Abstract
In this paper, we report a molecular diagnostic system—combining a colorimetric probe (RHthio-CuSO4) for pyrophosphate sensing and isothermal gene amplification (ramified rolling circle amplification)—that operates with high selectivity and sensitivity for clinical point-of-care diagnosis of SARS-CoV-2. During the polymerase phase of the DNA amplification process, pyrophosphate was released from the nucleotide triphosphate as a side product, which was then sensed by our RHthio-CuSO4 probe with a visible color change. This simple colorimetric diagnostic system allowed highly sensitive (1.13 copies/reaction) detection of clinical SARS-CoV-2 within 1 h, while also displaying high selectivity, as evidenced by its discrimination of two respiratory viral genomes (human rhino virus and respiratory syncytial virus) from that of SARS-CoV-2. All of the reactions in this system were performed at a single temperature, with positive identification being made by the naked eye, without requiring any instrumentation. The high sensitivity and selectivity, short detection time (1 h), simple treatment (one-pot reaction), isothermal amplification, and colorimetric detection together satisfy the requirements for clinical point-of-care detection of SARS-CoV-2. Therefore, we believe that this combination of a colorimetric probe and isothermal amplification will be useful for point-of-care testing to prevent the propagation of COVID-19.
Similar content being viewed by others
Introduction
Molecular diagnostic methods can help to prevent the propagation of infectious diseases prior to the development of effective drugs and vaccines [1]. These diagnostic methods require selectivity and sensitivity when identifying such diseases [2, 3]. The reverse transcription–based polymerization chain reaction (RT-PCR) has become the standard confirmatory method for diagnosing viral infectious diseases, including coronavirus disease 2019 (COVID-19) [4]. Nevertheless, its adaptability and utility have been limited because of its long turnaround time and the need for skilled laboratory personnel and specialized equipment [5]. Commercial RT-PCR processes demand gel electrophoresis [6], an exonuclease with the quenched probe oligonucleotide [7, 8], SYBR Green (as a fluorogenic duplex DNA–sensing material) for detection of the amplification products [9], or biosensing mediated by nano- and electro-based materials [10,11,12,13]. Therefore, despite its high accuracy, efficiency, and sensitivity, many barriers remain for on-site diagnosis using RT-PCR, due to the expensive equipment and manpower required to handle this complex diagnostic process. Accordingly, there is room to develop much more efficient genetic molecular diagnostic methods for diagnosing the viral diseases that are likely to arise periodically in the future, and to prevent their rapid spread. Thus, novel methods for rapid detection are being sought, as well as point-of-care (POC) devices displaying sufficient sensitivity and selectivity to prevent the rapid spread of disease [14]. From the viewpoint of rapidity, genetic molecular diagnostic methods based on isothermal amplification would appear to be useful alternatives.
The most important characteristic of POC diagnosis is that the process should be simple (i.e., with no need for expensive equipment), such that it could be adopted widely in resource-limited settings. In this respect, although genetic diagnostic methods based on fluorescence have displayed high sensitivity, they are less attractive because they require fluorescence detectors. In contrast, colorimetric methods are more appealing for on-spot diagnosis, particularly when the color changes can be detected by the naked eye, without the need for any detection instrumentation.
Recently, several colorimetric probing systems, combined with RT-PCR, CRISPR/Cas9, and reverse transcription loop-mediated isothermal amplification (RT-LAMP), have been developed for POC testing to minimize detection times and the need for expensive equipment [15,16,17,18,19]. As a simple sensing platform that operates without any equipment, we recently developed a novel pyrophosphate-sensing molecule for colorimetric detection based on Cu2+ complexation and decomplexation [20, 21].
Our pyrophosphate probe provided a color signal after performing DNA polymerase–assisted gene amplification, because the DNA polymerase released pyrophosphate after incorporating the nucleoside triphosphates. Nevertheless, most enzyme-based biological reactions require a variety of organic and inorganic substances, which may become obstacles in selectively detecting only pyrophosphate.
For example, dithiothreitol (DTT) is a key material for maintaining enzymatic function by preventing the intramolecular and intermolecular disulfide bonds of proteins from forming. Many DNA polymerases include DTT in their buffer solutions. Unfortunately, our previously reported probe reacted with the thiol groups of DTT and cysteine, and could not discriminate them from pyrophosphate [21, 22]. To resolve this problem, in this present study, we prepared RHthio-CuSO4, a new probe (based on CuSO4 chelation) that displays improved resistance toward DTT, cysteine, and other biological substrates and allows naked-eye detection of pyrophosphate (Scheme 1).
