Abstract
Key message
In wheat, multiple disease resistance meta-QTLs (MDR-MQTLs) and underlying candidate genes for the three rusts were identified which may prove useful for development of resistant cultivars.
Abstract
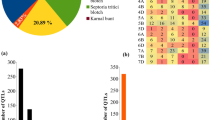
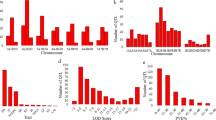
Rust diseases in wheat are a major threat to global food security. Therefore, development of multiple disease-resistant cultivars (resistant to all three rusts) is a major goal in all wheat breeding programs worldwide. In the present study, meta-QTLs and candidate genes for multiple disease resistance (MDR) involving all three rusts were identified using 152 individual QTL mapping studies for resistance to leaf rust (LR), stem rust (SR), and yellow rust (YR). From these 152 studies, a total of 1,146 QTLs for resistance to three rusts were retrieved, which included 368 QTLs for LR, 291 QTLs for SR, and 487 QTLs for YR. Of these 1,146 QTLs, only 718 QTLs could be projected onto the consensus map saturated with 2, 34,619 markers. Meta-analysis of the projected QTLs resulted in the identification of 86 MQTLs, which included 71 MDR-MQTLs. Ten of these MDR-MQTLs were referred to as the ‘Breeders’ MQTLs’. Seventy-eight of the 86 MQTLs could also be anchored to the physical map of the wheat genome, and 54 MQTLs were validated by marker-trait associations identified during earlier genome-wide association studies. Twenty MQTLs (including 17 MDR-MQTLs) identified in the present study were co-localized with 44 known R genes. In silico expression analysis allowed identification of several differentially expressed candidate genes (DECGs) encoding proteins carrying different domains including the following: NBS-LRR, WRKY domains, F-box domains, sugar transporters, transferases, etc. The introgression of these MDR loci into high-yielding cultivars should prove useful for developing high yielding cultivars with resistance to all the three rusts.









Similar content being viewed by others
Availability of data and material
Additional data relevant to this paper are available in the form of supplementary material.
References
Aduragbemi A, Soriano JM (2021) Unravelling consensus genomic regions conferring leaf rust resistance in wheat via meta-QTL analysis. BMC Genom. https://doi.org/10.1101/2021.05.11.443557
Afzal SN, Haque MI, Ahmedani MS, Bashir S, Rattu AR (2007) Assessment of yield losses caused by Puccinia striiformis triggering stripe rust in the most common wheat varieties. Pak J Bot 39:2127–2134
Agenbag GM, Pretorius ZA, Boyd LA, Bender CM, MacCormack R, Prins R (2014) High-resolution mapping and new marker development for adult plant stripe rust resistance QTL in the wheat cultivar Kariega. Mol Breed 34:2005–2020
Alemu A, Feyissa T, Maccaferri M, Sciara G, Tuberosa R, Ammar K, Badebo A, Acevedo M, Letta T, Abeyo B (2021) Genome-wide association analysis unveils novel QTLs for seminal root system architecture traits in Ethiopian durum wheat. BMC Genomics 22:1–16
Ali F, Pan Q, Chen G, Zahid KR, Yan J (2013) Evidence of multiple disease resistance (MDR) and implication of meta-analysis in marker assisted selection. PLoS ONE 8:e68150
Arcade A, Labourdette A, Falque M, Mangin B, Chardon F, Charcosset A, Joets J (2004) BioMercator: integrating genetic maps and QTL towards discovery of candidate genes. Bioinformatics 20:2324–2326
Ausemus ER (1943) Breeding for disease resistance in wheat, oats, barleyand flax. Bot Rev 9:207–260
