Abstract
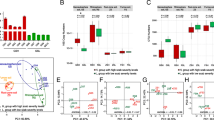
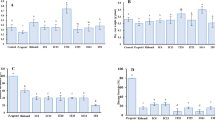
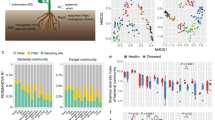
Common scab (CS) caused by pathogenic Streptomyces spp. plays a decisive role in the qualitative and quantitative production of potatoes worldwide. Although the CS pathogen is present in Assam’s soil, disease signs and symptoms are less obvious in the landrace Rongpuria potatoes that indicate an interesting interaction between the plant and the geocaulosphere microbial population. Toward this, a comparative metagenomics study was performed to elucidate the geocaulosphere microbiome assemblages and functions of low CS-severe (LSG) and moderately severe (MSG) potato plants. Alpha diversity indices showed that CS occurrence modulated microbiome composition and decreased overall microbial abundances. Functional analysis involving cluster of orthologous groups (COG) too confirmed reduced microbial metabolism under disease incidence. The top-three most dominant genera were Pseudomonas (relative abundance: 2.79% in LSG; 12.31% in MSG), Streptomyces (2.55% in LSG; 5.28% in MSG), and Pantoea (2.30% in LSG; 3.51% in MSG). As shown by the high Pielou’s J evenness index, the potato geocaulosphere core microbiome was adaptive and resilient to CS infection. The plant growth-promoting traits and potential antagonistic activity of major taxa (Pseudomonads, non-pathogenic Streptomyces spp., and others) against the CS pathogen, i.e., Streptomyces scabiei, point toward selective microbial recruitment and colonization strategy by the plants to its own advantage. KEGG Orthology analysis showed that the CS infection resulted in high abundances of ATP-binding cassette transporters and a two-component system, ubiquitous to the transportation and regulation of metabolites. As compared to the LSG metagenome, the MSG counterpart had a higher representation of important PGPTs related to 1-aminocyclopropane-1-carboxylate deaminase, IAA production, betaine utilization, and siderophore production.




Similar content being viewed by others
Data availability
The metagenomes data are available in the Sequence Read Archive hosted at the NCBI.
References
Andrade MHML, Niederheitmann M, de Paula Ribeiro SRR et al (2019) Development and validation of a standard area diagram to assess common scab in potato tubers. Eur J Plant Pathol 154:739–750. https://doi.org/10.1007/s10658-019-01697-z
Arseneault T, Goyer C, Filion M (2016) Biocontrol of Potato common scab is associated with high Pseudomonas fluorescens LBUM223 populations and phenazine-1-carboxylic acid biosynthetic transcript accumulation in the Potato geocaulosphere®. Phytopathology 106:963–970. https://doi.org/10.1094/PHYTO-01-16-0019-R
Beneduzi A, Ambrosini A, Passaglia LMP (2012) Plant growth-promoting rhizobacteria (PGPR): their potential as antagonists and biocontrol agents. Genet Mol Biol 35:1044–1051
Bulgarelli D, Garrido-Oter R, Münch PC et al (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17:392–403. https://doi.org/10.1016/j.chom.2015.01.011
Canarini A, Kaiser C, Merchant A et al (2019) Root exudation of primary metabolites: mechanisms and their roles in plant responses to environmental stimuli. Front Plant Sci. https://doi.org/10.3389/fpls.2019.00157
Carvalhais LC, Dennis PG, Badri DV et al (2015) Linking jasmonic acid signaling, root exudates, and rhizosphere microbiomes. Mol Plant-Microbe Interact 28:1049–1058. https://doi.org/10.1094/MPMI-01-15-0016-R
Chin-A-Woeng TFC, Bloemberg GV, Lugtenberg BJJ (2003) Phenazines and their role in biocontrol by Pseudomonas bacteria. New Phytol 157:503–523. https://doi.org/10.1046/j.1469-8137.2003.00686.x
Cole JR, Wang Q, Fish JA et al (2014) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633. https://doi.org/10.1093/nar/gkt1244