Many types of isothermal amplification have already been applied in systems for COVID-19 detection, including loop-mediated isothermal amplification (LAMP) [23,24,25], recombinase polymerase amplification (RPA) [26, 27], and rolling circle amplification (RCA) [28, 29]. Here, to ensure rapid and ultra-sensitive detection, we combined our pyrophosphate colorimetric diagnosis technique with an RCA-based ramification method: ramified rolling circle amplification (RAM), a double isothermal amplification system [30]. To the best of our knowledge, we report herein the first example of its application to the detection of RNA from severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).
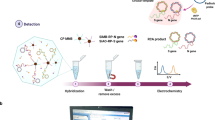
We designed a rapid and ultra-sensitive RAM reaction involving splint R Ligase, phi29 DNA polymerase, and three oligonucleotides and combined it with RHthio-CuSO4 for one-pot colorimetric sensing (Scheme 2). For an actual clinical application of this on-site diagnostic method, we extracted human materials from the nasopharynx of an actual COVID-19–infected patient and found that diagnosis was possible with very high sensitivity and selectivity. Accordingly, we believe that this method will be useful for POC diagnosis and for prevention of the propagation of a diverse range of viral diseases, including COVID-19.
Experiments
General information
All DNA oligonucleotides (RAM tem, RAM pri-1, RAM pri-2) were purchased from Bioneer (South Korea). The dNTP mixture (dATP, dTTP, dCTP, dGTP; each 2 mM) was purchased from Enzynomics (South Korea). The SARS-CoV-2 full genome was isolated using an AccuPlex™ SARS-CoV-2 Reference Material Kit from Seracare (USA). Splint R ligase and phi29 DNA polymerase were purchased from New England Biolabs (USA). RHthio and RHthio-CuSO4 were synthesized in-house; data are provided in the Supporting Information. Chemicals used to synthesize RHthio were purchased from Sigma–Aldrich (USA). UV–Vis absorption spectra were recorded using a Shimadzu (Japan) UV-1650PC spectrophotometer. All optical measurements were performed at room temperature, using a quartz cuvette (path length: 1 cm). Agarose gel electrophoresis was performed in 1% agarose gel. The gel was loaded in an electrophoresis instrument at 100 V for 25 min. The gels were stained in EtBr solution for 15 min; the stained gels were washed in water for 30 min. Photographs of the gels were recorded using an E-gel Imager (Invitrogen, USA); colorimetric detection images were captured using a mobile device.
RHthio-CuSO4 selectivity study with anions, DTT, and cysteine
Solutions of the anions and biomaterials were prepared at 200 mM in water. Solutions of RHthio and RHthio-CuSO4 were prepared at 25 mM in DMSO. The dilution buffer was prepared using water and acetonitrile in a 7:3 ratio. In the dilution buffer, 25 mM RHthio-CuSO4 (1 μL) and a 200 mM solution of a reactive material (1 μL) were added in a 1.5-mL tube. All tubes were shaken for 1 min and then transferred to a 1-cm cuvette cell to measure the absorbance spectra. All spectral absorbances were recorded at room temperature.
RAM reaction confirmation
The RAM reactions were performed in solutions having a total volume of 20 μL. The RAM oligo mixture included 1 μM of RAM tem, 10 μM of RAM pri-1, and 1 μM of RAM pri-2. The RAM reaction mixture was treated with 10 × phi29 DNA polymerase buffer (500 mM Tris–HCl, 100 mM MgCl2, 100 mM (NH4)2SO4, 10 mM DTT; 2 μL), 10 × Splint R ligase buffer (500 mM Tris–HCl, 100 mM MgCl2, 10 mM ATP; 2 μL), 10 × bovine serum albumin (BSA, 2 mg/mL; 2 μL), the dNTP mixture (2 mM of dATP, dCTP, dGTP, and dTTP; 6 μL), and RAM oligo mixture (2 μL) to give a total volume of 14 μL in one 1.5-mL tube. Prior to starting the reaction, the target (5 μL) was added, followed by Splint R ligase (0.5 μL) and phi29 DNA polymerase (10 U/μL; 0.5 μL). The reaction mixture was incubated for 1 h at 37 °C.
