Pietra G, Lin J, Widjaja JML, Harpur CM, Lester S, Rossjohn J, Szer J, Schwarer A, Bradstock K, Bardy PG, Mingari MC, Moretta L, Sullivan LC, Brooks AG (2013) Polymorphism in human cytomegalovirus UL40 impacts on recognition of human leukocyte antigen-E (HLA-E) by natural killer cells. J Biol Chem 288(12):8679–8690. https://doi.org/10.1074/jbc.M112.409672
Hendricks DW, Balfour HH, Dunmire SK, Schmeling DO, Hogquist KA, Lanier LL (2014) Cutting edge: NKG2C hi CD57 + NK cells respond specifically to acute infection with cytomegalovirus and not Epstein-Barr virus. J Immunol 192(10):4492–4496. https://doi.org/10.4049/jimmunol.1303211
Herman J, Jongeneel V, Kuznetsov D, Coulie PG (1999) Differences in the recognition by CTL of peptides presented by the HLA-B*4402 and the HLA-B*4403 molecules which differ by a single amino acid. Tissue Antigens 53(2):111–121. https://doi.org/10.1034/j.1399-0039.1999.530201.x
Hildesheim A, Apple RJ, Chen C-J, Wang SS, Cheng Y-J, Klitz W, Mack SJ, Chen IH, Hsu M-M, Yang C-S, Brinton LA, Levine PH, Erlich HA (2002) Association of HLA class I and II alleles and extended haplotypes with nasopharyngeal carcinoma in Taiwan. J Natl Cancer Inst 94(23):1780–1789. https://doi.org/10.1093/jnci/94.23.1780
Hilton HG, Parham P (2017) Missing or altered self: human NK cell receptors that recognize HLA-C. In (Vol. 69, pp. 567–579): Springer Verlag
Hjalgrim H, Rostgaard K, Johnson PCD, Shield L, Little AM, Ekstrom-Smedby K, Adami HO, Glimelius B, Hamilton-Dutoit S, Kane E, Malcolm Taylor G, McConnachie A, Ryder LP, Sundstrom C, Andersen PS, Chang ET, Alexander FE, Melbye M, Jarrett RF (2010) HLA-A alleles and infectious mononucleosis suggest a critical role for cytotoxic T-cell response in EBV-related Hodgkin lymphoma. Proc Natl Acad Sci USA 107(14):6400–6405. https://doi.org/10.1073/pnas.0915054107
Huang X, Hepkema B, Nolte I, Kushekhar K, Jongsma T, Veenstra R, Poppema S, Gao Z, Visser L, Diepstra A, van den Berg A (2012a) HLA-A*02:07 is a protective allele for EBV negative and a susceptibility allele for EBV positive classical Hodgkin lymphoma in China. PLoS ONE 7(2). https://doi.org/10.1371/journal.pone.0031865
Huang X, Kushekhar K, Nolte I, Kooistra W, Visser L, Bouwman I, Kouprie N, Veenstra R, van Imhoff G, Olver B, Houlston RS, Poppema S, Diepstra A, Hepkema B, van den Berg A (2012b) HLA associations in classical Hodgkin lymphoma: EBV status matters. PLoS ONE, 7(7). https://doi.org/10.1371/journal.pone.0039986
Hughes AL, Nei M (1988) Pattern of nucleotide substitution at major histocompatibility complex class I loci reveals overdominant selection. Nature 335(6186):167–170. https://doi.org/10.1038/335167a0
Huo L, Jiang MY, Li Q, Zhu YP (2015) Novel association of killer cell immunoglobulin-like receptor genes with EBV-infectious diseases in children. In (Vol. 28, pp. 303–307): Elsevier Ltd
Jiang P, Nolte IM, Hepkema BG, Stulp M, van den Berg A, Diepstra A (2022) Killer cell immunoglobulin-like receptor haplotype B modulates susceptibility to EBV-associated classic Hodgkin lymphoma. Front Immunol 13:829943. https://doi.org/10.3389/fimmu.2022.829943
Jing L, Haas J, Chong TM, Bruckner JJ, Dann GC, Dong L, Marshak JO, McClurkan CL, Yamamoto TN, Bailer SM, Laing KJ, Wald A, Verjans GMGM, Koelle DM (2012) Cross-presentation and genome-wide screening reveal candidate T cells antigens for a herpes simplex virus type 1 vaccine. J Clin Investig 122(2):654–673. https://doi.org/10.1172/JCI60556
Jing L, Laing KJ, Dong L, Russell RM, Barlow RS, Haas JG, Ramchandani MS, Johnston C, Buus S, Redwood AJ, White KD, Mallal SA, Phillips EJ, Posavad CM, Wald A, Koelle DM (2016) Extensive CD4 and CD8 T cell cross-reactivity between alphaherpesviruses. J Immunol 196(5):2205–2218. https://doi.org/10.4049/jimmunol.1502366
Johnson PCD, McAulay KA, Montgomery D, Lake A, Shield L, Gallagher A, Little AM, Shah A, Marsh SGE, Taylor GM, Jarrett RF (2015) Modeling HLA associations with EBV-positive and -negative Hodgkin lymphoma suggests distinct mechanisms in disease pathogenesis. Int J Cancer 137(5):1066–1075. https://doi.org/10.1002/ijc.29467
Jones DC, Peacock S, Hughes D, Traherne JA, Allen RL, Barnardo MCNM, Friend P, Taylor CJ, Fuggle S, Trowsdale J, Young NT (2014) Killer immunoglobulin-like receptor gene repertoire influences viral load of primary human cytomegalovirus infection in renal transplant patients. Genes Immun 15(8):562–568. https://doi.org/10.1038/gene.2014.53