Bajgain P, Rouse MN, Tsilo TJ, Macharia GK, Bhavani S, Jin Y, Anderson JA (2016) Nested association mapping of stem rust resistance in wheat using genotyping by sequencing. PLoS ONE 11:e0155760
Bemister DH, Semagn K, Iqbal M, Randhawa H, Strelkov SE, Spaner DM (2019) Mapping QTL associated with stripe rust, leaf rust, and leaf spotting in a Canadian spring wheat population. Crop Sci 59:650–658
Brueggeman R, Druka A, Nirmala J, Cavileer T, Drader T, Rostoks N, Mirlohi A, Bennypaul H, Gill U, Kudrna D, Whitelaw C (2008) The stem rust resistance gene Rpg5 encodes a protein with nucleotide-binding-site, leucine-rich, and protein kinase domains. Proc Natl Acad Sci 105:14970–14975
Cao A, Xing L, Wang X, Yang X, Wang W, Sun Y, Qian C, Ni J, Chen Y, Liu D, Wang X (2011) Serine/threonine kinase gene Stpk-V, a key member of powdery mildew resistance gene Pm21, confers powdery mildew resistance in wheat. Proc Natl Acad Sci 108:7727–7732
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci 110:8057–8062
Chardon F, Virlon B, Moreau L, Falque M, Joets J, Decousset L, Murigneux A, Charcosset A (2004) Genetic architecture of flowering time in maize as inferred from quantitative trait loci meta-analysis and synteny conservation with the rice genome. Genetics 168:2169–2185
Chen XM (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp. tritici] on wheat. Can J Plant Pathol 27:314–337
Chen S, Zhang W, Bolus S, Rouse MN, Dubcovsky J (2018) Identification and characterization of wheat stem rust resistance gene Sr21 effective against the Ug99 race group at high temperature. PLoS Genet 14:e1007287
Cloutier S, Wang Z, Banks TW, Jordan MC, McCallum BD (2008) Gene expression of plant defence pathways using Lr1 transgenic lines and the Affymetrix wheat Chip. In: The 11th international wheat genetics symposium proceedings edited by Rudi Appels Russell Eastwood Evans Lagudah Peter Langridge Michael Mackay Lynne. Sydney University Press
Coram TE, Settles ML, Chen X (2008) Transcriptome analysis of high-temperature adult-plant resistance conditioned by Yr39 during the wheat–Puccinia striiformis f. sp. tritici interaction. Mol Plant Pathol 9:479–493
Cui F, Zhang N, Fan XL, Zhang W, Zhao CH, Yang LJ, Pan RQ, Chen M, Han J, Zhao XQ, Ji J (2017) Utilization of a Wheat 660K SNP array-derived high-density genetic map for high-resolution mapping of a major QTL for kernel number. Sci Rep 7:1–12
Darvasi A, Soller M (1997) A simple method to calculate resolving power and confidence interval of QTL map location. Behav Genet 27:125–132
Dholakia BB, Rajwade AV, Hosmani P, Khan RR, Chavan S, Reddy DMR, Lagu MD, Bansal UK, Saini RG, Gupta VS (2013) Molecular mapping of leaf rust resistance gene Lr15 in hexaploid wheat. Mol Breed 31:743–747
Dobon A, Bunting DC, Cabrera-Quio LE, Uauy C, Saunders DG (2016) The host-pathogen interaction between wheat and yellow rust induces temporally coordinated waves of gene expression. BMC Genom 17:1–14
Edae EA, Rouse MN (2020) Association mapping of resistance to emerging stem rust pathogen races in spring wheat using genotyping-by-sequencing. The Plant Genome 13:e20050
Endelman JB, Plomion C (2014) LPmerge: an R package for merging genetic maps by linear programming. Bioinformatics 30:1623–1624
Eriksen L, Afshari F, Christiansen MJ, McIntosh RA, Jahoor A, Wellings CR (2004) Yr32 for resistance to stripe (yellow) rust present in the wheat cultivar Carstens V. Theor Appl Genet 108:567–575
Fatima F, McCallum BD, Pozniak CJ, Hiebert CW, McCartney CA, Fedak G, You FM, Cloutier S (2020) Identification of new leaf rust resistance loci in wheat and wild relatives by array-based SNP genotyping and association genetics. Front Plant Sci 11:1728