All RAM reactions were performed according to the RAM protocol. The target featured 20 copies/μL of SARS-CoV-2 full-genome material. To form the negative controls, one component (phi29 DNA polymerase, Splint R ligase, target SARS-CoV-2, RAM tem, RAM pri-1, or RAM pri-2) was neglected in each case. All other reaction conditions were the same as the general protocol. The reactions were confirmed using agarose gel electrophoresis (1%), with EtBr staining.
Sensitivity and selectivity measurements
An AccuPlex™ SARS-CoV-2 Reference Material Kit (Seracare, Milford, MA, USA), which was assigned as 5000 copies/mL, was used for spiked samples. The SARS-CoV-2 RNA was extracted using an eMAG system (bioMerieux, MarcylEtoile, France), following the extraction protocol provided by the manufacturer, with an input volume of 200 μL and an elution volume of 50 μL. The copy concentration in the extracted RNA was approximately 20 copies/μL. To increase the concentration, the SARS-CoV-2 RNA was lyophilized to give 50 copies/μL. For the sensitivity study, the RNA sample was diluted in distilled water to give concentrations varying from 0.1 to 50 copies/μL. All of the samples at each concentration were subjected to the RAM reaction with RHthio-CuSO4; the absorbances were measured to calculate the LOD. For the selectivity study, nine species of bacteria that are known normal flora in the upper respiratory tract (Staphylococcus aureus, Staphylococcus epidermidis, Enterococcus faecalis, Enterococcus feacium, Escherichia coli, Klebsiella pneumoniae, Enterobacter cloacae, Pseudomonas aeruginosa, and Acinetobacter baumannii) and two respiratory RNA viruses (human rhinovirus and respiratory syncytial virus) were obtained from clinical samples. All bacterial DNA was extracted using the boiling method, with DNA extraction buffer (Seegene, Seoul, South Korea); viral RNA was extracted in the same manner of validation with clinical samples. The extracted bacterial DNA and viral RNA were subjected to the RAM reaction with RHthio-CuSO4; their analytical results were compared with those for the SARS-CoV-2 detection. In the sensitivity and selectivity studies, water (180 μL) was added into a 250-μL reaction tube along with 25 mM RHthio-CuSO4 in DMSO (2 μL). For detailed analysis, the absorbance of each reaction mixture was measured in the presence of RHthio-CuSO4.
Clinical sample preparation and validation
The evaluation of clinical samples was approved by the Institutional Review Board of Jeonbuk National University Hospital (IRB approval number: CUH-2021–11-005). The requirement for patient consent was waived by the IRB according to local laws. A total of 40 residual samples for SARS-CoV-2 real-time reverse transcription PCR (rRT-PCR) were employed in this study: 20 positive samples and 20 negative samples. All nasopharyngeal samples were collected according to the standard protocol for the diagnosis of COVID-19 [4]. All swabs were sampled using an eNAT tube (Copan Italy, Brescia, Italy). After performing the clinical tests for the diagnosis of COVID-19, all samples were stored at − 20 °C. The nucleic acid was extracted using Magna Pure 24 (Roche Diagnostics, Basel, Switzerland) or eMAG (bioMérieux, Marcy-l’Étoile, France), following the manufacturer’s protocol. The rRT-PCRs were performed using the Allplex SARS-CoV-2 Assay (Seegene, Seoul, South Korea); interpretations were performed according to the manufacturer’s instructions and the Guidelines for Laboratory Diagnosis of Coronavirus Disease 2019 in Korea [4]. Samples having positive cycle threshold (Ct) values from 19.00 to 37.86 for the RdRp gene were selected. For clinical validation, 40 reactions were performed using the RAM reaction combined with RHthio-CuSO4 detection. For all clinical validation studies, water (180 μL) was added into a 250-μL reaction tube, followed by 25 mM RHthio-CuSO4 in DMSO (2 μL). For detailed analysis, the absorbance of each reaction with RHthio-CuSO4 was measured.
Results and discussion
Optimization
The RHthio-CuSO4 complex was prepared in two steps (Scheme S1). Rhodamine hydrazide was condensed with thiophene 2-carboxaldehyde in the presence of a catalytic amount of trifluoroacetic acid (TFA) in MeOH to give the hydrazone RHthio, which was then complexed with CuSO4 in DMF to give a deep-violet solid. This complex was characterized using IR spectroscopy and high-resolution mass spectrometry (HRMS); it was dissolved in DMSO (25 mM) for use in subsequent studies.