Jones K, Wockner L, Brennan RM, Keane C, Chattopadhyay PK, Roederer M, Price DA, Cole DK, Hassan B, Beck K, Gottlieb D, Ritchie DS, Seymour JF, Vari F, Crooks P, Burrows SR, Gandhi MK (2016) The impact of HLA class I and EBV latency-II antigen-specific CD8+ T cells on the pathogenesis of EBV+ Hodgkin lymphoma. Clin Exp Immunol 183(2):206–220. https://doi.org/10.1111/cei.12716
Jouand N, Bressollette-Bodin C, Gérard N, Giral M, Guérif P, Rodallec A, Oger R, Parrot T, Allard M, Cesbron-Gautier A, Gervois N, Charreau B (2018) HCMV triggers frequent and persistent UL40-specific unconventional HLA-E-restricted CD8 T-cell responses with potential autologous and allogeneic peptide recognition. PLoS Pathogens 14(4). https://doi.org/10.1371/journal.ppat.1007041
Jud A, Kotur M, Berger C, Gysin C, Nadal D, Lünemann A (2017) Tonsillar CD56brightNKG2A+ NK cells restrict primary Epstein-Barr virus infection in B cells via IFN-γ. Oncotarget 8(4):6130–6141. https://doi.org/10.18632/oncotarget.14045
Kelly A, Trowsdale J (2019) Genetics of antigen processing and presentation. In (Vol. 71, pp. 161–170): Springer Verlag
Khanna R, Burrows SR, Nicholls J, Poulsen LM (1998) Identification of cytotoxic T cell epitopes within Epstein-Barr virus (EBV) oncogene latent membrane protein 1 (LMP1): evidence for HLA A2 supertype-restricted immune recognition of EBV-infected cells by LMP1-specific cytotoxic T lymphocytes. Eur J Immunol 28(2):451–458. https://doi.org/10.1002/(SICI)1521-4141(199802)28:02%3c451::AID-IMMU451%3e3.0.CO;2-U
Kleinstein SE, Shea PR, Allen AS, Koelle DM, Wald A, Goldstein DB (2019) Genome-wide association study (GWAS) of human host factors influencing viral severity of herpes simplex virus type 2 (HSV-2). Genes Immun 20(2):112–120. https://doi.org/10.1038/s41435-018-0013-4
Koelle DM, Liu Z, McClurkan CL, Cevallos RC, Vieira J, Hosken NA, Meseda CA, Snow DC, Wald A, Corey L (2003) Immunodominance among herpes simplex virus-specific CD8 T cells expressing a tissue-specific homing receptor. Proc Natl Acad Sci USA 100(22):12899–12904. https://doi.org/10.1073/pnas.2131705100
Kovacic MB, Martin M, Gao X, Fuksenko T, Chen CJ, Cheng YJ, Chen JY, Apple R, Hildesheim A, Carrington M (2005) Variation of the killer cell immunoglobulin-like receptors and HLA-C genes in nasopharyngeal carcinoma. Canc Epidemiol Biomark Prevent 14(11 I):2673–2677. https://doi.org/10.1158/1055-9965.EPI-05-0229
Kuparinen T, Seppälä I, Jylhävä J, Marttila S, Aittoniemi J, Kettunen J, Viikari J, Kähönen M, Raitakari O, Lehtimäki T, Hurme M (2012) Genome-wide association study does not reveal major genetic determinants for anti-cytomegalovirus antibody response. Genes Immun 13(2):184–190. https://doi.org/10.1038/gene.2011.71
Laing KJ, Ouwendijk WJD, Koelle DM, Verjans GM (2018) Immunobiology of varicella-zoster virus infection. J Infect Dis 218(suppl_2):S68-S74. https://doi.org/10.1093/infdis/jiy403
Lee SP, Thomas WA, Murray RJ, Khanim F, Kaur S, Young LS, Rowe M, Kurilla M, Rickinson AB (1993) HLA A2.1-restricted cytotoxic T cells recognizing a range of Epstein-Barr virus isolates through a defined epitope in latent membrane protein LMP2. J Virol 67(12), 7428–7435. https://doi.org/10.1128/jvi.67.12.7428-7435.1993
Lee SP, Tierney RJ, Thomas WA, Brooks JM, Rickinson AB (1997) Conserved CTL epitopes within EBV latent membrane protein 2: a potential target for CTL-based tumor therapy. J Immun (Baltimore, Md. : 1950), 158(7):3325–3334
Lekstrom-Himes JA, Hohman P, Warren T, Wald A, Nam JM, Simonis T, Corey L, Straus SE (1999) Association of major histocompatibility complex determinants with the development of symptomatic and asymptomatic genital herpes simplex virus type 2 infections. J Infect Dis 179(5):1077–1085. https://doi.org/10.1086/314729
Li J, Zeng XH, Mo HY, Rolén U, Gao YF, Zhang XS, Chen QY, Zhang L, Zeng MS, Li MZ, Huang WL, Wang XN, Zang YX, Masucci MG (2007) Functional inactivation of EBV-specific T-lymphocytes in nasopharyngeal carcinoma: implications for tumor immunotherapy. PLoS ONE 2(11). https://doi.org/10.1371/journal.pone.0001122
Li Q, Bu W, Gabriel E, Aguilar F, Hoshino Y, Miyadera H, Hess C, Hornung RL, Roy A, Cohen JI (2017) HLA-DQ β1 alleles associated with Epstein-Barr virus (EBV) infectivity and EBV gp42 binding to cells. JCI Insight 2(4). https://doi.org/10.1172/jci.insight.85687