Flemmig EL (2012) Molecular markers to deploy and characterize stem rust resistance in wheat. Dissertation, North Carolina State University
Flor HH (1942) Inheritance of pathogenicity in Melampsora lini. Phytopathology 32:653–669
Flor HH (1971) Current status of the gene-for-gene concept. Annu Rev Phytopathol 9:275–296
Friesen TL, Xu SS, Harris MO (2008) Stem rust, tan spot, Stagonosporanodorum blotch, and Hessian fly resistance in Langdon durum-Aegilops tauschii synthetic hexaploid wheat lines. Crop Sci 48:1062–1070
Gao L, Turner MK, Chao S, Kolmer J, Anderson JA (2016) Genome wide association study of seedling and adult plant leaf rust resistance in elite spring wheat breeding lines. PLoS One 11:e0148671
Genievskaya Y, Turuspekov Y, Rsaliyev A, Abugalieva S (2020) Genome-wide association mapping for resistance to leaf, stem, and yellow rusts of common wheat under field conditions of South Kazakhstan. Peer J 8:e9820
Goffinet B, Gerber S (2000) Quantitative trait loci: a meta-analysis. Genetics 155:463–473
Gudi S, Saini DK, Singh G, Halladakeri P, Shamshad M, Tanin MJ, Kumar P, Sharma A (2021) Unravelling consensus genomic regions associated with quality traits in wheat (Triticum aestivum L.) using meta-analysis of quantitative trait loci. bioRxiv. https://doi.org/10.1101/2021.11.24.469810
Guo B, Sleper DA, Lu P, Shannon JG, Nguyen HT, Arelli PR (2006) QTLs associated with resistance to soybean cyst nematode in soybean: Meta-analysis of QTL locations. Crop Sci 46:595–602
Gupta PK, Balyan HS, Chhuneja P et al (2021) Pyramiding of genes for grain protein content, grain quality and rust resistance in eleven Indian bread wheat cultivars: a multi-Institutional effort. Mol Breed. https://doi.org/10.21203/rs.3.rs-637558/v1
Gurung S, Bonman JM, Ali S, Patel J, Myrfield M, Mergoum M, Singh PK, Adhikari TB (2009) New and diverse sources of multiple disease resistance in wheat. Crop Sci 49:1655–1666
Habib M, Awan FS, Sadia B, Zia MA (2020) Genome-wide association mapping for stripe rust resistance in Pakistani spring wheat genotypes. Plants 9:1056
Herrera-Foessel SA, Singh RP, Lillemo M, Huerta-Espino J, Bhavani S, Singh S, Lan C, Calvo-Salazar V, Lagudah ES (2014) Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor Appl Genet 127:781–789
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinform. https://doi.org/10.1155/2008/420747
Hu T, Zhong X, Yang Q, Zhou X, Li X, Yang S, Hou L, Yao Q, Guo Q, Kang Z (2020) Introgression of two quantitative trait loci for stripe rust resistance into three Chinese wheat cultivars. Agronomy 10:483
Huerta-Espino J, Singh RP, German S, McCallum BD, Park RF, Chen WQ, Bhardwaj SC, Goyeau H (2011) Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 179:143–160
Huerta-Espino J, Singh R, Crespo-Herrera LA, Villasenor-Mir HE, Rodriguez-Garcia MF, Dreisigacker S, Barcenas-Santana D, Lagudah E (2020) Adult plant slow rusting genes confer high levels of resistance to rusts in bread wheat cultivars from Mexico. Frontiers in Plant Sci 11:824
Jan I, Saripalli G, Kumar K, Kumar A, Singh R, Batra R, Sharma PK, Balyan HS, Gupta PK (2021) Meta-QTLs and genes for stripe rust resistance in wheat. Sci Rep 11:1–13
Jia M, Yang L, Zhang W, Rosewarne G, Li J, Yang E, Chen L, Wang W, Liu Y, Tong H, He W (2020) Genome wide association analysis of stripe rust resistance in modern Chinese wheat. BMC Plant Biol 20:1-13
Jighly A, Alagu M, Makdis F, Singh M, Singh S, Emebiri LC, Ogbonnaya FC (2016) Genomic regions conferring resistance to multiple fungal pathogens in synthetic hexaploid wheat. Mol Breed 36:1–19
