Abstract
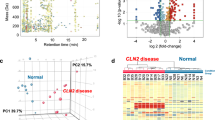
Mass spectrometry imaging (MSI) is a powerful tool to perform untargeted mapping of biomolecules in situ. In the current study, we performed matrix-assisted laser desorption/ionization-mass spectrometry imaging (MALDI-MSI) to evaluate lipid changes during disease progression (asymptomatic to symptomatic time points) in Niemann-Pick disease, type C1 (NPC1), a cerebellar neurodegenerative, lipid storage disorder. Our data show that gangliosides GM2 and GM3 are elevated in NPC1 disease and localize in the posterior lobules of the cerebellum, which is enhanced over a time-course analysis of the disease. Further analysis of sphingolipids in negative ion mode indicated reduction of sulfatides in white matter of the cerebellum and patterned distribution and co-localization of ceramide species Cer(d36:1), HexCer(d36:1), and the ganglioside GM1(d36:1) during disease progression. Finally, a putative lipid of unknown structure demonstrated similar patterning during NPC1 cerebellar degeneration. These studies provide insight into lipid markers of neurodegeneration in NPC1 and link lipid alterations to altered pathways that lead to cell death.




Similar content being viewed by others
References
Higgins JJ, Patterson MC, Dambrosia JM, Pikus AT, Pentchev PG, Sato S, et al. A clinical staging classification for type C Niemann-Pick disease. Neurology. 1992;42(12):2286–90. https://doi.org/10.1212/wnl.42.12.2286.
Patterson MC, Vanier MT, Suzuki K, Morris JA, Carstea E, Neufeld EB, et al. Niemann-Pick disease type C: a lipid trafficking disorder. In: Scriver CR, Beaudet AL, Sly WS, et al., editors. The metabolic and molecular bases of inherited disease. New York: Mc Graw Hill; 2001.
Vanier MT. Niemann-Pick disease type C. Orphanet J Rare Dis. 2010;5(1):16. https://doi.org/10.1186/1750-1172-5-16.
Naureckiene S, Sleat DE, Lackland H, Fensom A, Vanier MT, Wattiaux R, et al. Identification of HE1 as the second gene of Niemann-Pick C disease. Science. 2000;290(5500):2298–301. https://doi.org/10.1126/science.290.5500.2298.
Kwon HJ, Abi-Mosleh L, Wang ML, Deisenhofer J, Goldstein JL, Brown MS, et al. Structure of N-terminal domain of NPC1 reveals distinct subdomains for binding and transfer of cholesterol. Cell. 2009;137(7):1213–24. https://doi.org/10.1016/j.cell.2009.03.049.
Vanier MT. Biochemical studies in Niemann-Pick disease. I. Major sphingolipids of liver and spleen. Biochim Biophys Acta. 1983;750(1):178–84.
Zhou S, Davidson C, McGlynn R, Stephney G, Dobrenis K, Vanier MT, et al. Endosomal/lysosomal processing of gangliosides affects neuronal cholesterol sequestration in Niemann-Pick disease type C. Am J Pathol. 2011;179(2):890–902. https://doi.org/10.1016/j.ajpath.2011.04.017.
Vanier MT. Lipid changes in Niemann-Pick disease type C brain: personal experience and review of the literature. Neurochem Res. 1999;24(4):481–9.
Zervas M, Somers KL, Thrall MA, Walkley SU. Critical role for glycosphingolipids in Niemann-Pick disease type C. Curr Biol. 2001;11(16):1283–7.
Higashi Y, Murayama S, Pentchev PG, Suzuki K. Cerebellar degeneration in the Niemann-Pick type C mouse. Acta Neuropathol. 1993;85(2):175–84. https://doi.org/10.1007/BF00227765.
Xie C, Burns DK, Turley SD, Dietschy JM. Cholesterol is sequestered in the brains of mice with Niemann-Pick type C disease but turnover is increased. J Neuropathol Exp Neurol. 2000;59(12):1106–17. https://doi.org/10.1093/jnen/59.12.1106.
Xie C, Turley SD, Dietschy JM. Cholesterol accumulation in tissues of the Niemann-Pick type C mouse is determined by the rate of lipoprotein-cholesterol uptake through the coated-pit pathway in each organ. Proc Natl Acad Sci U S A. 1999;96(21):11992–7. https://doi.org/10.1073/pnas.96.21.11992.