Li Q, Spriggs MK, Kovats S, Turk SM, Comeau MR, Nepom B, Hutt-Fletcher LM (1997) Epstein-Barr virus uses HLA class II as a cofactor for infection of B lymphocytes. J Virol 71(6):4657–4662. https://doi.org/10.1128/jvi.71.6.4657-4662.1997
Lighten J, Papadopulos AST, Mohammed RS, Ward BJ, Paterson GI, Baillie L, Bradbury IR, Hendry AP, Bentzen P, Van Oosterhout C (2017) Evolutionary genetics of immunological supertypes reveals two faces of the Red Queen. Nat Commun 8(1):1294–1294. https://doi.org/10.1038/s41467-017-01183-2
Lipinski M, Fridman WH, Tursz T, Vincent C, Pious D, Fellous M (1979) Absence of allogeneic restriction in human t-cell-mediated cytotoxicity to epstein-barr virus-infected target cells: demonstration of an HLA-linked control at the effector level. J Exp Med 150(6):1310–1322. https://doi.org/10.1084/jem.150.6.1310
Lu SJ, Day NE, Degos L, Lepage V, Wang PC, Chan SH, Simons M, McKnight B, Easton D, Zeng Y, De-Thé G (1990) Linkage of a nasopharyngeal carcinoma susceptibility locus to the HLA region. Nature 346(6283):470–471. https://doi.org/10.1038/346470a0
Lünemann A, Vanoaica LD, Azzi T, Nadal D, Münz C (2013) A distinct subpopulation of human NK cells restricts B cell transformation by EBV. J Immunol 191(10):4989–4995. https://doi.org/10.4049/jimmunol.1301046
Lusso P, Malnati MS, Garzino-Demo A, Crowley RW, Long EO, Gallo RC (1993) Infection of natural killer cells by human herpesvirus 6. Nature 362(6419):458–462. https://doi.org/10.1038/362458a0
Magaret A, Dong L, John M, Mallal SA, James I, Warren T, Gaudieri S, Koelle DM, Wald A (2016) HLA class i and II alleles, heterozygosity and HLA-KIR interactions are associated with rates of genital HSV shedding and lesions. Genes Immun 17(7):412–418. https://doi.org/10.1038/gene.2016.42
Malnati MS, Lusso P, Ciccone E, Moretta A, Moretta L, Long EO (1993) Recognition of virus-lnfected cells by natural killer cell clones is controlled by polymorphic target cell elements. J Exp Med 178(3):961–969. https://doi.org/10.1084/jem.178.3.961
Malo K, Wank. (1998) Recurrent herpes simplex virus-induced erythema multiforme: different HLA-DQB1 alleles associate with severe mucous membrane versus skin attacks. Scand J Immunol 47(5):408–411. https://doi.org/10.1046/j.1365-3083.1998.00357.x
Mancao C, Hammerschmidt W (2007) Epstein-Barr virus latent membrane protein 2A is a B-cell receptor mimic and essential for B-cell survival. Blood 110(10):3715–3721. https://doi.org/10.1182/blood-2007-05-090142
Manczinger M, Boross G, Kemény L, Müller V, Lenz TL, Papp B, Pál C (2019) Pathogen diversity drives the evolution of generalist MHC-II alleles in human populations. PLoS Biology 17(1). https://doi.org/10.1371/journal.pbio.3000131
Manser AR, Scherenschlich N, Thöns C, Hengel H, Timm J, Uhrberg M (2019) KIR polymorphism modulates the size of the adaptive NK cell pool in human cytomegalovirus–infected individuals. J Immunol 203(8):2301–2309. https://doi.org/10.4049/jimmunol.1900423
Masala MV, Carcassi C, Cottoni F, Mulargia M, Contu L, Cerimele D (2005) Classic Kaposi’s sarcoma in Sardinia: HLA positive and negative associations. Int J Dermatol 44(9):743–745. https://doi.org/10.1111/j.1365-4632.2004.02206.x
McShane MP, Mullen MM, Haan KM, Jardetzky TS, Longnecker R (2003) Mutational analysis of the HLA class II interaction with Epstein-Barr virus glycoprotein 42. J Virol 77(13):7655–7662. https://doi.org/10.1128/jvi.77.13.7655-7662.2003
Mentzer AJ, Brenner N, Allen N, Littlejohns TJ, Chong AY, Cortes A, Almond R, Hill M, Sheard S, McVean G, Board UKBIA, Collins R, Hill AVS, Waterboer T (2022) Identification of host-pathogen-disease relationships using a scalable multiplex serology platform in UK Biobank. Nat Commun 13(1):1818. https://doi.org/10.1038/s41467-022-29307-3
Meyer D, Vitor VR, Bitarello BD, Débora DY, Nunes K (2018) A genomic perspective on HLA evolution. In (Vol. 70, pp. 5–27):Springer Verlag
Meysman P, Ogunjimi B, Naulaerts S, Beutels P, Van Tendeloo V, Laukens K (2015) Varicella-zoster virus-derived major histocompatibility complex class I-restricted peptide affinity is a determining factor in the HLA risk profile for the development of postherpetic neuralgia. J Virol 89(2):962–969. https://doi.org/10.1128/jvi.02500-14
Mori Y, Yamanishi K (2007) HHV-6A, 6B, and 7: pathogenesis, host response, and clinical disease. In A. Arvin, G. Campadelli-Fiume, E. Mocarski, P. S. Moore, B. Roizman, R. Whitley, & K. Yamanishi (Eds.), Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis. https://www.ncbi.nlm.nih.gov/pubmed/21348085
