Abstract
Pollinators face many stressors, including reduced floral diversity. A low-diversity diet can impair organisms’ ability to cope with additional stressors, such as pathogens, by altering the gut microbiome and/or immune function, but these effects are understudied for most pollinators. We investigated the impact of pollen diet diversity on two ecologically and economically important generalist pollinators, the social bumble bee (Bombus impatiens) and the solitary alfalfa leafcutter bee (Megachile rotundata). We experimentally tested the effect of one-, two-, or three-species pollen diets on gut bacterial communities in both species, and the melanization immune response in B. impatiens. Pollen diets included dandelion (Taraxacum officinale), staghorn sumac (Rhus typhina), and hawthorn (Crataegus sp.) alone, each pair-wise combination, or a mix of all three species. We fed bees their diet for 7 days and then dissected out guts and sequenced 16S rRNA gene amplicons to characterize gut bacterial communities. To assess melanization in B. impatiens, we inserted microfilament implants into the bee abdomen and measured melanin deposition on the implant. We found that pollen diet did not influence gut bacterial communities in M. rotundata. In B. impatiens, pollen diet composition, but not diversity, affected gut bacterial richness in older, but not newly-emerged bees. Pollen diet did not affect the melanization response in B. impatiens. Our results suggest that even a monofloral, low-quality pollen diet such as dandelion can support diverse gut bacterial communities in captive-reared adults of these bee species. These findings shed light on the effects of reduced diet diversity on bee health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Reduced diet diversity due to landscape simplification can negatively impact wildlife populations by limiting nutrient availability and increasing foraging time. Low-diversity diets lacking protein and other nutrients can impair animal performance, particularly by weakening strength and immune function (e.g., [1]). Such effects of diet are often mediated by the gut microbiome, which can modulate host immunity, digestion, and metabolism [2]. Diet can influence gut microbial communities (“microbiota” hereafter) by introducing foodborne microbes and/or nutrients that alter dynamics of the existing community [3]. Thus, diet can influence host health directly and through its effects on the gut microbiome.
Bees are critical pollinators, and many populations have shown recent declines [4]. Reduced floral diversity has been implicated in those declines [5], yet the mechanisms behind this trend remain poorly understood for most taxa. Studies on bee nutrition, immunity, and the gut microbiota have mostly focused on the social and commonly managed taxa, such as honey bees and bumble bees (reviewed in [6]). However, the vast majority of bee species (~ 80%), including many important wild bee pollinators, are solitary [7].
Previous studies have found varying effects of diet diversity on bee growth, development, and immune function. For example, bumble bees produced smaller and fewer offspring when fed a low-protein pollen (Cistus sp.) compared to those fed a mixed pollen diet [8]. In the field, Andrena bees in more diverse, natural landscapes were larger than those in homogenous agricultural landscapes [9] and Osmia nests with more diverse pollen provisions produced more female offspring [10]. Diet can also affect bee immunity. For example, bumble bees fed a sunflower pollen-only diet had different immune gene expression profiles compared to bees fed a polyfloral diet [11]. These studies demonstrate links between diet diversity, longevity, and pathogen resistance in a limited number of bee species, but the mechanistic links between diet diversity and bee health remain poorly understood.
Diet can influence the bee gut microbiota, but this has only been investigated in a few studies using social bees [12,13,14]; dietary effects on solitary bee gut microbes have not been assessed. Social behavior strongly influences routes of acquisition of gut microbes in bees and other animals. Thus, the gut microbiota in the highly social, corbiculate bee species is made up of a relatively small and consistent group of coevolved taxa [15] that contribute to individual digestion, growth, immune modulation, and detoxification (e.g., [16]). Solitary bees, however, typically host more diverse gut microbes. While some solitary bees host a core group of bacteria [17], which can overlap with social bees [18]), their microbes are primarily acquired from the environment rather than within-nest social contact [19]. The functions of solitary bee microbiota are also less understood, although microbes on pollen provisions are important for larval development [20]. Due to their different gut microbiota, bees that differ in sociality may respond differently to changes in diet breadth, such that studies on social bees may not translate to solitary bees.
We tested the effect of one-, two-, and three-species pollen diets on immunity and the gut microbiota of the social common eastern bumble bee, Bombus impatiens, and the gut microbiota of the solitary alfalfa leafcutter bee, Megachile rotundata. We used sterilized pollen to test solely the effects of pollen nutrient input on the existing microbial community, rather than evaluating pollen as a source of novel microbial diversity. We predicted that higher diet diversity would result in higher gut microbial diversity and a stronger immune response. Further, we predicted that the gut microbiota of M. rotundata would reflect changes in diet more than B. impatiens due to B. impatiens’ coevolution with a core group of bacterial taxa. Lastly, we predicted that dandelion pollen would result in a microbiota rich in Lactobacillaceae, since these bacteria have been associated with Asteraceae pollen in bumble bee pollen baskets [21] and Megachile nests [22]. Understanding the mechanisms by which diet diversity affects both social and solitary wild bees is critical for supporting their populations, particularly in agricultural settings where crop monocultures are common.