Joshi LM, Singh DV, Srivastava KD (1985) Status of rusts and smuts in India. Barley, Wheat and Triticale Newsletter. ISSN: 0255-6421
Juliana P, He X, Kabir MR, Roy KK, Anwar MB, Marza F, Poland J, Shrestha S, Singh RP, Singh PK (2020) Genome-wide association mapping for wheat blast resistance in CIMMYT’s international screening nurseries evaluated in Bolivia and Bangladesh. Sci Rep 10:1–14
Kankwatsa P, Singh D, Thomson PC, Babiker EM, Bonman JM, Newcomb M, Park RF (2017) Characterization and genome-wide association mapping of resistance to leaf rust, stem rust and stripe rust in a geographically diverse collection of spring wheat landraces. Mol Breed 37:1–24
Kaur S, Kaur J, Mavi GS, Dhillon GS, Sharma A, Singh R, Devi U, Chhuneja P (2020) Pyramiding of high grain weight with stripe rust and leaf rust resistance in elite Indian wheat cultivar using a combination of marker-assisted and phenotypic selection. Front Genet 11:1680
Kaur B, Bhatia D, Mavi GS (2021) Eighty years of gene-for-gene relationship and its applications in identification and utilization of R genes. J Genet 100:1–17
Knott DR (1989) The effect of transfers of alien genes for leaf rust resistance on the agronomic and quality characteristics of wheat. Euphytica 44:65–72
Kolmer JA, Garvin DF, Jin Y (2011) Expression of a Thatcher wheat adult plant stem rust resistance QTL on chromosome arm 2BL is enhanced by Lr34. Crop Sci 51:526–533
Kou Y, Wang S (2010) Broad-spectrum and durability: understanding of quantitative disease resistance. Curr Opin Plant Biol 13:181–185
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Kumar IS, Nadarajah K (2020) A meta-analysis of quantitative trait loci associated with multiple disease resistance in rice (Oryza sativa L). Plants 9:1491
Kumar A, Saripalli G, Jan I, Kumar K, Sharma PK, Balyan HS, Gupta PK (2020a) Meta-QTL analysis and identification of candidate genes for drought tolerance in bread wheat (Triticum aestivum L.). Physiol Mol Biol Plants 26:1713–1725
Kumar D, Kumar A, Chhokar V, Gangwar OP, Bhardwaj SC, Sivasamy M, Prasad SV, Prakasha TL, Khan H, Singh R, Sharma P (2020b) Genome-wide association studies in diverse spring wheat panel for stripe, stem, and leaf rust resistance. Front Plant Sci 11:748
Kumar A, Saripalli G, Jan I, Kumar K, Sharma PK, Balyan HS, Gupta PK (2020c) Meta-QTL analysis and identification of candidate genes for drought tolerance in bread wheat (Triticum aestivum L.). Physiol Mol Biol Plants 26:1713–1725
Kumar S, Singh VP, Saini DK, Sharma H, Saripalli G, Kumar S et al (2021) Meta-QTLs, ortho-MQTLs, and candidate genes for thermotolerance in wheat (Triticum aestivum L.). Mol Breed 41:1–22
Lagudah ES, Krattinger SG, Herrera-Foessel S, Singh RP, Huerta-Espino J, Spielmeyer W, Brown-Guedira G, Selter LL, Keller B (2009) Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119:889–898
Ledesma-Ramírez L, Solís-Moya E, Iturriaga G, Sehgal D, Reyes-Valdes MH, Montero-Tavera V, Sansaloni CP, Burgueno J, Ortiz C, Aguirre-Mancilla CL, Ramírez-Pimentel JG (2019) GWAS to identify genetic loci for resistance to yellow rust in wheat pre-breeding lines derived from diverse exotic crosses. Front Plant Sci 10:1390
Leonova IN, Skolotneva ES, Salina EA (2020) Genome-wide association study of leaf rust resistance in Russian spring wheat varieties. BMC Plant Biol 20:1–13
Letta T, Maccaferri M, Badebo A, Ammar K, Ricci A, Crossa J, Tuberosa R (2013) Searching for novel sources of field resistance to Ug99 and Ethiopian stem rust races in durum wheat via association mapping. Theor Appl Genet 126:1237–1256