Muriuki BM, Forconi CS, Oluoch PO, Bailey JA, Ghansah A, Moormann AM, Ong’echa JM (2021) Association of killer cell immunoglobulin-like receptors with endemic Burkitt lymphoma in Kenyan children. Sci Rep 11(1):11343. https://doi.org/10.1038/s41598-021-90596-7
Naiyer MM, Cassidy SA, Magri A, Cowton V, Chen K, Mansour S, Kranidioti H, Mbiribindi B, Rettman P, Harris S, Fanning LJ, Mulder A, Claas FHJ, Davidson AD, Patel AH, Purbhoo MA, Khakoo SI (2017) KIR2DS2 recognizes conserved peptides derived from viral helicases in the context of HLA-C. Sci Immun 2(15). https://doi.org/10.1126/sciimmunol.aal5296
Neefjes J, Jongsma MLM, Paul P, Bakke O (2011) Towards a systems understanding of MHC class i and MHC class II antigen presentation. In 11:823–836
Niehrs A, Garcia-Beltran WF, Norman PJ, Watson GM, Hölzemer A, Chapel A, Richert L, Pommerening-Röser A, Körner C, Ozawa M, Martrus G, Rossjohn J, Lee JH, Berry R, Carrington M, Altfeld M (2019) A subset of HLA-DP molecules serve as ligands for the natural cytotoxicity receptor NKp44. In 20:1129–1137: Nature Publishing Group
Niens M, Jarrett RF, Hepkema B, Nolte IM, Diepstra A, Platteel M, Kouprie N, Delury CP, Gallagher A, Visser L, Poppema S, Te Meerman GJ, Van Den Berg A (2007) HLA-A*02 is associated with a reduced risk and HLA-A*01 with an increased risk of developing EBV+ Hodgkin lymphoma. Blood 110(9):3310–3315. https://doi.org/10.1182/blood-2007-05-086934
Nowak J, Gwozdowicz S, Graczyk-Pol E, Mika-Witkowska R, Rogatko-Koros M, Nestorowicz K, Szlendak U, Malinowska A, Kaczmarek B, Nasilowska-Adamska B, Tormanowska M, Szczurowska N, Szypnicki J, Witkowska A, Lasota M, Malinowska E, Halaburda K (2019) Epstein-Barr virus infections are strongly dependent on activating and inhibitory KIR-HLA pairs after T-cell replate unrelated hematopoietic stem cell transplantation, the principles, and method of pairing analysis. HLA 94(S2):40–48. https://doi.org/10.1111/tan.13770
Ouwendijk WJD, Laing KJ, Verjans GM, Koelle DM (2013) T-cell immunity to human alphaherpesviruses. In 3:452–460:Elsevier B.V
Ozawa A, Sasao Y, Iwashita K, Miyahara M, Sugai J, Iizuka M, Kawakubo Y, Ohkido M, Naruse T, Anzai T, Takashige N, Ando A, Inoko H (1999) HLA-A33 and -B44 and susceptibility to postherpetic neuralgia (PHN). Tissue Antigens 53(3):263–268. https://doi.org/10.1034/j.1399-0039.1999.530306.x
Palmer WH, Hadfield JD, Obbard DJ (2018) RNA-interference pathways display high rates of adaptive protein evolution in multiple invertebrates. Genetics 208(4):1585–1599. https://doi.org/10.1534/genetics.117.300567
Palmer WH, Telford M, Navarro A, Santpere G, Norman PJ (2022) Human herpesvirus diversity is altered in HLA class I binding peptides. Proc Natl Acad Sci U S A 119(18):e2123248119. https://doi.org/10.1073/pnas.2123248119
Paludan SR, Bowie AG, Horan KA, Fitzgerald KA (2011) Recognition of herpesviruses by the innate immune system. In (Vol. 11, pp. 143–154): Nature Publishing Group
Pappworth IY, Wang EC, Rowe M (2007) The switch from latent to productive infection in Epstein-Barr virus-infected B cells is associated with sensitization to NK cell killing. J Virol 81(2):474–482. https://doi.org/10.1128/jvi.01777-06
Parham P, Lomen CE, Lawlor DA, Ways JP, Holmes N, Coppin HL, Salter RD, Wan AM, Ennis PD (1988) Nature of polymorphism in HLA-A, -B, and -C molecules. Proc Natl Acad Sci USA 85(11):4005–4009. https://doi.org/10.1073/pnas.85.11.4005
Parks S, Avramopoulos D, Mulle J, McGrath J, Wang R, Goes FS, Conneely K, Ruczinski I, Yolken R, Pulver AE, Pearce BD (2018) HLA typing using genome wide data reveals susceptibility types for infections in a psychiatric disease enriched sample. Brain Behav Immun 70:203–213. https://doi.org/10.1016/j.bbi.2018.03.001
Pazmany L, Mandelboim O, Valés-Gómez M, Davis DM, Reyburn HT, Strominger JL (1996) Protection from natural killer cell-mediated lysis by HLA-G expression on target cells. Science 274(5288):792–795. https://doi.org/10.1126/science.274.5288.792
Picarda G, Benedict CA (2018) Cytomegalovirus: shape-shifting the immune system. J Immunol 200(12):3881–3889. https://doi.org/10.4049/jimmunol.1800171
Pierini F, Lenz TL (2018) Divergent allele advantage at human MHC genes: signatures of past and ongoing selection. Mol Biol Evol 35(9):2145–2158. https://doi.org/10.1093/molbev/msy116