Methods
Overview
We tested the effect of pollen diet diversity on gut bacterial communities in Bombus impatiens and Megachile rotundata, and on the melanization immune response in B. impatiens (Fig. 1). For all experiments, we placed bees in individual containers with assigned pollen diets (sterilized) and 30% sucrose solution (not sterilized). Sucrose was accessible to the bee through a cotton wick. We replaced sucrose and pollen every other day and measured consumption over a 48-hr period. After 7 d, we anesthetized bees on ice and froze them at -80 °C for later processing. We collected the right forewing and measured marginal cell length as a proxy for bee size. Additional details for all methods can be found in the supplement.
Pollen Diets
We used pollen from dandelion (Taraxacum officinale, Asteraceae), staghorn sumac (Rhus typhina, Anacardiaceae), and hawthorn (Crataegus sp., Rosaceae). Pollens were honey-bee collected from Quebec, Canada and confirmed to be > 95% one species by microscopic assessment of morphology (using fuchsin dyed glycerin jelly). We chose these species in part because they are abundant bee-collected resources in an area where B. impatiens is common, and in part because we could obtain a sufficient quantity from these species of single-species pollen for feeding trials. These pollens differ in nutritional content (see Supplement for references), and Taraxacum pollen may have chemical or mechanical defenses that reduce performance in B. terrestris [23]. Pollens were sterilized by ethylene oxide (USDA, Logan, Utah [24]; see Supplement for protocol), ground by mortar and pestle, and sorted into seven treatments: each species alone (three 1-species treatments), each pair-wise combination (three 2-species treatments), and one combination of all three species. We made multifloral treatments by combining the pollen species in equal parts by weight and then mixed each treatment with deionized water to make a paste (see Table S2 for pollen:water ratios). We measured pollen and sucrose consumption per bee in the first 48 h by weighing before and after delivery.
Bombus impatiens Gut Microbiota
We used Bombus impatiens (Apidae) workers derived from wild-caught queens collected in Amherst, Massachusetts in spring of 2021. Bombus impatiens are generalist foragers, and workers consume pollen in addition to foraging for pollen for the colony (personal observations). We reared queens in the laboratory until colonies were large enough to remove workers without affecting colony survival. Colonies were fed non-sterilized, honey bee-collected wildflower mix pollen (Koppert Biological Systems, Howell, Michigan). All queens were screened for Crithidia sp. infections via microscopy of feces, and only uninfected queens were used.
We used 90 workers from six colonies, of which 56 were newly emerged, also known as callows (identified by silver body hair, which turns black after about 1 day post-eclosion). We placed callows in individual vials for 2 h, and hand-inoculated them with 10–15 µL of pooled feces from five nestmates mixed with 50% sucrose. Similar inoculation protocols result in microbiomes indistinguishable from those of mature, in-colony workers [25, 26]. We initially only used callows to control for worker age and starting microbiome. However, callows were reluctant to consume fecal inoculum, and we switched to non-callow adults for the last 34 bumble bees. The non-callows were taken straight from their colony and did not receive fecal inoculum; we presume they were inoculated with nestmate feces prior to removal (bumble bees’ gut microbiomes stabilize within 4 days of eclosion; [27]). We included age (callow/non-callow) in analyses. Bees were housed in 16-oz deli cups in darkness with access to 30% sucrose and their pollen diet. Bumble bees were placed in the experiment on ten start dates between August 11–26, 2021. Five bees died, resulting in 85 samples (Table S1).
Megachile rotundata Gut Microbiota
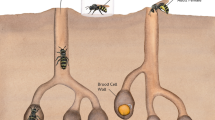
We used commercial Megachile rotundata (Megachilidae; masonbeesforsale.com, Deweyville, Utah). Bees arrived as pupae, which we kept at 7 °C until use. We then placed pupae in darkness inside a mesh cage at 27 °C until individuals emerged (based on methods from [28]. Megachile rotundata is a solitary bee that nests in cavities and lines brood cells with leaves. Females provision cells for a single egg, then seal the cell and provide no additional care to the developing larva. Megachile rotundata are highly valued and managed in North America for alfalfa pollination, but they are also generalist foragers [29].
To ensure that the leafcutter bees were exposed to bacterial cells for an initial microbiome, we created an ecologically-relevant microbial inoculum using deionized water sonicated with flowers collected from Amherst, Massachusetts on the first trial date and frozen at -80 °C in a glycerol solution. The flower species included Solidago sp. (Asteraceae), Cosmos sp. (Asteraceae), Impatiens capensis (Balsaminaceae), Lobelia siphilitica. (Campanulaceae), Trifolium repens (Fabaceae), Pycnanthemum virginianum (Lamiaceae), Satureja hortensis (Lamiaceae), and Allium tuberosum (Amaryllidaceae). We prepared inoculum for each trial by mixing 1:1 flower water and 50% sucrose (see Supplement for additional details). While we are unaware of previous studies that have used such a floral inoculum, the concept is similar to inoculating individual Bombus with feces from nestmates (as in [25, 26]); our goal here was to simulate exposure to a floral microbiome that M. rotundata might encounter from flowers. We placed the bees in their individual containers (60 mm diameter petri dish; with drilled holes) and pipetted 10 µL of the flower inoculum directly onto a cotton wick, which was placed in a 30% sucrose solution. We entered 74 bees in the experiment on September 14, 15 and 16, 2021. Four bees died and two samples could not be PCR amplified, resulting in 68 samples (Table S1).