First, we examined the selectivity of RHthio-CuSO4 toward pyrophosphate (Fig. 1a), testing the effects of various anions that would be present in the biological reaction. In the presence of pyrophosphate, the absorption of RHthio-CuSO4 at 550 nm (λmax) underwent a clear decrease in intensity, implying the exchange of its Cu2+ cation to pyrophosphate; the other anions did not induce any dramatic changes in absorption. Furthermore, we did not observe any dramatic decreases in absorption when DTT (a key material for enzyme function) and cysteine were present, although they did induce some small changes in the absorption spectra. Although the exact mechanism is unknown, we suspect that the thiophene unit in RHthio-CuSO4 influenced the strength of the coordinative bonds of the imino and C = O groups toward the Cu2+ cation.
a Selectivity of RHthio-CuSO4. Absorbance spectra of RHthio and RHthio-CuSO4 recorded in the presence of various anions and biomaterials. All reaction mixtures included 25 mM RHthio or RHthio-CuSO4 (1 μL) and a reactive material (200 mM, 1 μL) in water (1 mL). b Agarose gel electrophoresis of the RAM reaction. Lane M: DNA marker; lane 1: RAM reaction without phi29 DNA polymerase; lane 2: RAM reaction without Splint R ligase; lane 3: RAM reaction without SARS-CoV-2 target; lane 4: RAM reaction without RAM template; lane 5: RAM reaction without RAM pri-1; lane 6: RAM reaction without pri-2; lane 7: RAM reaction. The gel was stained with EtBr
Next, we examined the RAM process targeting SARS-CoV-2. Here, we prepared a circular template sequence (RAM tem) and pri-1 (RAM pri-1) and pri-2 (RAM pri-2) sequences for the RAM reaction, and screened different target regions in SARS-CoV-2. Designing a primer that binds to a specific region of viral RNA for reverse transcription is the key pivotal step for selective detection of SARS-CoV-2 when using RT-PCR, RT-LAMP, and RT-RPA for amplification. Accordingly, we screened five different target binding regions: RdRp gene (Table S1), RdRp-2 gene, E gene, S gene, and N gene (Table S2). We attempted to find the selective binding region by using 20 copies of the SARS-CoV-2 genome and analyzed the outcome through agarose gel electrophoresis. Figure S11 reveals that the E gene and S gene target regions displayed low sensitivity relative to other three targets (RdRp gene, RdRp-2 gene, and N gene). Thus, we excluded the E gene and S gene as target binding regions. To further identify the most selective region, we prepared nine species of bacteria that are known as normal flora in the upper respiratory tract (Staphylococcus aureus, Staphylococcus epidermidis, Enterococcus faecalis, Enterococcus feacium, Escherichia coli, Klebsiella pneumoniae, Enterobacter cloacae, Pseudomonas aeruginosa, and Acinetobacter baumannii) and two respiratory RNA viruses (human rhinovirus and respiratory syncytial virus) from clinical samples. Interestingly, our colorimetric RAM system could discriminate SARS-CoV-2 selectively from the bacterial genomes in all three target regions (RdRp gene, RdRp-2 gene, and N gene; Fig. S12). In the presence of the respiratory viruses, however, our colorimetric RAM system could identify SARS-CoV-2 selectively only in the RdRp gene region; that is, it could not discriminate the viral genome in the RdRp-2 gene and N gene regions (Fig. S13). Thus, we chose the RdRp gene as the target region for subsequent diagnoses of SARS-CoV-2.
To examine the success of the RAM reaction, we performed it under several negative conditions: in the absence of an enzyme, target, primer, or template, as negative controls (Fig. 1b). We used agarose gel electrophoresis to compare the results with those from the positive RAM reaction. Lanes 1 to 5 (see Fig. 1b for the components in each lane of the gel) did not feature any amplification bands, but lane 6 (lacking RAM pri-2) and lane 7 (containing RAM pri-2) revealed the amplification pattern. The reason why lane 6 contained amplification bands is because Splint R ligase successfully produced the circular DNA template, which was amplified with RAM pri-1 through the general RCA reaction. Compared with lane 7, however, the band in lane 6 was weaker. Thus, by adding RAM pri-2, an additional amplification step—ramified amplification—occurred during the RCA reaction. Accordingly, we confirmed that the RAM reaction was sensitive and faster than the RCA reaction.