Love S, Bridges LR, Case CP. Neurofibrillary tangles in Niemann-Pick disease type C. Brain. 1995;118 ( Pt 1:119–29. https://doi.org/10.1093/brain/118.1.119.
Kodam A, Maulik M, Peake K, Amritraj A, Vetrivel KS, Thinakaran G, et al. Altered levels and distribution of amyloid precursor protein and its processing enzymes in Niemann-Pick type C1-deficient mouse brains. Glia. 2010;58(11):1267–81. https://doi.org/10.1002/glia.21001.
Yan X, Yang F, Lukas J, Witt M, Wree A, Rolfs A, et al. Hyperactive glial cells contribute to axonal pathologies in the spinal cord of Npc1 mutant mice. Glia. 2014;62(7):1024–40. https://doi.org/10.1002/glia.22659.
Pentchev PG, Gal AE, Booth AD, Omodeo-Sale F, Fouks J, Neumeyer BA, et al. A lysosomal storage disorder in mice characterized by a dual deficiency of sphingomyelinase and glucocerebrosidase. Biochim Biophys Acta. 1980;619(3):669–79.
Weintraub H, Abramovici A, Sandbank U, Booth AD, Pentchev PG, Sela B. Dysmyelination in NCTR-Balb/C mouse mutant with a lysosomal storage disorder. Morphological survey. Acta Neuropathol. 1987;74(4):374–81.
Beltroy EP, Richardson JA, Horton JD, Turley SD, Dietschy JM. Cholesterol accumulation and liver cell death in mice with Niemann-Pick type C disease. Hepatology. 2005;42(4):886–93. https://doi.org/10.1002/hep.20868.
Yu T, Shakkottai VG, Chung C, Lieberman AP. Temporal and cell-specific deletion establishes that neuronal Npc1 deficiency is sufficient to mediate neurodegeneration. Hum Mol Genet. 2011;20(22):4440–51. https://doi.org/10.1093/hmg/ddr372.
Elrick MJ, Pacheco CD, Yu T, Dadgar N, Shakkottai VG, Ware C, et al. Conditional Niemann-Pick C mice demonstrate cell autonomous Purkinje cell neurodegeneration. Hum Mol Genet. 2010;19(5):837–47. https://doi.org/10.1093/hmg/ddp552.
Karten B, Vance DE, Campenot RB, Vance JE. Cholesterol accumulates in cell bodies, but is decreased in distal axons, of Niemann-Pick C1-deficient neurons. J Neurochem. 2002;83(5):1154–63. https://doi.org/10.1046/j.1471-4159.2002.01220.x.
Hawes CM, Wiemer H, Krueger SR, Karten B. Pre-synaptic defects of NPC1-deficient hippocampal neurons are not directly related to plasma membrane cholesterol. J Neurochem. 2010;114(1):311–22. https://doi.org/10.1111/j.1471-4159.2010.06768.x.
Peake KB, Vance JE. Normalization of cholesterol homeostasis by 2-hydroxypropyl-beta-cyclodextrin in neurons and glia from Niemann-Pick C1 (NPC1)-deficient mice. J Biol Chem. 2012;287(12):9290–8. https://doi.org/10.1074/jbc.M111.326405.
Mizutani Y, Kihara A, Igarashi Y. LASS3 (longevity assurance homologue 3) is a mainly testis-specific (dihydro) ceramide synthase with relatively broad substrate specificity. The Biochemical Journal. 2006;398(3):531–8. https://doi.org/10.1042/BJ20060379.
Zhao L, Spassieva SD, Jucius TJ, Shultz LD, Shick HE, Macklin WB, et al. A deficiency of ceramide biosynthesis causes cerebellar Purkinje cell neurodegeneration and lipofuscin accumulation. PLoS Genet. 2011;7(5):e1002063. https://doi.org/10.1371/journal.pgen.1002063.
Laviad EL, Albee L, Pankova-Kholmyansky I, Epstein S, Park H, Merrill AH Jr, et al. Characterization of ceramide synthase 2: tissue distribution, substrate specificity, and inhibition by sphingosine 1-phosphate. J Biol Chem. 2008;283(9):5677–84. https://doi.org/10.1074/jbc.M707386200.