Microbiome Processing and Analysis
Sample Processing and Sequencing
We stored each bee at -80 °C until gut dissection and DNA extraction. We dissected out the gut of each bee under sterile conditions and placed each (excluding the crop and rectum) into a tissue collection plate (Qiagen, Germantown, Maryland). We included four blank extractions as no-template controls in all downstream procedures and analyses. To characterize bacterial communities, we prepared amplicon libraries using the 799 F (CMGGATTAGATACCCKGG) and 1115R (AGGGTTGCGCTCGTTG) 16S rRNA gene primers (e.g., [30]). The Genomics Core at the University of California, Riverside checked DNA quality and concentration using the 2100 Bioanalyzer (Agilent, Santa Clara, California) and then sequenced the libraries in a single run on the MiSeq (Illumina, San Diego, California) using the V3 2 × 300 reagent kit.
Bioinformatics
We used QIIME2 to process the Illumina fastq files [31]. We removed the barcodes and concatenated them into a separate file to be compatible with QIIME2, and then demultiplexed the sequences. To trim low quality sequences and bin reads into amplicon sequence variants (ASVs), we ran DADA2 with default parameters and read trimming of 253 bases for forward reads and 211 bases for reverse reads [32]. We assigned taxonomy to genus level by using the QIIME2 sklearn classifier trained to the 799 to 1115 region of the SILVA 16S rRNA gene database [33, 34]. We additionally used the NCBI database to conduct local BLAST searches for ASVs that required further classification. We used R version 4.2.1 for decontamination and QIIME2 for final filtering, including the removal of singletons [35]. We identified contaminants using the “prevalence” method in the decontam package [36], with a conservative threshold of 0.5, which identifies ASVs that were more prevalent in negative controls than in samples. We then filtered out the 22 identified contaminants as well as mitochondria and chloroplast ASVs. We rarefied to 8000 reads per sample.
Statistics
To assess community diversity, we used the vegan [37] and phyloseq packages [38] in R [31]. We first ran models to assess community metrics in response to host bee species. After finding significant effects of host species (see Results), we analyzed effects of diet within each host. Models for each species included diet composition (7 levels) or diet diversity (3 levels; 1-, 2- or 3-species), but not both.
To investigate alpha diversity, we modeled the number of ASVs (“ASV richness” hereafter) and Shannon index by diet treatment (both composition and diversity in separate models). For B. impatiens, full models included age (callow/non-callow) and bee size (estimated by wing marginal cell length) as additional fixed effects and start date and colony as random effects. For M. rotundata, full models included bee size as an additional fixed effect, and start date and PCR plate ID as random effects. All final models (except those with host as a predictor) retained diet treatment as a fixed effect since it was our variable of interest. To analyze pollen consumption, we used a similar approach, using a linear model with a normal distribution (lme4 package) [39] and analyzed survival using the coxme function (coxme package [40]), survfit, and Surv functions (survival package [41]). For pollen consumption and survival models, we used pollen diet composition (not diversity) as the treatment predictor. We ran full linear mixed models (lme4 package) [42] and then used Akaike information criterion (AIC) for model selection [43]. We report results from the best-fitting models in the tables. We tested the significance of terms using likelihood ratio χ2 tests. We validated models using the simulateResiduals function [44] and produced figures with emmeans [39] and ggplot [45].
To investigate beta diversity, we calculated Bray-Curtis distance matrices (results using unweighted UniFrac, and weighted UniFrac were generally similar and are reported in Supplement Tables S4, S5). We then analyzed three community metrics: (1) community dispersion among treatment groups using the betadisper function, (2) overall community composition between treatment groups using the adonis function, and (3) similarity percentages to identify taxa that contributed most to the dissimilarity between groups using the simper function. Betadisper models are limited to one predictor variable at a time, therefore we ran dispersion models for each predictor. For the B. impatiens adonis models, we included diet composition, age, and colony as predictors. We did not include start date as a block since it was confounded with age. For M. rotundata, we included diet as a predictor and start date as a block. We visualized beta diversity using Principal Coordinates Analysis (PCoA) ordinations.