Li W, Deng Y, Ning Y, He Z, Wang GL (2020) Exploiting broad-spectrum disease resistance in crops: from molecular dissection to breeding. Annu Rev Plant Biol 71:575–603
Liu W, Frick M, Huel R, Nykiforuk CL, Wang X, Gaudet DA, Eudes F, Conner RL, Kuzyk A, Chen Q, Kang Z (2014) The stripe rust resistance gene Yr10 encodes an evolutionary-conserved and unique CC–NBS–LRR sequence in wheat. Mol Plant 7:1740–1755
Liu Y, Salsman E, Wang R, Galagedara N, Zhang Q, Fiedler JD, Liu Z, Xu S, Faris J, Li X (2020) Meta-QTL analysis of tan spot resistance in wheat. Theor Appl Genet 133:2363–2375
Maccaferri M, Zhang J, Bulli P, Abate Z, Chao S, Cantu D, Bossolini E, Chen X, Pumphrey M, Dubcovsky J (2015) A genome-wide association study of resistance to stripe rust (Puccinia striiformis f. sp. tritici) in a worldwide collection of hexaploid spring wheat (Triticum aestivum L.). G3-Genes Genomes Genet 5(3):449–465
Mago R, Tabe L, McIntosh RA, Pretorius Z, Kota R, Paux E, Wicker T, Breen J, Lagudah ES, Ellis JG, Spielmeyer W (2011) A multiple resistance locus on chromosome arm 3BS in wheat confers resistance to stem rust (Sr2) leaf rust (Lr27) and powdery mildew. Theor Appl Genet 123:615–623
Marchal C, Zhang J, Zhang P, Fenwick P, Steuernagel B, Adamski NM, Boyd L, McIntosh R, Wulff BB, Berry S, Lagudah E (2018) BED-domain-containing immune receptors confer diverse resistance spectra to yellow rust. Nat Plant 4:662–668
Marone D, Russo MA, Laidò G, De Vita P, Papa R, Blanco A, Gadaleta A, Rubiales D, Mastrangelo AM (2013) Genetic basis of qualitative and quantitative resistance to powdery mildew in wheat: from consensus regions to candidate genes. BMC Genom 14:1–17
McIntosh RA, Dubcovsky J, Rogers W, Morris C, Appels R, Xia X (2016) Catalogue of gene symbols for wheat: 2015–2016 supplement. Annual Wheat Newsletter 58:1–18
Megerssa SH, Ammar K, Acevedo M, Brown-Guedira G, Ward B, Degete AG, Randhawa MS, Sorrells ME (2020) Multiple-race stem rust resistance loci identified in durum wheat using genome-wide association mapping. Front Plant Sci 11:1934
Miedaner T, Rapp M, Flath K, Longin CFH, Würschum T (2019) Genetic architecture of yellow and stem rust resistance in a durum wheat diversity panel. Euphytica 215:1–17
Miedaner T, Akel W, Flath K, Jacobi A, Taylor M, Longin F, Würschum T (2020) Molecular tracking of multiple disease resistance in a winter wheat diversity panel. Theor Appl Genet 133:419–431
Moore JW, Herrera-Foessel S, Lan C, Schnippenkoetter W, Ayliffe M, Huerta-Espino J, Lillemo M, Viccars L, Milne R, Periyannan S, Kong X (2015) A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat Genet 47:1494–1498
Nene YL (1988) Multiple-disease resistance in grain legumes. Annu Rev Phytopathol 26:203–217
Oliver RP (2014) A reassessment of the risk of rust fungi developing resistance to fungicides. Pest Manag Sci 70:1641–1645
Pal N, Saini DK, Kumar S (2021) Meta-QTLs, ortho-MQTLs and candidate genes for the traits contributing to salinity stress tolerance in common wheat (Triticum aestivum L.). Physiol Mol Biol Plants 27:2767–2786
Pal N, Saini DK, Kumar S (2022) Breaking yield ceiling in wheat: progress and future prospects. In (Ed.), Wheat. IntechOpen. https://doi.org/10.5772/intechopen.102919
Paterson J (2014) Multiple disease resistance the holy grail. https://grdc.com.au/resources-and-publications/groundcover/ground-cover-supplements/gcs110/multiple-disease-resistance-the-holy-grail. Accessed 25 Jan 2019
Pooja S, Sharma RB, Singh AK, Rakesh S, Sundeep K (2014) Multiple disease resistance in wheat: need of today. Wheat Inf Serv 118:7–16