Ginkel C, Hartmann D, vom Dorp K, Zlomuzica A, Farwanah H, Eckhardt M, et al. Ablation of neuronal ceramide synthase 1 in mice decreases ganglioside levels and expression of myelin-associated glycoprotein in oligodendrocytes. J Biol Chem. 2012;287(50):41888–902. https://doi.org/10.1074/jbc.M112.413500.
Levy M, Futerman AH. Mammalian ceramide synthases. IUBMB Life. 2010;62(5):347–56. https://doi.org/10.1002/iub.319.
Merrill AH, Jr. (2002) De novo sphingolipid biosynthesis: a necessary, but dangerous, pathway. J Biol Chem 277 (29):25843–25846. doi:https://doi.org/10.1074/jbc.R200009200.
Linke T, Wilkening G, Lansmann S, Moczall H, Bartelsen O, Weisgerber J, et al. Stimulation of acid sphingomyelinase activity by lysosomal lipids and sphingolipid activator proteins. Biol Chem. 2001;382(2):283–90. https://doi.org/10.1515/BC.2001.035.
Brady RO, Kanfer JN, Shapiro D. Metabolism of glucocerebrosides. Ii. Evidence of an enzymatic deficiency in Gaucher’s disease. Biochem Biophys Res Commun. 1965;18(2):221–5. https://doi.org/10.1016/0006-291X(65)90743-6.
Heinrich M, Wickel M, Winoto-Morbach S, Schneider-Brachert W, Weber T, Brunner J, et al. Ceramide as an activator lipid of cathepsin D. In: Langner J, Ansorge S, editors. Cellular peptidases in immune functions and diseases, vol. 2. Boston, MA: Springer US; 2002. p. 305–15. https://doi.org/10.1007/0-306-46826-3_33.
Zaidi N, Maurer A, Nieke S, Kalbacher H. Cathepsin D: a cellular roadmap. Biochem Biophys Res Commun. 2008;376(1):5–9. https://doi.org/10.1016/j.bbrc.2008.08.099.
Heinrich M, Neumeyer J, Jakob M, Hallas C, Tchikov V, Winoto-Morbach S, et al. Cathepsin D links TNF-induced acid sphingomyelinase to bid-mediated caspase-9 and -3 activation. Cell Death Differ. 2004;11(5):550–63. https://doi.org/10.1038/sj.cdd.4401382.
Benes P, Vetvicka V, Fusek M. Cathepsin D--many functions of one aspartic protease. Crit Rev Oncol Hematol. 2008;68(1):12–28. https://doi.org/10.1016/j.critrevonc.2008.02.008.
Amritraj A, Peake K, Kodam A, Salio C, Merighi A, Vance JE, et al. Increased activity and altered subcellular distribution of lysosomal enzymes determine neuronal vulnerability in Niemann-Pick type C1-deficient mice. Am J Pathol. 2009;175(6):2540–56. https://doi.org/10.2353/ajpath.2009.081096.
Cluzeau CV, Watkins-Chow DE, Fu R, Borate B, Yanjanin N, Dail MK, et al. Microarray expression analysis and identification of serum biomarkers for Niemann-Pick disease, type C1. Hum Mol Genet. 2012;21(16):3632–46. https://doi.org/10.1093/hmg/dds193.
Fan M, Sidhu R, Fujiwara H, Tortelli B, Zhang J, Davidson C, et al. Identification of Niemann-Pick C1 disease biomarkers through sphingolipid profiling. J Lipid Res. 2013;54(10):2800–14. https://doi.org/10.1194/jlr.M040618.
Tobias F, Olson MT, Cologna SM. Mass spectrometry imaging of lipids: untargeted consensus spectra reveal spatial distributions in Niemann-Pick disease type C1. J Lipid Res. 2018;59(12):2446–55. https://doi.org/10.1194/jlr.D086090.
Angel PM, Spraggins JM, Baldwin HS, Caprioli R. Enhanced sensitivity for high spatial resolution lipid analysis by negative ion mode matrix assisted laser desorption ionization imaging mass spectrometry. Anal Chem. 2012;84(3):1557–64. https://doi.org/10.1021/ac202383m.
Robichaud G, Garrard KP, Barry JA, Muddiman DC. MSiReader: an open-source interface to view and analyze high resolving power MS imaging files on Matlab platform. J Am Soc Mass Spectrom. 2013;24(5):718–21. https://doi.org/10.1007/s13361-013-0607-z.