Bombus impatiens Melanization
Experimental Design
We measured melanization of an implant in the bee abdomen, a commonly used method for estimating immune function in bumble bees (e.g., [46]). We used 135 B. impatiens workers from three commercial colonies (Koppert Biological Systems, Howell, Michigan), reared in the lab. We placed non-callow workers into individual deli cups with their pollen diet (same as previous experiments) and 30% sucrose. On day 7, we anesthetized bees by placing them in the freezer for 3–5 min, punctured the pleural membrane between the 3rd and 4th abdominal tergite using a sterile needle, and then inserted a sterile, transparent 1-mm long microfilament (Sufix Elite) into the abdominal cavity using forceps. The bee then recovered in its container for 2 h in a dark room with access to sucrose and its pollen diet. After 2 h, bees were frozen at -20 °C. We later dissected abdomens to recover implants, mounted implants to microscope slides using clear nail polish, and photographed implants using a dissecting microscope at 20X magnification. Bees initiated the experiment on five dates between November 29-December 13, 2021. Twelve bees died, 7 escaped, 9 died after being anesthetized, and we could not find the implants in 5 bees, resulting in 102 samples (Table S1). We imported photographs of each implant to ImageJ [47] and estimated percent of the implant covered by melanin (see Supplement for further methods and discussion).
Statistics
We modeled proportion melanization with a generalized linear mixed model with a beta error distribution [48]. Initial models included diet treatments (composition or diversity) and bee size as fixed effects and start date and natal colony as random effects. We built separate models using diet composition or diversity as predictors and performed model selection using AIC.
Results
Gut Microbiomes in Both Host Species
After quality control and removing contaminants, we retained an average of 25,177 reads per sample across 153 samples. We identified 334 ASVs. Top models and results assessing effect of host species are in Table 1. The host species differed significantly in community composition (Fig. 2C). Compared to M. rotundata, B. impatiens had higher alpha diversity (Fig. 2A and B) and wider dispersion among replicates (Fig. 2C). The bacterial taxa that appear to drive dissimilarities between hosts, based on the simper function, were Neokomagataea, Rosenbergiella, Pantoea, and ASVs that matched to genera within the Enterobacteriaceae, which were prominent in M. rotundata guts, while B. impatiens harbored more Candidatus Schmidhempelia and Snodgrassella, which are common members of their core microbiota.
Bombus impatiens Gut Microbiome
Bumble bee guts contained previously-documented core members, including Snodgrassella, Bifidobacterium, Lactobacillus, and Candidatus Schmidhempelia (Fig. 3A). However, they were missing a commonly dominant member, Gilliamella [15]. Communities also included other genera previously associated with bees and flowers, Acinetobacter, Asaia, Pantoea [19], Neokomagataea [49]) and soil bacteria (ASVs in Enterobacteriaceae, multiple of which shared 100% sequence identity to Citrobacter, Enterobacter, and Pantoea species).
For B. impatiens, top models and results are reported in Table 2. We found a significant diet composition by age interaction on ASV richness (Fig. 2D), and subsequently assessed effect of diet within callows and non-callows using the joint_tests function. Diet composition affected ASV richness in non-callows, but not callows; non-callows fed the sumac + hawthorn diet had particularly low ASV richness, with an average of 16.2 ASVs (all pairwise comparisons were non-significant except dandelion; z = -3.05, P = 0.035). When diets were combined by levels of diversity, we found no significant effects of diet diversity, bee size, or the diet by age interaction on ASV richness (Fig. 2E). Diet (whether composition or diversity) did not affect Shannon diversity.
Beta diversity was significantly affected by colony, but not diet (composition or diversity), or age (Table S4, Figure S3). Community composition was affected by colony, but not diet composition (Fig. 3A), diversity, or age (Figure S3). One bacterial genus, Bombella, was only present in bees from one colony (detected in 11 out of 18 bees from this colony; seven of which were callows and four non-callows; Figure S4). We also note that Bifidobacterium was detected in 18 bees, however was only found at abundances > 1% in non-callows (Figure S4).
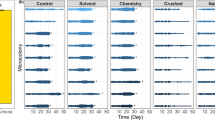
Bee species differed in their gut bacterial communities. Compared to M. rotundata, B. impatiens had higher (A) ASV richness, (B) Shannon diversity, and (C) community variation, visualized as a PCoA ordination of Bray-Curtis dissimilarities. Ellipses are 95% confidence intervals. The effect of diet on bacterial ASV richness in B. impatiens depended on worker age (D, E). Diet did not significantly affect bacterial ASV richness in M. rotundata(F, G). (Diets are D = Dandelion, H = Hawthorn, S = Sumac.)
Megachile rotundata Gut Microbiome
Leafcutter bees guts contained flower-associated bacteria, including Neokomagataea, Rosenbergiella, Acinetobacter, and Asaia, as well as soil-associates, including multiple genera in the Enterobacteriaceae (Klebsiella, Kluyvera, and ASVs that matched Citrobacter, Enterobacter, Klebsiella, and Pantoea; Fig. 3B; [19]). Megachile rotundata contained other genera at lower abundances that were also in bumble bee guts, including Acinetobacter and Asaia (Fig. 3B) as well as Snodgrassella and Candidatus Schmidhempelia (found at < 1% abundance and therefore not included in Fig. 3).