Pietra G, Romagnani C, Mazzarino P, Falco M, Millo E, Moretta A, Moretta L, Mingari MC (2003) HLA-E-restricted recognition of cytomegalovirus-derived peptides by human CD8+ cytolytic T lymphocytes. Proc Natl Acad Sci USA 100(19):10896–10901. https://doi.org/10.1073/pnas.1834449100
Pollock NR, Harrison GF, Norman PJ (2022) Immunogenomics of killer cell immunoglobulin-like receptor (KIR) and HLA class I: coevolution and consequences for human health. J Allergy Clin Immunol Pract 10(7):1763–1775. https://doi.org/10.1016/j.jaip.2022.04.036
Price AM, Luftig MA (2015) To be or not IIb: a multi-step process for Epstein-Barr virus latency establishment and consequences for B cell tumorigenesis. In (Vol. 11): Public Library of Science
Prod’homme, V., Griffin, C., Aicheler, R. J., Wang, E. C. Y., McSharry, B. P., Rickards, C. R., Stanton, R. J., Borysiewicz, L. K., López-Botet, M., Wilkinson, G. W. G., & Tomasec, P. (2007) The human cytomegalovirus MHC class I homolog UL18 inhibits LIR-1 + but activates LIR-1 − NK cells. J Immunol 178(7):4473–4481. https://doi.org/10.4049/jimmunol.178.7.4473
Qiang Q, Zhengde X, Chunyan L, Zhizhuo H, Junmei X, Junhong A, Zheng C, Henter JI, Kunling S (2012) Killer cell immunoglobulin-like receptor gene polymorphisms predispose susceptibility to Epstein-Barr virus associated hemophagocytic lymphohistiocytosis in Chinese children. Microbiol Immunol 56(6):378–384. https://doi.org/10.1111/j.1348-0421.2012.00443.x
Rabinstein AA (2017) Herpes virus encephalitis in adults: current knowledge and old myths. Neurol Clin 35(4):695–705. https://doi.org/10.1016/j.ncl.2017.06.006
Racimo F, Sankararaman S, Nielsen R, Huerta-Sánchez E (2015) Evidence for archaic adaptive introgression in humans. In (Vol. 16, pp. 359–371): Nature Publishing Group
Radwan J, Babik W, Kaufman J, Lenz TL, Winternitz J (2020) Advances in the evolutionary understanding of MHC polymorphism. In (Vol. 36, pp. 298–311): Elsevier Ltd
Reche PA, Reinherz EL (2003) Sequence variability analysis of human class I and class II MHC molecules: functional and structural correlates of amino acid polymorphisms. J Mol Biol 331(3):623–641. https://doi.org/10.1016/S0022-2836(03)00750-2
Ressing ME, Van Leeuwen D, Verreck FAW, Gomez R, Heemskerk B, Toebes M, Mullen MM, Jardetzky TS, Longnecker R, Schilham MW, Ottenhoff THM, Neefjes J, Schumacher TN, Hutt-Fletcher LM, Wiertz EJHJ (2003) Interference with T cell receptor-HLA-DR interactions by Epstein-Barr virus gp42 results in reduced T helper cell recognition. Proc Natl Acad Sci USA 100(20):11583–11588. https://doi.org/10.1073/pnas.2034960100
Ribechini E, Fortini C, Marastoni M, Traniello S, Spisani S, Monini P, Gavioli R (2006) Identification of CD8 + T cell epitopes within lytic antigens of human herpes virus 8. J Immunol 176(2):923–930. https://doi.org/10.4049/jimmunol.176.2.923
Rizzo R, Bortolotti D, Gentili V, Rotola A, Bolzani S, Caselli E, Tola MR, Di Luca D (2019) KIR2DS2/KIR2DL2/HLA-C1 haplotype is associated with Alzheimer’s disease: implication for the role of herpesvirus infections. J Alzheimers Dis 67(4):1379–1389. https://doi.org/10.3233/JAD-180777
Robey RC, Mletzko S, Gotch FM (2010) The T-cell immune response against Kaposi’s sarcoma-associated herpesvirus. Advances in Virology 2010:340356–340356. https://doi.org/10.1155/2010/340356
Robinson J, Barker DJ, Georgiou X, Cooper MA, Flicek P, Marsh SGE (2020) IPD-IMGT/HLA database. Nucleic Acids Res 48(D1):D948–D955. https://doi.org/10.1093/nar/gkz950
Rock KL, Reits E, Neefjes J (2016) Present yourself! By MHC class I and MHC class II molecules. In (Vol. 37, pp. 724–737): Elsevier Ltd
Rölle A, Pollmann J, Ewen EM, Le VTK, Halenius A, Hengel H, Cerwenka A (2014) IL-12-producing monocytes and HLA-E control HCMV-driven NKG2C+ NK cell expansion. J Clin Investig 124(12):5305–5316. https://doi.org/10.1172/JCI77440
Rubicz R, Yolken R, Drigalenko E, Carless MA, Dyer TD, Bauman L, Melton PE, Kent JW, Harley JB, Curran JE, Johnson MP, Cole SA, Almasy L, Moses EK, Dhurandhar NV, Kraig E, Blangero J, Leach CT, Göring HHH (2013) A genome-wide integrative genomic study localizes genetic factors influencing antibodies against Epstein-Barr virus nuclear antigen 1 (EBNA-1). PLoS Genetics 9(1). https://doi.org/10.1371/journal.pgen.1003147