Pradhan AK, Kumar S, Singh AK, Budhlakoti N, Mishra DC, Chauhan D, Mittal S, Grover M, Kumar S, Gangwar OP, Kumar S (2020) Identification of QTLs/defense genes effective at seedling stage against prevailing races of wheat stripe rust in India. Front Genet 11:572975
Prins R, Pretorius ZA, Bender CM, Lehmensiek A (2011) QTL mapping of Stripe, leaf and stem rust resistance genes in a Kariega x Avocet S doubled haploid wheat population. Mol Breed 27:259–270
Ramirez-Gonzalez RH, Borrill P, Lang D, Harrington SA, Brinton J, Venturini L, Davey M, Jacobs J, Van Ex F, Pasha A, Khedikar Y (2018) The transcriptional landscape of polyploid wheat. Science 361:6403
Rana M, Kaldate R, Nabi SU, Wani SH, Khan H (2021) Marker-assisted breeding for resistance against wheat rusts. Physiological, molecular, and genetic perspectives of wheat improvement. Springer, Cham, pp 229–262
Randhawa MS, Lan C, Basnet BR, Bhavani S, Huerta-Espino J, Forrest KL, Hayden MJ, Singh RP (2018) Interactions among genes Sr2/Yr30, Lr34/Yr18/Sr57 and Lr68 confer enhanced adult plant resistance to rust diseases in common wheat (Triticum aestivum L.) line’Arula’. Aust J Crop Sci 12:1023–1033
Rani R, Singh R, Yadav NR (2019) Evaluating stripe rust resistance in Indian wheat genotypes and breeding lines using molecular markers. CR Biol 342:154–174
Rouse MN, Nirmala J, Jin Y, Chao S, Fetch TG, Pretorius ZA, Hiebert CW (2014) Characterization of Sr9h, a wheat stem rust resistance allele effective to Ug99. Theor Appl Genet 127:1681–1688
Rutter WB, Salcedo A, Akhunova A, He F, Wang S, Liang H, Bowden RL, Akhunov E (2017) Divergent and convergent modes of interaction between wheat and Puccinia graminis f. sp. tritici isolates revealed by the comparative gene co-expression network and genome analyses. BMC Genom 18:1–20
Saini DK, Chopra Y, Pal N, Chahal A, Srivastava P, Gupta PK (2021) Meta-QTLs, ortho-MQTLs and candidate genes for nitrogen use efficiency and root system architecture in bread wheat (Triticum aestivum L.). Physiol Mol Biol Plants 27:2245–2267
Saini DK, Chahal A, Pal N, Srivastava P, Gupta PK (2022a) Meta-analysis reveals consensus genomic regions associated with multiple disease resistance in wheat (Triticum aestivum L.). Mol Breed 42:11
Saini DK, Chopra Y, Singh J, Sandhu KS, Kumar A, Bazzer S, Srivastava P (2022b) Comprehensive evaluation of mapping complex traits in wheat using genome-wide association studies. Mol Breed 42:1–52
Saini DK, Srivastava P, Pal N, Gupta PK (2022c) Meta-QTLs, ortho-meta-QTLs and candidate genes for grain yield and associated traits in wheat (Triticum aestivum L). Theor Appl Genet 135(3):1049–1081
Salcedo A, Rutter W, Wang S, Akhunova A, Bolus S, Chao S, Anderson N, De Soto MF, Rouse M, Szabo L, Bowden RL (2017) Variation in the AvrSr35 gene determines Sr35 resistance against wheat stem rust race Ug99. Science 358:1604–1606
Sapkota S, Hao Y, Johnson J, Buck J, Aoun M, Mergoum M (2019) Genome-wide association study of a worldwide collection of wheat genotypes reveals novel quantitative trait loci for leaf rust resistance. Plant Genome 12:190033
Saremirad A, Bihamta M, Malihipour A, Mostafavi K, Alipour H (2020) Association mapping of Iranian wheat genotypes to detect QTLs conferring resistance to stem rust at seedling stage. https://doi.org/10.21203/rs.3.rs25893/v1
Schweizer P, Stein N (2011) Large-scale data integration reveals colocalization of gene functional groups with meta-QTL for multiple disease resistance in barley. Mol Plant Microbe Interact 24:1492–1501