For M. rotundata, top models and results are reported in Table 3. Neither ASV richness nor Shannon diversity was affected by diet composition (Fig. 2F), diversity (Fig. 2G), or bee size (Table S5). Community dispersion and composition were not affected by diet composition, diversity, or start date (Fig. 3B, Figures S5, S6, Table S5).
Bombus impatiens Melanization
Most implant samples (93.2%) showed evidence of melanization. On average, bees melanized about 11% (± 0.984 SE) of the implant surface area. The top models included diet (composition or diversity in separate models) as the predictor and start date and colony as random effects. Proportion melanization was not affected by diet composition (χ2 = 2.693, d.f. = 6, P = 0.846; Fig. 4) nor diversity (χ2 = 1.219, d.f. = 2, P = 0.544).
Discussion
We predicted that a low-diversity pollen diet would result in less diverse gut microbial communities due to lower nutritional diversity, but our results did not support this prediction. We found no obvious relationship between pollen diversity and gut bacterial diversity, suggesting that initial bacterial communities were relatively stable despite nutritional change. Even single-species diets still contributed adequate resources for the persistence of diverse communities over seven days. We used sterile pollen to rule out effects of foodborne microbes, thereby allowing us to attribute any effects of diet on microbial communities to nutritional differences. We recognize this may not be reflective of nature, since pollen commonly harbors its own microbial community [50], but nonetheless sheds light on diet-microbiome dynamics. The stability of these communities suggests that priority effects and drift play important roles in long-term assemblages. Additionally, pollen diet may not affect gut microbiota if these microbes are more dependent on carbohydrates, such as pectin of the pollen wall [51] or nectar sugars, than the proteins and lipids inside pollen. Indeed, nectar qualities such as the concentration of different sugars can affect bumble bee gut bacterial composition [13]. While the relevance of these results may be limited for wild bees, our results suggest that simple, low-diversity diets of sterile pollen can still support established microbial communities.
Our results are consistent with previous studies that found little to no correlation between diets and host microbial diversity. For example, multiple studies found weak or no correlation between landscape, pollen and microbial diversity in pollen provisions of several bee species [22, 30, 52]. Conversely, our results contradict studies that found a positive relationship between diet and host microbial diversity, for example in antelope [53]. These contrasting patterns may be due to host taxa differing in levels of microbial filtering, which can also vary by ecological context [54]. While correlations between dietary and microbial diversity vary, short-term dietary changes can alter gut microbiome composition, demonstrating that certain diets are associated with certain bacterial taxa. For example, in humans, a shift to meat-eating increased clusters of bile-resistant bacteria [3]. In nests of Ceratina bees, Acinetobacter, Lactobacillus, Pantoea, and Sodalis bacteria were positively correlated with pollen from multiple plant genera [30]. Lactobacillus was also positively correlated with Asteraceae pollen in Megachile nests and Bombus corbicula [21, 22]. We thus predicted that dandelion diet would be relatively rich in Lactobacillus, but our results did not support this (Fig. 3, S4), suggesting that gut-inhabiting Lactobacillus does not preferentially feed on Asteraceae pollen. Lactobacillus may be associated with Asteraceae resources in the wild; we would not have observed this relationship due to sterilizing pollen.
Gut bacterial communities did not show distinct clusters by diet treatments in either species (Figure S3, S5), however this was dependent on worker age in bumble bees. Consistent with previous work [13], diet influenced bacterial diversity in older, but not newly emerged workers (Fig. 2D). A recent study found that bumble bee gut microbiomes exhibit a predictable successional trajectory, growing logistically in both abundance and diversity, and plateauing around 4 days old [27]. Given the non-callow group is a much wider age range (any age > 24 h), and therefore has opportunity for more variation, it is surprising that we found a significant effect of diet on diversity in this group and not callows. However, because the callows received a fecal inoculum pooled from multiple non-callows, they did not experience this succession process and may have developed a more diverse and therefore stable microbiome than expected naturally. Future studies using germ-free bees inoculated with controlled microbiomes could assess how bee gut microbiomes are influenced by diet at different life stages and why certain diets, such as sumac and hawthorn, yielded relatively low ASV richness in non-callows.
We found that bumble bees hosted many core bacterial taxa, including Snodgrassella, Lactobacillus, Candidatus Schmidhempelia, and Bifidobacterium, which have been shown to play important roles for the bee host including digestion, detoxification, and pathogen defense (reviewed in [6, 55]). We also found multiple flower-associated bacteria including Neokomagataea, Acinetobacter, Asaia, and Pantoea in both host species [14, 19, 49]. This is interesting given that we did not directly feed floral microbes to the bumble bees. They may have acquired floral microbes through their wild-caught queen and/or the non-sterilized honey bee-collected pollen they were fed prior to the experiment. Colony had a significant effect on community composition in bumble bees, and one bacterial genus, Bombella, which is an acetic acid bacteria with antifungal effects [56], only occurred in workers from one colony. This suggests that the first microbes that arrive in the gut, likely originating from the queen during spring foraging, become established and then remain stable in the face of varying nutritional input. This significant effect of colony on bumble bee gut communities contrasts with recent work showing that honey bees from different locations around the United States harbored similar gut bacteria down to the strain level [57], suggesting that environmental factors play a stronger role in shaping the gut microbiota of B. impatiens compared to A. mellifera. Megachile rotundata, which were dominated by nectar-inhabiting taxa, may have acquired floral microbes from the flower inoculum that we provided, or from larval pollen provisions (although bees shed their larval microbiome during metamorphosis), the nest cavity, or leaves lining the nest. This is consistent with the hypothesis that flowers are transmission hubs for pollinator microbes [19].