Russell AS, Schlaut J (1975) HL-A transplantation antigens in subjects susceptible to recrudescent herpes labialis. Tissue Antigens 6(3):257–261. https://doi.org/10.1111/j.1399-0039.1975.tb00640.x
Sallah N, Carstensen T, Wakeham K, Bagni R, Labo N, Pollard MO, Gurdasani D, Ekoru K, Pomilla C, Young EH, Fatumo S, Asiki G, Kamali A, Sandhu M, Kellam P, Whitby D, Barroso I, Newton R (2017) Whole-genome association study of antibody response to Epstein-Barr virus in an African population: a pilot. Global Health, Epidemiology and Genomics 2:e18–e18. https://doi.org/10.1017/gheg.2017.16
Sallah N, Miley W, Labo N, Carstensen T, Fatumo S, Gurdasani D, Pollard MO, Dilthey AT, Mentzer AJ, Marshall V, Cornejo Castro EM, Pomilla C, Young EH, Asiki G, Hibberd ML, Sandhu M, Kellam P, Newton R, Whitby D, Barroso I (2020) Distinct genetic architectures and environmental factors associate with host response to the γ2-herpesvirus infections. Nat Commun 11(1):3849–3849. https://doi.org/10.1038/s41467-020-17696-2
Samandary S, Kridane-Miledi H, Sandoval JS, Choudhury Z, Langa-Vives F, Spencer D, Chentoufi AA, Lemonnier FA, BenMohamed L (2014) Associations of HLA-A, HLA-B and HLA-C alleles frequency with prevalence of herpes simplex virus infections and diseases across global populations: implication for the development of an universal CD8+ T-cell epitope-based vaccine. Hum Immunol 75(8):715–729. https://doi.org/10.1016/j.humimm.2014.04.016
Santpere G, Darre F, Blanco S, Alcami A, Villoslada P, Mar Albà M, Navarro A (2014) Genome-wide analysis of wild-type Epstein-Barr virus genomes derived from healthy individuals of the 1,000 Genomes Project. Genome Biol Evol 6(4):846–860. https://doi.org/10.1093/gbe/evu054
Sato M, Ohashi J, Tsuchiya N, Kashiwase K, Ishikawa Y, Arita H, Hanaoka K, Tokunaga K, Yabe T (2002) Association of HLA-A*3303-B*4403-DRB1*1302 haplotype, but not of TNFA promoter and NKp30 polymorphism, with postherpetic neuralgia (PHN) in the Japanese population. Genes Immun 3(8):477–481. https://doi.org/10.1038/sj.gene.6363890
Sato-Takeda M, Ihn H, Ohashi J, Tsuchiya N, Satake M, Arita H, Tamaki K, Hanaoka K, Tokunaga K, Yabe T (2004) The human histocompatibility leukocyte antigen (HLA) haplotype is associated with the onset of postherpetic neuralgia after herpes zoster. Pain 110(1–2):329–336. https://doi.org/10.1016/j.pain.2004.04.010
Schmidt NJ, Lennette EH, Magoffin RL (1969) Immunological relationship between herpes simplex and varicella-zoster viruses demonstrated by complement-fixation, neutralization and fluorescent antibody tests. J Gen Virol 4(3):321–328. https://doi.org/10.1099/0022-1317-4-3-321
Shukla SA, Rooney MS, Rajasagi M, Tiao G, Dixon PM, Lawrence MS, Stevens J, Lane WJ, Dellagatta JL, Steelman S, Sougnez C, Cibulskis K, Kiezun A, Hacohen N, Brusic V, Wu CJ, Getz G (2015) Comprehensive analysis of cancer-associated somatic mutations in class i HLA genes. Nat Biotechnol 33(11):1152–1158. https://doi.org/10.1038/nbt.3344
Sim MJW, Rajagopalan S, Altmann DM, Boyton RJ, Sun PD, Long EO (2019) Human NK cell receptor KIR2DS4 detects a conserved bacterial epitope presented by HLA-C. Proc Natl Acad Sci USA 116(26):12964–12973. https://doi.org/10.1073/pnas.1903781116
Simons MJ, Wee GB, Chan SH, Shanmugaratnam K, Day NE, de-Thé G (1975) Immunogenetic aspects of nasopharyngeal carcinoma (NPC) III. HL-a type as a genetic marker of NPC predisposition to test the hypothesis that Epstein-Barr virus is an etiological factor in NPC. IARC Sci Pub (11 Pt 2):249–258
Sirianni MC, Vincenzi L, Topino S, Giovannetti A, Mazzetta F, Libi F, Scaramuzzi D, Andreoni M, Pinter E, Baccarini S, Rezza G, Monini P, Ensoli B (2002) NK cell activity controls human herpesvirus 8 latent infection and is restored upon highly active antiretroviral therapy in AIDS patients with regressing Kaposi’s sarcoma. Eur J Immunol 32(10):2711–2720. https://doi.org/10.1002/1521-4141(2002010)32:10%3c2711::AID-IMMU2711%3e3.0.CO;2-3
Smith C, Khanna R (2013) Immune regulation of human herpesviruses and its implications for human transplantation. Am J Transplant 13(SUPPL. 3):9–23. https://doi.org/10.1111/ajt.12005
Srivastava R, Khan AA, Spencer D, Vahed H, Lopes PP, Thai NTU, Wang C, Pham TT, Huang J, Scarfone VM, Nesburn AB, Wechsler SL, BenMohamed L (2015) HLA-A02:01–restricted epitopes identified from the herpes simplex virus tegument protein VP11/12 preferentially recall polyfunctional effector memory CD8 + T cells from seropositive asymptomatic individuals and protect humanized HLA-A*02:01 transgenic mice against ocular herpes. J Immunol 194(5):2232–2248. https://doi.org/10.4049/jimmunol.1402606