da Rocha UN, van Elsas JD, van Overbeek LS (2010) Real-time PCR detection of Holophagae (Acidobacteria) and Verrucomicrobia subdivision 1 groups in bulk and leek (Allium porrum) rhizosphere soils. J Microbiol Methods 83:141–148. https://doi.org/10.1016/j.mimet.2010.08.003
Dintner S, Heermann R, Fang C et al (2014) A Sensory complex consisting of an ATP-binding cassette transporter and a two-component regulatory system controls bacitracin resistance in bacillus subtilis. J Biol Chem 289:27899–27910. https://doi.org/10.1074/jbc.M114.596221
Dintner S, Staroń A, Berchtold E et al (2011) Coevolution of ABC transporters and two-component regulatory systems as resistance modules against antimicrobial peptides in Firmicutes bacteria. J Bacteriol 193:3851–3862. https://doi.org/10.1128/JB.05175-11
Edwards J, Johnson C, Santos-Medellín C et al (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci 112:E911–E920. https://doi.org/10.1073/pnas.1414592112
Elango D (2019) Gender responsive participatory varietal selection for sustainable seed potato systems in Assam, India
Ezekiel R, Rana G (2009) Physicochemical properties of potato starch in relation to cultivar, growing location, season and crop maturity. Adv Hortic Sci 23:94–100
Faucher E, Otrysko B, Paradis É et al (1993) Characterization of streptomycetes causing russet scab in quebec. Plant Dis 77:1217. https://doi.org/10.1094/PD-77-1217
Feng G, Xie T, Wang X et al (2018) Metagenomic analysis of microbial community and function involved in cd-contaminated soil. BMC Microbiol 18:11. https://doi.org/10.1186/s12866-018-1152-5
Foyer CH, Noctor G, Hodges M (2011) Respiration and nitrogen assimilation: targeting mitochondria-associated metabolism as a means to enhance nitrogen use efficiency. J Exp Bot 62:1467–1482. https://doi.org/10.1093/jxb/erq453
Gómez Expósito R, de Bruijn I, Postma J, Raaijmakers JM (2017) Current insights into the role ofrhizosphere bacteria in disease suppressive soils. Front Microbiol. https://doi.org/10.3389/fmicb.2017.02529
Gopalakrishnan S, Sathya A, Vijayabharathi R et al (2015) Plant growth promoting rhizobia: challenges and opportunities. 3 Biotech. 5:355–377. https://doi.org/10.1007/s13205-014-0241-x
Gupta S, Pandey S (2019) ACC Deaminase producing bacteria with multifarious plant growth promoting traits alleviates salinity stress in French Bean (Phaseolus vulgaris) plants. Front Microbiol. https://doi.org/10.3389/fmicb.2019.01506
Hakim S, Naqqash T, Nawaz MS et al (2021) Rhizosphere engineering With plant growth-promoting microorganisms for agriculture and ecological sustainability. Front Sustain Food Syst. https://doi.org/10.3389/fsufs.2021.617157
Hamel C, Gan Y, Sokolski S, Bainard LD (2018) High frequency cropping of pulses modifies soil nitrogen level and the rhizosphere bacterial microbiome in 4-year rotation systems of the semiarid prairie. Appl Soil Ecol 126:47–56. https://doi.org/10.1016/j.apsoil.2018.01.003
Hudec C, Novinscak A, Filion M (2021) Diversity and virulence of Streptomyces spp. causing potato common scab in Prince Edward Island®. Canada. Phytopathology. 111:617–626. https://doi.org/10.1094/PHYTO-08-20-0339-R
Huerta-Cepas J, Szklarczyk D, Forslund K et al (2016) EGGNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res 44:D286–D293. https://doi.org/10.1093/nar/gkv1248
Hunziker L, Bönisch D, Groenhagen U et al (2015) Pseudomonas strains naturally associated with potato plants produce volatiles with high potential for inhibition of Phytophthora infestans. Appl Environ Microbiol 81:821–830. https://doi.org/10.1128/AEM.02999-14
Hyatt D, Chen GL, LoCascio PF et al (2010) Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119. https://doi.org/10.1186/1471-2105-11-119