Bokhart MT, Nazari M, Garrard KP, Muddiman DC. MSiReader v1.0: evolving open-source mass spectrometry imaging software for targeted and untargeted analyses. J Am Soc Mass Spectrom. 2018;29(1):8–16. https://doi.org/10.1007/s13361-017-1809-6.
Vanier MT. Complex lipid trafficking in Niemann-Pick disease type C. J Inherit Metab Dis. 2015;38(1):187–99. https://doi.org/10.1007/s10545-014-9794-4.
Fields RD. Neuroscience Change in the brain’s white matter. Science. 2010;330(6005):768–9. https://doi.org/10.1126/science.1199139.
Abi-Mosleh L, Infante RE, Radhakrishnan A, Goldstein JL, Brown MS. Cyclodextrin overcomes deficient lysosome-to-endoplasmic reticulum transport of cholesterol in Niemann-Pick type C cells. Proc Natl Acad Sci U S A. 2009;106(46):19316–21. https://doi.org/10.1073/pnas.0910916106.
Zhang M, Strnatka D, Donohue C, Hallows JL, Vincent I, Erickson RP. Astrocyte-only Npc1 reduces neuronal cholesterol and triples life span of Npc1(−/−) mice. J Neurosci Res. 2008;86(13):2848–56. https://doi.org/10.1002/jnr.21730.
Takikita S, Fukuda T, Mohri I, Yagi T, Suzuki K. Perturbed myelination process of premyelinating oligodendrocyte in Niemann-Pick type C mouse. J Neuropathol Exp Neurol. 2004;63(6):660–73. https://doi.org/10.1093/jnen/63.6.660.
Kaya I, Brinet D, Michno W, Syvanen S, Sehlin D, Zetterberg H, et al. Delineating amyloid plaque associated neuronal sphingolipids in transgenic Alzheimer’s disease mice (tgArcSwe) using MALDI imaging mass spectrometry. ACS Chem Neurosci. 2017;8(2):347–55. https://doi.org/10.1021/acschemneuro.6b00391.
Hsu FF. Complete structural characterization of ceramides as [M-H](−) ions by multiple-stage linear ion trap mass spectrometry. Biochimie. 2016;130:63–75. https://doi.org/10.1016/j.biochi.2016.07.012.
Jennemann R, Sandhoff R, Wang S, Kiss E, Gretz N, Zuliani C, et al. Cell-specific deletion of glucosylceramide synthase in brain leads to severe neural defects after birth. Proc Natl Acad Sci U S A. 2005;102(35):12459–64. https://doi.org/10.1073/pnas.0500893102.
Costello CE, Juhasz P, Perreault H. New mass spectral approaches to ganglioside structure determinations. Prog Brain Res. 1994;101:45–61.
Kotani M, Kawashima I, Ozawa H, Terashima T, Tai T. Differential distribution of major gangliosides in rat central nervous system detected by specific monoclonal antibodies. Glycobiology. 1993;3(2):137–46. https://doi.org/10.1093/glycob/3.2.137.
Heffer-Lauc M, Lauc G, Nimrichter L, Fromholt SE, Schnaar RL. Membrane redistribution of gangliosides and glycosylphosphatidylinositol-anchored proteins in brain tissue sections under conditions of lipid raft isolation. Biochim Biophys Acta. 2005;1686(3):200–8. https://doi.org/10.1016/j.bbalip.2004.10.002.
Vajn K, Viljetic B, Degmecic IV, Schnaar RL, Heffer M. Differential distribution of major brain gangliosides in the adult mouse central nervous system. PLoS One. 2013;8(9):e75720. https://doi.org/10.1371/journal.pone.0075720.
Sarna JR, Larouche M, Marzban H, Sillitoe RV, Rancourt DE, Hawkes R. Patterned Purkinje cell degeneration in mouse models of Niemann-Pick type C disease. J Comp Neurol. 2003;456(3):279–91. https://doi.org/10.1002/cne.10522.
Vance JE, Karten B. Niemann-Pick C disease and mobilization of lysosomal cholesterol by cyclodextrin. J Lipid Res. 2014;55(8):1609–21. https://doi.org/10.1194/jlr.R047837.