Stern H (1978) Correlation between cytomegalovirus infection and HLA-BW15. In (Vol. 2, pp. 126–126): BMJ Publishing Group
Stern M, Hadaya K, Hönger G, Martin PY, Steiger J, Hess C, Villard J (2011) Telomeric rather than centromeric activating KIR genes protect from cytomegalovirus infection after kidney transplantation. Am J Transplant 11(6):1302–1307. https://doi.org/10.1111/j.1600-6143.2011.03516.x
Stewart CA, Laugier-Anfossi F, Vély F, Saulquin X, Riedmuller J, Tisserant A, Gauthiers L, Romagné F, Ferracci G, Arosa FA, Moretta A, Sun PD, Ugolini S, Vivier E (2005) Recognition of peptide-MHC class I complexes by activating killer immunoglobulin-like receptors. Proc Natl Acad Sci USA 102(37):13224–13229. https://doi.org/10.1073/pnas.0503594102
Strowig T, Brilot F, Arrey F, Bougras G, Thomas D, Muller WA, Münz C (2008) Tonsilar NK cells restrict B cell transformation by the Epstein-Barr virus via IFN-γ. PLoS Pathogens 4(2). https://doi.org/10.1371/journal.ppat.0040027
Su WH, Hildesheim A, Chang YS (2013) Human leukocyte antigens and Epstein-Barr virus-associated nasopharyngeal carcinoma: old associations offer new clues into the role of immunity in infection-associated cancers. In (Vol. 3 DEC): Front Med SA
Sumiyama D, Kikkawa EF, Kita YF, Shinagawa H, Mabuchi T, Ozawa A, Inoko H (2008) HLA alleles are associated with postherpetic neuralgia but not with herpes zoster. Tokai J Exp Clin Med 33(4):150–153
Sylwester AW, Mitchell BL, Edgar JB, Taormina C, Pelte C, Ruchti F, Sleath PR, Grabstein KH, Hosken NA, Kern F, Nelson JA, Picker LJ (2005) Broadly targeted human cytomegalovirus-specific CD4+ and CD8+ T cells dominate the memory compartments of exposed subjects. J Exp Med 202(5):673–685. https://doi.org/10.1084/jem.20050882
Takahata N, Nei M (1990) Allelic genealogy under overdominant and frequency-dependent selection and polymorphism of major histocompatibility complex loci. Genetics 124(4):967–978
Tang M, Lautenberger JA, Gao X, Sezgin E, Hendrickson SL, Troyer JL, David VA, Guan L, McIntosh CE, Guo X, Zheng Y, Liao J, Deng H, Malasky M, Kessing B, Winkler CA, Carrington M, dé The G, Zeng Y, O'Brien SJ (2012) The principal genetic determinants for nasopharyngeal carcinoma in China involve the HLA class I antigen recognition groove. PLoS Genetics 8(11). https://doi.org/10.1371/journal.pgen.1003103
Taylor GS, Long HM, Brooks JM, Rickinson AB, Hislop AD (2015) The immunology of Epstein-Barr virus–induced disease. Annu Rev Immunol 33(1):787–821. https://doi.org/10.1146/annurev-immunol-032414-112326
Thorsby E (2009) A short history of HLA. In (Vol. 74, pp. 101–116)
Tian C, Hromatka BS, Kiefer AK, Eriksson N, Noble SM, Tung JY, Hinds DA (2017) Genome-wide association and HLA region fine-mapping studies identify susceptibility loci for multiple common infections. Nat Commun 8(1):599–599. https://doi.org/10.1038/s41467-017-00257-5
Tomasec P, Braud VM, Rickards C, Powell MB, McSharry BP, Gadola S, Cerundolo V, Borysiewicz LK, McMichae AJ, Wilkinson GWG (2000) Surface expression of HLA-E, an inhibitor of natural killer cells, enhanced by human cytomegalovirus gpUL40. Science 287(5455):1031–1033. https://doi.org/10.1126/science.287.5455.1031
Tse KP, Su WH, Chang KP, Tsang NM, Yu CJ, Tang P, See LC, Hsueh C, Yang ML, Hao SP, Li HY, Wang MH, Liao LP, Chen LC, Lin SR, Jorgensen TJ, Chang YS, Shugart YY (2009) Genome-wide association study reveals multiple nasopharyngeal carcinoma-associated loci within the HLA region at chromosome 6p21.3. Am J Human Gen 85(2):194–203. https://doi.org/10.1016/j.ajhg.2009.07.007
Unanue ER, Turk V, Neefjes J (2016) Variations in MHC class II antigen processing and presentation in health and disease. Annu Rev Immunol 34(1):265–297. https://doi.org/10.1146/annurev-immunol-041015-055420
van der Ploeg K, Chang, C, Ivarsson MA, Moffett A, Wills MR, Trowsdale J (2017) Modulation of human leukocyte antigen-C by human cytomegalovirus stimulates KIR2DS1 recognition by natural killer cells. Front Immunol 8(MAR). https://doi.org/10.3389/fimmu.2017.00298
van Duin D, Avery RK, Hemachandra S, Yen-Lieberman B, Zhang A, Jain A, Butler RS, Barnard J, Schold JD, Fung J, Askar M (2014) KIR and HLA interactions are associated with control of primary CMV infection in solid organ transplant recipients. Am J Transplant 14(1):156–162. https://doi.org/10.1111/ajt.12532