Sensitivity and selectivity
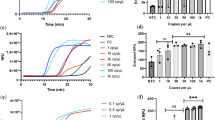
Next, we examined the sensitivity and selectivity of the detection when using our RHthio-CuSO4 probe and RAM system (Fig. 2). To ensure similar conditions for calculation of the detection limit, we used a full-genome SARS-CoV-2 materials kit. We extracted the RNA from the materials with assigned copy numbers, and prepared samples of various concentrations for the sensitivity study. First, we performed all of the RAM reactions using SARS-CoV-2 RNA; subsequently, we added the RHthio-CuSO4 probe. We measured the sensitivity in terms of the absorbance at 560 nm. Figure 2a presents the average change in absorbance at each specific concentration; the inset reveals that linear relationship used to calculate the limit of detection (LOD) through the 3σ method (LOD = 3 × (SD/S), where SD is the standard deviation and S is the slope of the plot). We obtained an LOD of 1.13 copies/rxn. Thus, our RHthio-CuSO4/RAM system was extremely sensitive, functioning even in the presence of only a small copy number of viral RNA from SARS-CoV-2. In general, RNA viruses have very low concentrations of their genomes; therefore, ultra-sensitive detection is necessary. In our system, 1.13 copies/μL was sufficient for detection of the RNA. Corman reported 2.31 log copies/mL based on a 25-μL reaction volume [31], and Pfefferle reported 2.83 log copies/mL as the LOD for SARS-CoV-2 when using real-time RT-PCR [32]. Thus, our RHthio-CuSO4/RAM system is much more sensitive than those used previously with other types of isothermal amplification [26,27,28,29].
Sensitivity and selectivity of the RAM reaction performed with the RHthio-CuSO4 probe. a Sensitivity study, using full-genome SARS-CoV-2 at concentrations from 0.5 to 250 copies/reaction (copies/rxn). All reactions were repeated three times; error bars are presented in the graph. Inset: linear relationship between the change in absorbance and the concentration of SARS-CoV-2 from 0.5 to 12.5 copies/rxn. All absorbances were measured at 560 nm. A, absorbance in the presence of target; A0, absorbance in the absence of target; y-axis, absolute value. b Selectivity study, performed in the presence of bacterial genomes, determined in terms of the absorbance at 560 nm. All reactions were repeated three times; error bars are presented in the chart. The tubes in the color photograph of the samples correspond to the bars below, revealing the relationship between color and absorbance
For the selectivity study, we employed several bacterial genomes to examine whether cross-reactions occurred with our amplification system (Fig. 2b). We performed the RAM reactions with nine types of bacterial genomes that are found in the nose and mouth, notably in the mucous membrane [33,34,35]. Because COVID-19 samples are typically taken from the mouth and nose, such bacteria would be present if there were any bacterial infection. Any diagnostic system that reacted with these bacteria would diagnose the patient as a false-positive; therefore, selective detection would be necessary for practical application. Figure 2b reveals that these bacteria did not react with the RAM system; there were no changes in absorbance because no amplification occurred and, therefore, the system could not produce pyrophosphate. The colorimetric detection was consistent with the absorbance change. We also examined the effects of two other respiratory viruses: human rhinovirus and respiratory syncytial virus. Again, our RHthio-CuSO4/RAM system could diagnose and discriminate the SARS-CoV-2 virus from these other respiratory viruses.
Clinical validation
Finally, we studied the suitability of our RHthio-CuSO4/RAM system for clinical application by testing clinical samples. We prepared 40 samples—20 positive (P01 to P20) and 20 negative (N01 to N20)—obtained from patients. When we applied our diagnostic system to the real samples, the positive test samples examined with our RHthio-CuSO4/RAM system revealed a dramatic color change in all 20 cases, even though there were variations in their copy numbers (Ct values from 19 to 37); although 18 of the negative samples displayed a violet color, two of the negative samples provided a positive response (Fig. 3a). Thus, we could clearly validate the positive samples and negative samples in terms of their color. The absorbance data were consistent with the results from the color images (Fig. 3b). As a result, our diagnostic method functioned with 100% sensitivity and 90% selectivity in the 40 tested samples. A good correlation also existed between the observed absorption values from the diagnoses performed with our system and the Ct values of the RdRp gene determined through real-time RT-PCR in Fig. S14 (Pearson’s r = 0.90). Our RHthio-CuSO4/RAM system provided a clear positive value even for samples with low viral titer—a Ct value of approximately 37.8 (positive cut-off Ct value for Allplex SARS-CoV-2 Assay: 40; maximum amplification: 45 cycles).