Sharma A, Srivastava P, Mavi GS, Kaur S, Kaur J, Bala R, Sohu VS, Chhuneja P, Bains NS (2021) Resurrection of wheat cultivar ‘PBW343’using marker assisted gene pyramiding for rust resistance. Front Plant Sci 12:42
Shewabez E, Bekele E, Alemu A, Mugnai L, Tadesse W (2021) Genetic characterization and genome-wide association mapping for stem rust resistance in spring bread wheat. BMC Genomic Data. https://doi.org/10.21203/rs.3.rs-147919/v1
Singh S, Hernandez MV, Crossa J, Singh PK, Bains NS, Sngh K, Sharma I (2012) Multi-trait and multi-environment QTL analyses for resistance to wheat diseases. PLoS ONE 7:e38008
Singh A, Pandey MP, Singh AK, Knox RE, Ammar K, Clarke JM, Clarke FR, Singh RP, Pozniak CJ, DePauw RM, McCallum BD (2013) Identification and mapping of leaf, stem and Stripe rust resistance quantitative trait loci and their interactions in durum wheat. Mol Breed 31:405–418
Singh K, Batra R, Sharma S, Saripalli G, Gautam T, Singh R, Pal S, Malik P, Kumar M, Jan I, Singh S (2021) WheatQTLdb: a QTL database for wheat. Mol Genet Genom 296:1051–1056
Soko T, Bender CM, Prins R, Pretorius ZA (2018) Yield loss associated with different levels of stem rust resistance in bread wheat. Plant Dis 102:2531–2538
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Soriano JM, Royo C (2015) Dissecting the genetic architecture of leaf rust resistance in wheat by QTL MQTL analysis. Phytopathology 105:1585–1593. https://doi.org/10.1094/PHYTO-05-15-0130-R
Sosnowski O, Charcosset A, Joets J (2012) BioMercator V3: an upgrade of genetic map compilation and quantitative trait loci meta-analysis algorithms. Bioinformatics 28:2082–2083
Venske E, dos Santos RS, da Rosa FD, Rother V, Maia LC, Pegoraro C, Costa De Oliveira A (2019) MQTL analysis of the QTLome of Fusarium head blight resistance in bread wheat: refining the current puzzle. Front Plant Sci 10:727
Veyrieras JB, Goffinet B, Charcosset A (2007) Meta QTL: a package of new computational methods for the MQTL analysis of QTL mapping experiments. BMC Bioinformatics 8:49. https://doi.org/10.1186/1471-2105-8-49
Visscher PM, Goddard ME (2004) Prediction of the confidence interval of quantitative trait loci location. Behav Genet 34:477–482
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM (2014) Characterization of polyploid wheat genomic diversity using a high density 90000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Wang J, Li Z, Shi L, Zhu L, Ren Z, Li LD, Shah SJA (2015) QTL mapping for adult-plant leaf rust resistance genes in Chinese wheat cultivar Weimai 8. Czech J Genet Plant Breed 51:79–85
Wang H, Zou S, Li Y, Lin F, Tang D (2020) An ankyrin-repeat and WRKY-domain-containing immune receptor confers stripe rust resistance in wheat. Nat Commun 11:1–11
Wiesner-Hanks T, Nelson R (2016) Multiple disease resistance in plants. Annu Rev Phytopathol 54:229–252
Winfield MO, Allen AM, Burridge AJ, Barker GL, Benbow HR, Wilkinson PA, Coghill J, Waterfall C, Davassi A, Scopes G, Pirani A (2016) High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J 14:1195–1206
Xing L, Hu P, Liu J, Witek K, Zhou S, Xu J, Zhou W, Gao L, Huang Z, Zhang R, Wang X (2018) Pm21 from Haynaldia villosa encodes a CC-NBS-LRR protein conferring powdery mildew resistance in wheat. Mol Plant 11:874–878
Yadav IS, Sharma A, Kaur S, Nahar N, Bhardwaj SC, Sharma TR, Chhuneja P (2016) Comparative temporal transcriptome profiling of wheat near isogenic line carrying Lr57 under compatible and incompatible interactions. Front Plant Sci 7:1943