Captive rearing likely affects the bee gut microbiota. In our study, individuals from both species were provisioned by wild mothers, but were then exposed to a subset of the microbes they would normally encounter as adults since they had no direct contact with the natural environment. Although we did not compare our bees to wild-caught bees, captive rearing likely limits the diversity of the adult gut microbiome. This may affect the health of captive-reared bees, especially if bees are multiple generations removed from the wild or exposed to stressors, such as agrochemicals, that can further disrupt their native microbiota. There is new research on potential probiotics for honey bees (reviewed in [58]) that could be considered by bee rearing companies, although further studies are needed to assess their efficacy and in more bee species.
We predicted that a low-diversity pollen diet would result in reduced melanization in bumble bees, but our results did not support this prediction (Fig. 4). Although we only assessed a small portion of the bee immune system, our results are consistent with previous studies that found that monofloral and polyfloral pollen diets resulted in similar immune function in bumble bees (e.g., [59]). However, bumble bees with no access to pollen had reduced immune gene expression [60] and immunity was positively correlated with amount of pollen consumed [59]. This suggests that pollen is important for immune function in adult bees, but the type or diversity of pollen may be less so. However, in nature certain pollen types are correlated with certain microbes, and these microbes likely have important functions for training the bee immune system and/or inhibiting pathogens through other mechanisms [16].
Our results suggest that even single-species, low-protein pollen diets such as dandelion and sumac provided adequate nutrition to support diverse bacterial communities in adults of two distantly related bee species. However, older bumble bee workers may be more vulnerable to dietary quality than callows, since their gut microbiome was more sensitive to diet composition. This study is the first to assess the gut microbiome in a solitary bee species in an experimental setting and contributes to our understanding of the composition, variation, and drivers in solitary bee gut microbes. Our results further demonstrate that variation in bee gut microbiomes in the wild are not likely driven by pollen nutritional resources, and may be more strongly influenced by other factors such as the introduction of new microbes from flowers, nests, and social interactions. This study improves our understanding of wild bee gut microbiomes and biology, as well as drivers of bee health in the face of reduced biodiversity and emergent pathogens.
Data Availability
All data and code are available at https://github.com/aefowler/bee-microbiome-diet-diversity and on UMass ScholarWorks.
References
Cummings CR, Hernandez SM, Murray M, Ellison T, Adams HC, Cooper RE, Curry S, Navara KJ (2020) Effects of an anthropogenic diet on indicators of physiological challenge and immunity of white ibis nestlings raised in captivity. Ecol Evol 10(15):8416–8428
Turnbaugh PJ, Ridaura VK, Faith JJ, Rey FE, Gordon JI (2010) The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci Transl Med1:6ra14.
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE, Ling AV, Devlin AS, Varma Y, Fischbach MA, Biddinger SB (2014) Diet rapidly and reproducibly alters the human gut microbiome. Nature 505(7484):559–563
Potts SG, Biesmeijer JC, Kremen C, Neumann P, Schweiger O, Kunin WE (2010) Global pollinator declines: Trends, impacts and drivers. Trends Ecol Evol 25(6):345–353
Goulson D, Nicholls E, Botias C, Rotheray EL (2015) Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Sci 347(6229):1255957–1255957
Fowler AE, Irwin RE, Adler LS (2019) Parasite defense mechanisms in bees: behavior, immunity, antimicrobials, and symbionts. Emerg Top Life Sci 1–18
Danforth BN, Minckley RL, Neff JL (2019) The Solitary bees: biology, evolution, conservation. Princeton University Press, Princeton, New Jersey
Dance C, Botías C, Goulson D (2017) The combined effects of a monotonous diet and exposure to thiamethoxam on the performance of bumblebee micro-colonies. Ecotoxicol Environ Saf 139:194–201
Renauld M, Hutchinson A, Loeb G, Poveda K, Connelly H (2016) Landscape simplification constrains adult size in a native groundnesting bee. PloS ONE 11(3):e0150946
Centrella M, Russo L, Moreno Ramírez N, Eitzer B, van Dyke M, Danforth B, Poveda K (2020) Diet diversity and pesticide risk mediate the negative effects of land use change on solitary bee offspring production. J Appl Ecol 57(6):1031–1042
Giacomini JJ, Adler LS, Reading BJ, Irwin RE (2023) Differential bumble bee gene expression associated with pathogen infection and pollen diet. BMC genomics 24(1):1–8
Ricigliano VA, Williams ST, Oliver R (2022) Effects of different artificial diets on commercial honey bee colony performance, health biomarkers, and gut microbiota. BMC Vet Res 18(1):1–14
Billiet A, Meeus I, Van Nieuwerburgh F, Deforce D, Wäckers F, Smagghe (2016) Impact of sugar syrup and pollen diet on the bacterial diversity in the gut of indoor-reared bumblebees (Bombus terrestris). Apidologie 47(4):548–560
Figueroa LL, Maccaro JJ, Krichilsky E, Yanega D, McFrederick QS (2021) Why did the Bee eat the chicken? Symbiont Gain, loss, and Retention in the Vulture Bee Microbiome. MBio 12(6):e02317–21.