Clinical validation of the RAM system combined with RHthio-CuSO4. The color photographs and absorbance measurements were both performed using 40 clinical samples (20 positive and 20 negative). a Colorimetric detection assays. Positive RHthio reactions appeared as colorless; negative RHthio reactions provided a violet color. b Absorbance diagnosis at 560 nm, presented as a bar chart. Positive samples appear as red bars; negative samples appear as blue bars. RT-PCR Ct values of the positive samples are presented in parentheses next to their sample numbers. Black dotted line represents the negative line
Our RHthio-CuSO4/RAM system displayed ultra-sensitivity and sufficient selectivity for POC detection purposes within 1 h, even for real clinical samples, when applying only two enzymes and three oligonucleotides, at only a single temperature, with detection by the naked eye. Nevertheless, we cannot say that this system is perfect for diagnosis, because its specificity was not 100%, and because we could not perform the validation with a sufficient number of samples. For clinical use in the real world, a well-designed clinical validation study would be necessary.
Conclusion
We have developed RHthio-CuSO4, a new colorimetric probe for the sensing of pyrophosphate with sufficient selectivity over other types of anions, as well as DTT and cysteine. We applied RHthio-CuSO4 with a RAM reaction for the diagnosis of SARS-CoV-2—particularly targeting the RdRp gene region. Gratifyingly, the RHthio-CuSO4/RAM system exhibited extremely high sensitivity for the detection of SARS-CoV-2 (1.13 copies/rxn). Our RHthio-CuSO4/RAM system also displayed high selectivity toward SARS-CoV-2 against various bacterial genomes and other human respiratory viruses. Finally, we validated the performance of RHthio-CuSO4/RAM using real clinical samples from COVID-19 patients. Our RHthio-CuSO4/RAM system displayed 100% sensitivity and 90% selectivity when diagnosing the real COVID-19 clinical samples. This RHthio-CuSO4/RAM system has several attractive features making it suitable for application in POC tests: high sensitivity and selectivity, a processing time within 1 h in total, simple treatment (one-pot reaction), isothermal amplification, and colorimetric detection by the naked eye. Furthermore, we suspect that this ultra-sensitive, selective, and colorimetric detection method will also be applicable to the POC diagnoses of a diverse range of other RNA-based diseases.
References
Caly L, Druce J, Roberts J, Bond K, Tran T, Kostecki R, Yoga Y, Naughton W, Taiaroa G, Seemann T, Schultz MB, Howden BP, Korman TM, Lewin SR, Williamson DA, Catton MG. Isolation and rapid sharing of the 2019 novel coronavirus (SARS-CoV-2) from the first patient diagnosed with COVID-19 in Australia. Med J Aust. 2020;212:459–62.
Wu AH, McKay C, Broussard LA, Hoffman RS, Kwong TC, Moyer TP, Otten EM, Welch SL, Wax P. National academy of clinical biochemistry laboratory medicine practice guidelines: recommendations for the use of laboratory tests to support poisoned patients who present to the emergency department. Clin Chem. 2003;49:357–79.
Giljohann DA, Mirkin CA. Drivers of biodiagnostic development. Nature. 2009;462:461–4.
Hong KH, Lee SW, Kim TS, Huh HJ, Lee J, Kim SY, Pack JS, Kim GJ, Sung H, Roh KH, Kim JS, Kim HS, Lee ST, Seong MW, Ryoo N, Lee H, Kwon KC, Yoo CK. Guidelines for laboratory diagnosis of coronavirus disease 2019 (COVID-19) in Korea. Ann Lab Med. 2020;40:351–60.
Vandenberg O, Martiny D, Rochas O, van Belkum A, Kozlakidis Z. Considerations for diagnostic COVID-19 tests. Nat Rev Microbiol. 2021;19:171–83.
Poddar SK. Influenza virus types and subtypes detection by single step single tube multiplex reverse transcription-polymerase chain reaction (RT-PCR) and agarose gel electrophoresis. J Virol Methods. 2002;99:63–70.
Perchetti GA, Nalla AK, Huang ML, Jerome KR, Greninger AL. Multiplexing primer/probe sets for detection of SARS-CoV-2 by qRT-PCR. J Clin Virol. 2020;129: 104499.
Behrmann O, Bachmann I, Spiegel M, Schramm M, Abd El Wahed A, Dobler G, Dame G, Hufert FT. Rapid detection of SARS-CoV-2 by low volume real-time single tube reverse transcription recombinase polymerase amplification using an exo probe with an internally linked quencher (exo-IQ). Clin Chem. 2020;66:1047–1054.