İnceoğlu Ö, Al-Soud WA, Salles JF et al (2011) Comparative analysis of bacterial communities in a potato field as determined by pyrosequencing. PLoS ONE 6:e23321. https://doi.org/10.1371/journal.pone.0023321
Kielak AM, Barreto CC, Kowalchuk GA et al (2016) The ecology of Acidobacteria: moving beyond genes and genomes. Front Microbiol. https://doi.org/10.3389/fmicb.2016.00744
Köberl M, Dita M, Martinuz A et al (2017) Members of Gammaproteobacteria as indicator species of healthy banana plants on Fusarium wilt-infested fields in Central America. Sci Rep 7:45318. https://doi.org/10.1038/srep45318
Koco Š, Vilček J, Torma S et al (2020) Optimising potato (Solanum tuberosum L.) cultivation by selection of proper soils. Agriculture 10:155. https://doi.org/10.3390/agriculture10050155
Leontidou K, Genitsaris S, Papadopoulou A et al (2020) Plant growth promoting rhizobacteria isolated from halophytes and drought-tolerant plants: genomic characterisation and exploration of phyto-beneficial traits. Sci Rep 10:14857. https://doi.org/10.1038/s41598-020-71652-0
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25:1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li J, Wang C, Liang W, Liu S (2021) Rhizosphere microbiome: the emerging barrier in plant-pathogen interactions. Front Microbiol. https://doi.org/10.3389/fmicb.2021.772420
Liu D (1995) Biological control of potato scab in the field with antagonistic Streptomyces scabies. Phytopathology 85:827. https://doi.org/10.1094/Phyto-85-827
Ma L, Cao YH, Cheng MH et al (2013) Phylogenetic diversity of bacterial endophytes of Panax notoginseng with antagonistic characteristics towards pathogens of root-rot disease complex. Antonie Van Leeuwenhoek 103:299–312. https://doi.org/10.1007/s10482-012-9810-3
Mardanova A, Lutfullin M, Hadieva G et al (2019) Structure and variation of root-associated microbiomes of potato grown in alfisol. World J Microbiol Biotechnol 35:181. https://doi.org/10.1007/s11274-019-2761-3
Morrison CK, Novinscak A, Gadkar VJ et al (2016) Complete genome sequence of Pseudomonas fluorescens LBUM636, a strain with biocontrol capabilities against late blight of potato. Genome Announc 4. https://doi.org/10.1128/genomeA.00446-16
Neeno-Eckwall EC, Kinkel LL, Schottel JL (2001) Competition and antibiosis in the biological control of potato scab. Can J Microbiol 47:332–340. https://doi.org/10.1139/w01-010
Pascale A, Proietti S, Pantelides IS, Stringlis IA (2020) Modulation of the root microbiome by plant molecules: the basis for targeted disease suppression and plant growth promotion. Front Plant Sci. https://doi.org/10.3389/fpls.2019.01741
Pfeiffer S, Mitter B, Oswald A et al (2017) Rhizosphere microbiomes of potato cultivated in the High Andes show stable and dynamic core microbiomes with different responses to plant development. FEMS Microbiol Ecol 93:242. https://doi.org/10.1093/femsec/fiw242
Pielou EC (1974) Population and community ecology. Gordon and Breach Science Publishers
Pieterse CMJ, Zamioudis C, Berendsen RL et al (2014) Induced systemic resistance by beneficial microbes. Annu Rev Phytopathol 52:347–375. https://doi.org/10.1146/annurev-phyto-082712-102340
Qu XS, Wanner LA, Christ BJ (2011) Multiplex real-time PCR (TaqMan) assay for the simultaneous detection and discrimination of potato powdery and common scab diseases and pathogens. J Appl Microbiol 110:769–777. https://doi.org/10.1111/j.1365-2672.2010.04930.x
Razaq M, Zhang P, Shen H, Salahuddin (2017) Influence of nitrogen and phosphorous on the growth and root morphology of Acer mono. PLoS ONE 12:e0171321. https://doi.org/10.1371/journal.pone.0171321
Rolfe SA, Griffiths J, Ton J (2019) Crying out for help with root exudates: adaptive mechanisms by which stressed plants assemble health-promoting soil microbiomes. Curr Opin Microbiol 49:73–82. https://doi.org/10.1016/j.mib.2019.10.003