Kwong WK, Moran NA (2016) Gut microbial communities of social bees. Nat Rev Microbiol 14(6):374–384
Kwong WK, Mancenido AL, Moran NA (2017) Immune system stimulation by the native gut microbiota of honey bees. R Soc Open Sci 4(170003):1–9
Graystock P, Rehan SM, Mcfrederick QS (2017) Hunting for healthy microbiomes: determining the core microbiomes of Ceratina, Megalopta, and Apis bees and how they associate with microbes in bee collected pollen. Conserv Genet 18(3):701–711
Gu Y, Han W, Wang Y, Liang D, Gao J, Zhong Y, Zhao S, Wang S (2023) Xylocopa caerulea and Xylocopa auripennis harbor a homologous gut microbiome related to that of eusocial bees. Front Microbiol 14
McFrederick QS, Thomas JM, Neff JL, Vuong HQ, Russell KA, Hale AR, Mueller UG (2017) Flowers and wild megachilid bees share microbes. Microb Ecol 188–200
Dharampal PS, Carlson C, Currie CR, Steffan SA (1904) Pollen-borne microbes shape bee fitness. Proc R Soc B Biol Sci 286:20182894
Sookhan N, Lorenzo A, Tatsumi S, Yuen M, MacIvor JS (2020) Linking bacterial diversity to floral identity in the bumble bee pollen basket. Environ DNA no August 1–12
Voulgari-Kokota A, Ankenbrand MJ, Grimmer G, Steffan-Dewenter I, Keller A (2019) Linking pollen foraging of megachilid bees to their nest bacterial microbiota. Ecol Evol 9(18):10788–10800
Vanderplanck M, Gilles H, Nonclercq D, Duez P, Gerbaux P (2020) Asteraceae Paradox: Chemical and Mechanical Protection of Taraxacum Pollen. Insects 11(5):304
Strange JP, Tripodi AD, Huntzinger C, Knoblett J, Klinger E, Herndon JD, Vuong HQ, McFrederick QS, Irwin RE, Evans JD, Giacomini JJ (2023) Comparative analysis of 3 pollen sterilization methods for feeding bumble bees. J. Econ. Entomol 116:662–673
Zheng H, Powell JE, Steele MI, Dietrich C, Moran NA (2017) Honeybee gut microbiota promotes host weight gain via bacterial metabolism and hormonal signaling. Proc Natl Acad Sci 114(18)
Koch H, Schmid-Hempel P (2011) Socially transmitted gut microbiota protect bumble bees against an intestinal parasite. Proc Natl Acad Sci 108(48):19288–92
Hammer TJ, Easton-Calabria A, Moran NA (2023) Microbiome assembly and maintenance across the lifespan of bumble bee workers. Mol Ecol 32(3):724–740
Pinilla-Gallego MS, Irwin RE (2022) Effects of an alternative host on the prevalence and intensity of Infection of a bumble bee parasite. Parasitology 149(4):562–567
Pitts-Singer TL, Cane JH (2011) The Alfalfa Leafcutting Bee, Megachile rotundata: the World’s most intensively managed Solitary Bee. Annu Rev Entomol 56:221–237
McFrederick QS, Rehan SM (2016) Characterization of pollen and bacterial community composition in brood provisions of a small carpenter bee. Mol Ecol 25(10):2302–2311
Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37(8):852–857
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, Huttley GA, Gregory Caporaso J (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-featureclassifier plugin. Microbiome 6(1):1–17
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(D1):590–596
R Core Team (2022) A Language and Environment for Statistical Computing. R found stat comput. Vienna, Austria. https://www.R-project.org
Davis NM, Proctor DM, Holmes SP, Relman DA, Callahan BJ (2018) Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6(1):1–14
Oksanen J, Kindt R, Legendre P, O'Hara B, Simpson GL, Solymos PM, Stevens MH, Wagner H (2008) The vegan package, Community Ecol Packag 190
McMurdie PJ, Holmes S (2013) Phyloseq: an R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 8(4):e61217
Lenth R (2020) Emmeans: Estimated marginal means, aka least-squares means
Therneau TM (2020) Coxme: Mixed Effects Cox Models. R package version
Therneau T (2022) A package for survival analysis in R
Fox J, Weisberg S (2019) An R companion to applied regression, Third Edition. Sage, Thousand Oaks, CA
Mazerolle MJ (2020) AICcmodavg. R Vignette
Hartig F (2022) “_DHARMa: residual diagnostics for hierarchical (Multi-Level / Mixed) Regression Models_.” R package version 0.4.6
Wickham H (2016) Ggplot: elegant graphics for data analysis. Springer-Verlag, New York