Marinowic DR, Zanirati G, Rodrigues FVF, Grahl MVC, Alcará AM, Machado DC, Da Costa JC. A new SYBR Green real-time PCR to detect SARS-CoV-2. Sci Rep. 2021;11:2224.
El-Sherif DM, Abouzid M, Gaballah MS, Ahmed AA, Adeel M, El-Sheikh SM. New approach in SARS-CoV-2 surveillance using biosensor technology: a review. Environ Sci Pollut Res. 2022;29:1677–95.
Qiu A, Gai Z, Tao Y, Schmitt J, Kullak-Ublick GA, Wang J. Dual-functional plasmonic photothermal biosensors for highly accurate severe acute respiratory syndrome coronavirus 2 detection. ACS Nano. 2020;14:5268–77.
El-Sheikh SM, Osman DI, Ali OI, Shousha WG, Shoeib MA, Shawky SM, Sheta SM. A novel Ag/Zn bimetallic MOF as a superior sensitive biosensing platform for HCV-RNA electrochemical detection. Appl Surf Sci. 2021;562: 150202.
Raibiee N, Bagherzadeh M, Ghasemi A, Zare H, Ahmadi S, Fatahi Y, Dinarvand R, Rabiee M, Ramakrishna S, Shokouhimehr M, Varma RS. Point-of-use rapid detection of SARS-CoV-2: nanotechnology-enabled solutions for the COVID-19 pandemic. Int J Mol Sci. 2020;21:5126.
Nayak S, Blumenfeld NR, Laksanasopin T, Sia SK. Point-of-care diagnostics: recent developments in a connected age. Anal Chem. 2017;89:102–23.
Fakhri N, Abarghoei S, Dadmehr M, Hosseini M, Sabahi H, Ganjali MR. Paper based colorimetric detection of miRNA-21 using Ag/Pt nanoclusters. Spectrochim Acta A Mol Biomol Spectrosc. 2020;227: 117529.
Moon J, Kwon HJ, Yong D, Lee IC, Kim H, Kang H, Lim EK, Lee KS, Jung J, Park HG, Kang T. Colorimetric detection of SARS-CoV-2 and drug-resistant pH1N1 using CRISPR/dCas9. ACS Sens. 2020;5:4017–26.
Karami A, Hasani M, Jalilian FA, Ezati R. Conventional PCR assisted single-component assembly of spherical nucleic acids for simple colorimetric detection of SARS-CoV-2. Sens Actuators B Chem. 2021;328: 128971.
Choi MH, Lee J, Seo YJ. Dual-site ligation-assisted loop-mediated isothermal amplification (dLig-LAMP) for colorimetric and point-of-care determination of real SARS-CoV-2. Microchim Acta. 2022;189:176.
Kozlowski HN, Abdou Mohamed MA, Kim J, Bell NG, Zagorovsky K, Mubareka S, Chan WC. A colorimetric test to differentiate patients infected with influenza from COVID-19. Small Structures. 2021;2:2100034.
Pandith A, Bhattarai KR, Siddappa RKG, Chae HJ, Seo YJ. Novel fluorescent C2-symmetric sequential on-off-on switch for Cu2+ and pyrophosphate and its application in monitoring of endogenous alkaline phosphatase activity. Sens Actuators B Chem. 2019;282:730–42.
Pandith A, Seo YJ. Label-free sensing platform for miRNA-146a based on chromo-fluorogenic pyrophosphate recognition. J Inorg Biochem. 2020;203: 110867.
Choi MH, Seo YJ. Rapid and highly sensitive hairpin structure-mediated colorimetric detection of miRNA. Anal Chim Acta. 2021;1176: 338765.
Broughton JP, Deng X, Yu G, Fasching CL, Servellita V, Singh J, Miao X, Streithorst JA, Granados A, Sotomayor-Gonzalez A, Zorn K, Gopez A, Hsu E, Gu W, Miller S, Pan CY, Guevara H, Wadford DA, Chen JS, Chiu CY. CRISPR–Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020;38:870–4.
Yu L, Wu S, Hao X, Dong X, Mao L, Pelechano V, Chen WH, Yin X. Rapid detection of COVID-19 coronavirus using a reverse transcriptional loop-mediated isothermal amplification (RT-LAMP) diagnostic platform. Clin Chem. 2020;66:975–7.