van Velzen M, Jing L, Osterhaus ADME, Sette A, Koelle DM, Verjans GMGM (2013) Local CD4 and CD8 T-cell reactivity to HSV-1 antigens documents broad viral protein expression and immune competence in latently infected human trigeminal ganglia. PLoS Pathogens 9(8). https://doi.org/10.1371/journal.ppat.1003547
Vider-Shalit T, Fishbain V, Raffaeli S, Louzoun Y (2007) Phase-dependent immune evasion of herpesviruses. J Virol 81(17):9536–9545. https://doi.org/10.1128/JVI.02636-06
Vider-Shalit T, Sarid R, Maman K, Tsaban L, Levi R, Louzoun Y (2009) Viruses selectively mutate their CD8+ T-cell epitopes—a large-scale immunomic analysis. Bioinformatics 25(12):i39–i44. https://doi.org/10.1093/bioinformatics/btp221
Vietzen H, Rückert T, Hartenberger S, Honsig C, Jaksch P, Geleff S, Hammer Q, Romagnani C, Segura-Wang M, Puchhammer-Stöckl E, Paraskevis D (2021) Extent of cytomegalovirus replication in the human host depends on variations of the HLA-E/UL40 axis. mBio 12(2):e02996–02920. https://doi.org/10.1128/mBio.02996-20
Vivier E, Raulet DH, Moretta A, Caligiuri MA, Zitvogel L, Lanier LL, Yokoyama WM, Ugolini S (2011) Innate or adaptive immunity? The example of natural killer cells. Science 331(6013):44–49. https://doi.org/10.1126/science.1198687
Wegner F, Lassalle F, Depledge DP, Balloux F, Breuer J (2019) Co-evolution of sites under immune selection shapes Epstein-Barr virus population structure. Mol Biol Evol. https://doi.org/10.1093/molbev/msz152
Weinberg A, Zhang JH, Oxman MN, Johnson GR, Hayward AR, Caulfield MJ, Irwin MR, Clair J, Smith JG, Stanley H, Marchese RD, Harbecke R, Williams HM, Chan ISF, Arbeit RD, Gershon AA, Schödel F, Morrison VA, Kauffman CA, Levin MJ (2009) Varicella-zoster virus–specific immune responses to herpes zoster in elderly participants in a trial of a clinically effective zoster vaccine. J Infect Dis 200(7):1068–1077. https://doi.org/10.1086/605611
White DW, Suzanne Beard R, Barton ES (2012) Immune modulation during latent herpesvirus infection. Immunol Rev 245(1):189–208. https://doi.org/10.1111/j.1600-065X.2011.01074.x
Williams H, McAulay K, Macsween KF, Gallacher NJ, Higgins CD, Harrison N, Swerdlow AJ, Crawford DH (2005) The immune response to primary EBV infection: a role for natural killer cells. Br J Haematol 129(2):266–274. https://doi.org/10.1111/j.1365-2141.2005.05452.x
Wills MR, Ashiru O, Reeves MB, Okecha G, Trowsdale J, Tomasec P, Wilkinson GWG, Sinclair J, Sissons JGP (2005) Human cytomegalovirus encodes an MHC class I-like molecule (UL142) that functions to inhibit NK cell lysis1. J Immunol 175(11):7457–7465. https://doi.org/10.4049/jimmunol.175.11.7457
Yu K, Davidson CL, Wójtowicz A, Lisboa L, Wang T, Airo AM, Villard J, Buratto J, Sandalova T, Achour A, Humar A, Boggian K, Cusini A, Van Delden C, Egli A, Manuel O, Mueller N, Bochud PY, Burshtyn DN (2018) LILRB1 polymorphisms influence posttransplant HCMV susceptibility and ligand interactions. J Clin Investig 128(4):1523–1537. https://doi.org/10.1172/JCI96174
Zaia JA, Sun JY, Gallez-Hawkins GM, Thao L, Oki A, Lacey SF, Dagis A, Palmer J, Diamond DJ, Forman SJ, Senitzer D (2009) The effect of single and combined activating killer immunoglobulin-like receptor genotypes on cytomegalovirus infection and immunity after hematopoietic cell transplantation. Biol Blood Marrow Transplant 15(3):315–325. https://doi.org/10.1016/j.bbmt.2008.11.030
Zerboni L, Sen N, Oliver SL, Arvin AM (2014) Molecular mechanisms of varicella zoster virus pathogenesis. Nat Rev Microbiol 12(3):197–210. https://doi.org/10.1038/nrmicro3215
Zhu J, Koelle DM, Cao J, Vazquez J, Meei LH, Hladik F, Wald A, Corey L (2007) Virus-specific CD8+ T cells accumulate near sensory nerve endings in genital skin during subclinical HSV-2 reactivation. J Exp Med 204(3):595–603. https://doi.org/10.1084/jem.20061792
Funding
This work was supported by National Institutes of Health (NIH) NIAID grant R01 AI158410 (to P.J.N.). W.H.P. is supported by NIH NIAID grant F32 AI161790.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Palmer, W.H., Norman, P.J. The impact of HLA polymorphism on herpesvirus infection and disease. Immunogenetics 75, 231–247 (2023). https://doi.org/10.1007/s00251-022-01288-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-022-01288-z