Lopez ME, Klein AD, Scott MP. Complement is dispensable for neurodegeneration in Niemann-Pick disease type C. J Neuroinflammation. 2012;9(1):216. https://doi.org/10.1186/1742-2094-9-216.
Osiecki-Newman K, Legler G, Grace M, Dinur T, Gatt S, Desnick RJ, et al. Human acid beta-glucosidase: inhibition studies using glucose analogues and pH variation to characterize the normal and Gaucher disease glycon binding sites. Enzyme. 1988;40(4):173–88.
Goni FM, Alonso A. Sphingomyelinases: enzymology and membrane activity. FEBS Lett. 2002;531(1):38–46.
Linke T, Wilkening G, Sadeghlar F, Mozcall H, Bernardo K, Schuchman E, et al. Interfacial regulation of acid ceramidase activity. Stimulation of ceramide degradation by lysosomal lipids and sphingolipid activator proteins. J Biol Chem. 2001;276(8):5760–8. https://doi.org/10.1074/jbc.M006846200.
Bernardo K, Hurwitz R, Zenk T, Desnick RJ, Ferlinz K, Schuchman EH, et al. Purification, characterization, and biosynthesis of human acid ceramidase. J Biol Chem. 1995;270(19):11098–102. https://doi.org/10.1074/jbc.270.19.11098.
Schulze H, Sandhoff K. Lysosomal lipid storage diseases. Cold Spring Harb Perspect Biol. 3(6):a004804. https://doi.org/10.1101/cshperspect.a004804.
Tamura A, Nishida K, Yui N. Lysosomal pH-inducible supramolecular dissociation of polyrotaxanes possessing acid-labile N-triphenylmethyl end groups and their therapeutic potential for Niemann-Pick type C disease. Sci Technol Adv Mater. 2016;17(1):361–74. https://doi.org/10.1080/14686996.2016.1200948.
Pergande MR, Nguyen TTA, Haney-Ball C, Davidson CD, Cologna SM. Quantitative, label-free proteomics in the symptomatic Niemann-Pick, type C1 mouse model using standard flow liquid chromatography and thermal focusing electrospray ionization. Proteomics. 2019;19(9):e1800432. https://doi.org/10.1002/pmic.201800432.
Heinrich M, Wickel M, Schneider-Brachert W, Sandberg C, Gahr J, Schwandner R, et al. Cathepsin D targeted by acid sphingomyelinase-derived ceramide. EMBO J. 1999;18(19):5252–63. https://doi.org/10.1093/emboj/18.19.5252.
Li H, Repa JJ, Valasek MA, Beltroy EP, Turley SD, German DC, et al. Molecular, anatomical, and biochemical events associated with neurodegeneration in mice with Niemann-Pick type C disease. J Neuropathol Exp Neurol. 2005;64(4):323–33. https://doi.org/10.1093/jnen/64.4.323.
German DC, Liang CL, Song T, Yazdani U, Xie C, Dietschy JM. Neurodegeneration in the Niemann-Pick C mouse: glial involvement. Neuroscience. 2002;109(3):437–50.
Jin LW, Shie FS, Maezawa I, Vincent I, Bird T. Intracellular accumulation of amyloidogenic fragments of amyloid-beta precursor protein in neurons with Niemann-Pick type C defects is associated with endosomal abnormalities. Am J Pathol. 2004;164(3):975–85.
Liao G, Yao Y, Liu J, Yu Z, Cheung S, Xie A, et al. Cholesterol accumulation is associated with lysosomal dysfunction and autophagic stress in Npc1 −/− mouse brain. Am J Pathol. 2007;171(3):962–75. https://doi.org/10.2353/ajpath.2007.070052.
Acknowledgments
The authors would like to acknowledge the support from the University of Illinois at Chicago, Department of Chemistry and College of Liberal Arts and Sciences, and the Ara Parseghian Medical Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
ABC Highlights: authored by Rising Stars and Top Experts.
Electronic supplementary material
ESM 1
(PDF 2122 kb)
Rights and permissions
About this article
Cite this article
Tobias, F., Pathmasiri, K.C. & Cologna, S.M. Mass spectrometry imaging reveals ganglioside and ceramide localization patterns during cerebellar degeneration in the Npc1−/− mouse model. Anal Bioanal Chem 411, 5659–5668 (2019). https://doi.org/10.1007/s00216-019-01989-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-019-01989-7