Klein S, Müller TG, Khalid D, Sonntag-Buck V, Heuser AM, Glass B, Meurer M, Molales I, Schillak A, Freistaedter A, Ambiel I, Winter SL, Zimmermann L, Naumoska T, Bubeck F, Kirrmaier D, Ullrich S, Miranda IB, Anders S, Grimm D, Schnitzler P, Knop M, Krausslich HG, Thi VLD, Borner K, Chlanda P. SARS-CoV-2 RNA extraction using magnetic beads for rapid large-scale testing by RT-qPCR and RT-LAMP. Viruses. 2020;12:863.
Patchsung M, Jantarug K, Pattama A, Aphicho K, Suraritdechachai S, Meesawat P, Sappakhaw K, Leelahakorn N, Ruenkam T, Wongsatit T, Athipanyasilp N, Eiamthong B, Lakkanasirorat B, Phoodokmai T, Niljianskul N, Pakotiprapha D, Chanarat S, Homchan A, Tinikul R, Kamutira P, Phiwkaow K, Soithongcharoen S, Kantiwiriyawanitch C, Pongsupasa V, Trisrivirat D, Jaroensuk J, Wongnate T, Maenpuen S, Chaiyen P, Kamnerdnakta S, Swangsri J, Chuthapisith S, Sirivatanauksorn Y, Chaimayo C, Sutthent R, Kantakamalakul W, Joung J, Ladha A, Jin X, Gootenberg JS, Abudayyeh OO, Zhang F, Horthongkham N, Uttamapinant C. Clinical validation of a Cas13-based assay for the detection of SARS-CoV-2 RNA. Nat Biomed Eng. 2020;4:1140–9.
Choi MH, Lee J, Seo YJ. Combined recombinase polymerase amplification/rkDNA–graphene oxide probing system for detection of SARS-CoV-2. Anal Chim Acta. 2021;1158: 338390.
Chaibun T, Puenpa J, Ngamdee T, Boonapatcharoen N, Athamanolap P, O’Mullane AP, Vongpunsawad S, Poovorawan Y, Lee SY, Lertanantawong B. Rapid electrochemical detection of coronavirus SARS-CoV-2. Nat Commun. 2021;12:802.
Jiao J, Duan C, Xue L, Liu Y, Sun W, Xiang Y. DNA nanoscaffold-based SARS-CoV-2 detection for COVID-19 diagnosis. Biosens Bioelectron. 2020;167: 112479.
Yao B, Li J, Huang H, Sun C, Wang Z, Fan Y, Chang Q, Li S, Xi J. Quantitative analysis of zeptomole microRNAs based on isothermal ramification amplification. RNA. 2009;15:1787–94.
Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DK, Bleicker T, Brunink S, Schneider J, Schmidt ML, Mulders DG, Haagmans BL, van der Veer B, van der Brink S, Wijsman L, Goderski G, Romette JL, Ellis J, Zambon M, Peiris M, Goossens H, Reusken C, Koopmans MP, Drosten C. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance. 2020;25:2000045.
Pfefferle S, Reucher S, Nörz D, Lütgehetmann M. Evaluation of a quantitative RT-PCR assay for the detection of the emerging coronavirus SARS-CoV-2 using a high throughput system. Eurosurveillance. 2020;25:2000152.
Dahlén G. Bacterial infections of the oral mucosa. Periodontol. 2000;2009(49):13–38.
Lai CC, Chen SY, Ko WC, Hsueh PR. Increased antimicrobial resistance during the COVID-19 pandemic. Int J Antimicrob Agents. 2021;57: 106324.
Song W, Jia X, Zhang X, Ling Y, Yi Z. Co-infection in COVID-19, a cohort study. J Infec. 2021;82:414–51.
Funding
This study was supported by the Basic Science Research Program through the National Research Foundation of Korea (2021R1A2C1003804), funded by the Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
The evaluation of clinical samples was approved by the Institutional Review Board of Jeonbuk National University Hospital (IRB approval number: CUH-2021–11-005). The requirement for patient consent was waived by the IRB according to local laws.
Conflict of interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
216_2022_4156_MOESM1_ESM.pdf
Supplementary file1 Synthetic scheme and method for synthesizing RHthio; characterization of RHthio; oligonucleotide sequence information; UV–Vis absorption spectra; target sequence selection. (PDF 1338 KB)
Rights and permissions
About this article
Cite this article
Choi, M.H., Kumara, G.S.R., Lee, J. et al. Point-of-care COVID-19 testing: colorimetric diagnosis using rapid and ultra-sensitive ramified rolling circle amplification. Anal Bioanal Chem 414, 5907–5915 (2022). https://doi.org/10.1007/s00216-022-04156-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-022-04156-7