Yang Y, Amo A, Wei D, Chai Y, Zheng J, Qiao P, Cui C, Lu S, Chen L, Hu YG (2021) Large-scale integration of meta-QTL and genome-wide association study discovers the genomic regions and candidate genes for yield and yield-related traits in bread wheat. Theor Appl Genet 134:1–27
Yin JL, Fang ZW, Sun C, Zhang P, Zhang X, Lu C, Wang SP, Ma DF, Zhu YX (2018) Rapid identification of a stripe rust resistant gene in a space-induced wheat mutant using specific locus amplified fragment (SLAF) sequencing. Sci Rep 8:1–9
Yu LX, Morgounov A, Wanyera R, Keser M, Singh SK, Sorrells M (2012) Identification of Ug99 stem rust resistance loci in winter wheat germplasm using genome-wide association analysis. Theor Appl Genet 125:749–758
Zegeye H, Rasheed A, Makdis F, Badebo A, Ogbonnaya FC (2014) Genome-wide association mapping for seedling and adult plant resistance to stripe rust insynthetic hexaploid wheat. PloS one 9:e105593
Zhang H, Yang Y, Wang C, Liu M, Li H, Fu Y, Wang Y, Nie Y, Liu X, Ji W (2014) Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genom 15:1–14
Zhang P, Qi A, Zhou Y, Xia X, He Z, Li Z, Liu D (2016) Quantitative trait loci mapping of adult-plant resistance to leaf rust in a Fundulea 900בThatcher’wheat cross. Plant Breed 136:1–7
Zhang J, Zhang P, Dodds P, Lagudah E (2020) How target-sequence enrichment and sequencing (TEnSeq) pipelines have catalyzed resistance gene cloning in the wheat-rust pathosystem. Front Plant Sci 11:678
Zhang P, Yan X, Gebrewahid TW, Zhou Y, Yang E, Xia X, He Z, Li Z, Liu D (2021) Genome-wide association mapping of leaf rust and stripe rust resistance in wheat accessions using the 90K SNP array. Theor Appl Genet 134:1233–1251
Zwart RS, Bansal UK, Thompson JP, Williamson PM, Bariana HS (2008) QTL mapping of multiple disease resistance traits in a synthetic hexaploid x bread wheat population. In: Proceedings 11th international wheat genetics symposium, Brisbane, pp 1–3
Zwart RS, Thompson JP, Milgate AW, Bansal UK, Williamson PM, Raman H, Bariana HS (2010) QTL mapping of multiple foliar disease and root-lesion nematode resistances in wheat. Mol Breed 26:107–124
Acknowledgements
Thanks are due to Department of Biotechnology (DBT), Government of India, for financial support (BT/PR21024/AGIII/103/925/2016) and (BT/NABI-Flagship/2018). Thanks are also due to Indian National Science Academy (INSA), New Delhi, for the award of the positions of INSA-Senior Scientist and INSA Honorary Scientist to HSB.
Funding
No funding was received from any source for conducting the present study.
Author information
Authors and Affiliations
Contributions
PKG, HSB, PKS, and SK conceived and planned this study. NP, IJ, AK, and KK collected the literature and tabulated the QTL data for meta-QTL analysis. NP, IJ, DKS prepared the input files and performed QTL projection and meta-QTL analysis. NP, IJ and DKS interpreted the results and wrote the first draft of the manuscript. PKG, HSB, PKS, and SK edited and finalized the manuscript with the help of NP, IJ and DKS.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Communicated by Thomas Miedaner.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Fig. S1
Distribution of the markers on the consensus map used for meta-QTL analysis in the present study. Supplementary file2 (JPG 425 kb)
Rights and permissions
About this article
Cite this article
Pal, N., Jan, I., Saini, D.K. et al. Meta-QTLs for multiple disease resistance involving three rusts in common wheat (Triticum aestivum L.). Theor Appl Genet 135, 2385–2405 (2022). https://doi.org/10.1007/s00122-022-04119-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-022-04119-7