Davis SE, Malfi RL, Roulston H (2015) Species differences in bumblebee immune response predict developmental success of a parasitoid fly. Oecologia 178:1017–1032
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to imageJ: 25 years of image analysis. Nat Methods 9(7):671–675
Brooks ME, Kristensen K, Van Benthem KJ, Magnusson A, Berg CW, Nielsen A, Skaug HJ, Machler M, Bolker BM (2017) GlmmTMB balances speed and flexibility among packages for zero-inflated generalized Linear mixed modeling. R J 9(2):378–400
Yukphan P, Malimas T, Muramatsu Y, Potacharoen W, Tanasupawat S, Nakagawa Y, Tanticharoen M, Yamada Y (2011) Neokomagataea gen. Nov., with descriptions of Neokomagataea thailandica sp. nov. and Neokomagataea tanensis sp. Nov. Biosci Biotechnol Biochem 75(3):419–426
Ambika Manirajan B, Ratering S, Rusch V, Schwiertz A, Geissler-Plaum R, Cardinale M, Schnell S (2016) Bacterial microbiota associated with flower pollen is influenced by pollination type, and shows a high degree of diversity and species-specificity. Environ Microbiol 18(12):5161–5174
Engel P, Martinson VG, Moran NA (2012) Functional diversity within the simple gut microbiota of the honey bee. Proc Natl Acad Sci 109(27):11002–11007
Voulgari-Kokota A, Grimmer G, Steffan-Dewenter I, Keller A (2018) Bacterial community structure and succession in nests of two megachilid bee genera. FEMS Microbiol Ecol 95(1):1–11
Kartzinel TR, Hsing JC, Musili PM, Brown BRP, Pringle RM (2019) Covariation of diet and gut microbiome in African megafauna. Proc Natl Acad Sci 116(47):23588–23593
Hammer TJ, Sanders JG, Fierer N (2019) Not all animals need a microbiome. FEMS Microbiol Lett 366:1–11
Hammer TJ, Le E, Martin AN, Moran NA (2021) The gut microbiota of bumblebees. Insectes Soc 0123456789
Miller DL, Smith EA, Newton ILG (2021) A bacterial symbiont protects honey bees from fungal Disease. MBio 12(3)
Bobay L, Wissel EF, Raymann K (2020) Strain structure and Dynamics revealed by targeted deep sequencing of the honey bee gut microbiome. mSphere 5:e00694–20
Motta EVS, Powell JE, Leonard SP, Moran NA (2022) Prospects for probiotics in social bees. Philos Trans R Soc Lond B Biol Sci 377:20210156
Fowler AE, Sadd BM, Bassingthwaite T, Irwin RE, Adler LS (2022) Consuming sunflower pollen reduced pathogen Infection but did not alter measures of immunity in bumblebees. Philos Trans R Soc B Biol Sci 377:1853
Brunner FS, Schmid-Hempel P, Barribeau SM (2014) Proteinpoor diet reduces host-specific immune gene expression in Bombus terrestris. Proc R Soc B Biol Sci 281(1786):20140128
Acknowledgements
We thank M. Argueta Guzman, M. Hilliard, M. Garcia Morales, and L. Aguirre for bioinformatics help; E. Kola, L. Ngor, and A. Zymaris for lab help; M. Tissier, Y. Lauranger, J. Koch, and T. Lindsay for supplying and sterilizing the pollen; S. Pinilla-Gallego for advice about leafcutter bees; B. Sadd for advice about bumble bee gut microbes; R. Wick and P. Pearson for equipment; and A. Porter and the Adler Lab for comments on the manuscript.
Funding
This project was funded by the Northeast Sustainable Agriculture Research and Education program graduate student grant GNE19-200-33243, the US Department of Agriculture AFRI-2018-08591, and the Graduate Research Fellowship from the National Science Foundation. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the funding agencies.
Author information
Authors and Affiliations
Contributions
AEF and LSA designed the study. AEF conducted the experiments in the lab of LSA and the dissections, DNA extractions, and library preparations in the lab of QSM. AEF analyzed the data with guidance from QSM and wrote the first draft of the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fowler, A.E., McFrederick, Q.S. & Adler, L.S. Pollen Diet Diversity does not Affect Gut Bacterial Communities or Melanization in a Social and Solitary Bee Species. Microb Ecol 87, 25 (2024). https://doi.org/10.1007/s00248-023-02323-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-023-02323-6