Sarwar A, Latif Z, Zhang S et al (2018) Biological control of potato common scab with rare isatropolone C compound produced by plant growth promoting Streptomyces A1RT. Front Microbiol. https://doi.org/10.3389/fmicb.2018.01126
Sharma SB, Sayyed RZ, Trivedi MH, Gobi TA (2013) Phosphate solubilizing microbes: sustainable approach for managing phosphorus deficiency in agricultural soils. Springerplus 2:587. https://doi.org/10.1186/2193-1801-2-587
Shi W, Li M, Wei G et al (2019) The occurrence of potato common scab correlates with the community composition and function of the geocaulosphere soil microbiome. Microbiome 7:14. https://doi.org/10.1186/s40168-019-0629-2
Tsurumaru H, Okubo T, Okazaki K et al (2015) Metagenomic analysis of the bacterial community associated with the taproot of sugar beet. Microbes Environ 30:63–69. https://doi.org/10.1264/jsme2.ME14109
Wagner A, Norris S, Chatterjee P et al (2018) Aquatic Pseudomonads inhibit Oomycete plant pathogens of Glycine max. Front Microbiol. https://doi.org/10.3389/fmicb.2018.01007
Wagner MR, Lundberg DS, del Rio TG et al (2016) Host genotype and age shape the leaf and root microbiomes of a wild perennial plant. Nat Commun 7:12151. https://doi.org/10.1038/ncomms12151
Wu S, Zhu Z, Fu L et al (2011) WebMGA: a customizable web server for fast metagenomic sequence analysis. BMC Genomics 12:444. https://doi.org/10.1186/1471-2164-12-444
Xie C, Mao X, Huang J et al (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:316. https://doi.org/10.1093/nar/gkr483
Yazaki K (2006) ABC transporters involved in the transport of plant secondary metabolites. FEBS Lett 580:1183–1191. https://doi.org/10.1016/j.febslet.2005.12.009
Yossen V, Rojo R, Chiessa G et al (2011) Effect of green manure and biocontrol agents on potato crop in Cordoba, Argentina. J Plant Pathol 93:713–717
Zeng Y, Charkowski AO (2021) The role of ATP-binding cassette transporters in bacterial phytopathogenesis. Phytopathology 111:600–610. https://doi.org/10.1094/PHYTO-06-20-0212-RVW
Zengerer V, Schmid M, Bieri M et al (2018) Pseudomonas orientalis F9: a potent antagonist against phytopathogens with phytotoxic effect in the Apple flower. Front Microbiol. https://doi.org/10.3389/fmicb.2018.00145
Acknowledgements
The authors kindly acknowledge the timely suggestions and encouragement received from Professor B. K. Sarmah (Director, DBT-NECAB Centre, AAU, Jorhat) and Prof. M. K. Modi (Head, Department of Agricultural Biotechnology, AAU, Jorhat) throughout the research work.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
MB had conceived and designed the study. SSB, MB, DJH, AC, and AD carried out the experiments and collected the data. RN, RC, HK, SSB, and MB had analyzed the field data. All the authors contributed in preparing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare that they have no conflict of interest. The authors have no relevant financial or non-financial interests to disclose.
Ethics approval
This study involved microbiomes associated with cultivated potato tubers only.
Consent to participate
NA.
Consent for publication
No public participants were involved in this study. All the authors have given consent for publication.
Additional information
Communicated by Yusuf Akhter.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bora, S.S., Hazarika, D.J., Churaman, A. et al. Common scab disease-induced changes in geocaulosphere microbiome assemblages and functional processes in landrace potato (Solanum tuberosum var. Rongpuria) of Assam, India. Arch Microbiol 205, 44 (2023). https://doi.org/10.1007/s00203-022-03380-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-022-03380-0