Abstract
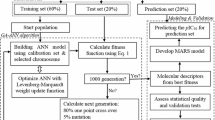
Tyrosine Kinase 2 (TYK2) inhibition is of potential therapeutic value for treating autoimmune diseases. An elaborate ligand-based computational workflow was employed to explore structural requirements for TYK2 inhibition. Genetic function algorithm (GFA) was coupled to k-nearest neighbor (kNN) and multiple linear regression (MLR) analyses to search for predictive QSAR models based on optimal pharmacophore(s)/physicochemical descriptors combinations. QSAR-selected pharmacophores were validated by receiver operating characteristic (ROC) curve analysis and by comparison with crystallographic structures of known inhibitors complexed within the TYK2 binding pocket. Optimal QSAR models and their associated pharmacophore hypotheses were used to identify new TYK2 inhibitory leads retrieved from the National Cancer Institute (NCI) structural database. The most potent hit exhibited experimental anti-TYK2 IC50 of 7.1 µM.















Similar content being viewed by others
References
Aboalhaija NH, Zihlif MA, Taha MO (2016) Discovery of new selective cytotoxic agents against Bcl-2 expressing cancer cells using ligand-based modeling. Chem-Biol Interact 250:12–26
Abuhammad A, Taha MO (2016) QSAR studies in the discovery of novel type-II diabetic therapies. Expert Opin Drug Discov 11:197–214
Abutayeh RF, Taha MO (2019) Discovery of novel Flt3 inhibitory chemotypes through extensive ligand-based and new structure-based pharmacophore modelling methods. J Mol Graph Model 88:128–151
Alabed SJ, Khanfar M, Taha MO (2016) Computer-aided discovery of new FGFR-1 inhibitors followed by in vitro validation. Future Med Chem 8:1841–1869
Al-Nadaf A, Sheikha GA, Taha MO (2010) Elaborate ligand-based pharmacophore exploration and QSAR analysis guide the synthesis of novel pyridinium-based potent β-secretase inhibitory leads. Bioorganic. Med Chem 18:3088–3115
Beeley N, Sage C (2003) GPCRs: an update on structural approaches to drug discovery. Targets 2:19–25
Bernstein CN, Wajda A, Blanchard JF (2005) The clustering of other chronic inflammatory diseases in inflammatory bowel disease: a population-based study. Gastroenterology 129:827–836
Blench TJ, Ellwood C, Goodacre SC, Lai Y, Liang J, Macleod C, Magnuson SR, Tsui VH, Williams K, Zhang B (2012) Azabenzothiazole compounds, compositions and methods of use, US Patent 2012/O2O2788, filled March 22, 2012, issued January 8, 2013
Blench T, Goodacre S, Lai Y, Liang Y, Macleod C, Magnuson S, Tsui V, Williams K, Zhang B (2012) Pyrazolopyridines and pyrazolopyridines and their use as TYK2 inhibitors. WO Patent Appl. WO2012/066061, filled May 24, 2012, issued May 19, 2013.
Brzozowski Z (1998) 2-Mercapto N-(azolyl)benzenesulfonamides. VI. Synthesis and anti-HIV activity of some new 2-mercapto-N-(1,2,4-triazol-3-yl)benzenesulfonamide derivatives containing the 1,2,4-triazole moiety fused with a variety of heteroaromatic rings. Acta Pol Pharm 55:473–480
Chrencik J, Patny A, Leung I, Korniski B, Emmons T, Hall T, Weinberg R, Gormley J, Williams J, Day J, Hirsch J, Kiefer J, Leone J, Fischer H, Sommers C, Huang H, Jacobsen E, Tenbrink R, Tomasselli A, Benson T (2010) Structural and thermodynamic characterization of the TYK2 and JAK3 kinase domains in complex with CP-690550 and CMP-6. J Mol Biol 400:413–433
Dassault Systèmes BIOVIA (2009) Discovery studio modeling environment, Release 2.5.5, San Diego, USA
De Mesquita MB, Ferrante M, Henckaerts L, Joossens M, Janssens V, Hlavaty T, Pierik M, Joossens S, Van Schuerbeek N, Van Assche G (2009) Clustering of (auto) immune diseases with early-onset and complicated inflammatory bowel disease. Eur J Pediatr 168:575–583
Depristo M (2004) Heterogeneity and inaccuracy in protein structures solved by X-ray crystallography. Structure 12:831–838
Dimova D, Stumpfe D, Hu Y, Bajorath J (2015) Activity cliff clusters as a source of structure–activity relationship information. Expert Opin Drug Discov 10:441–447
Duerr RH, Taylor KD, Brant SR, Rioux JD, Silverberg MS, Daly MJ, Steinhart AH, Abraham C, Regueiro M, Griffiths A (2006) A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314:1461–1463
Ellinghaus D, Ellinghaus E, Nair RP, Stuart PE, Esko T, Metspalu A, Debrus S, Raelson JV, Tejasvi T, Belouchi M (2012) Combined analysis of genome-wide association studies for Crohn disease and psoriasis identifies seven shared susceptibility loci. Am J Hum Genet 90:636–647
Fensome A, Ambler C, Arnold E, Banker M, Brown M, Chrencik J, Clark J, Dowty M, Efremov I, Flick A, Gerstenberger B, Gopalsamy A, Hayward M, Hegen M, Hollingshead B, Jussif J, Knafels J, Limburg D, Lin D, Lin T, Pierce B, Saiah E, Sharma R, Symanowicz P, Telliez J, Trujillo J, Vajdos F, Vincent F, Wan Z, Xing L, Yang X, Yang X, Zhang L (2018) Dual inhibition of TYK2 and JAK1 for the treatment of autoimmune diseases: discovery of ((S)-2,2-Difluorocyclopropyl)((1R,5S)-3-(2-((1-methyl-1H-pyrazol-4-yl)amino)pyrimidin-4-yl)-3,8-diazabicyclo[3.2.1]octan-8-yl)methanone (PF-06700841). J Med Chem 61:8597–8612
Fisher R (1971) The principle of experimentation illustrated by a psycho-physical experiment. The Design of Experiments. Hafner Publishing Company, New York
Frush E, Sekharan S, Keinan S (2017) In silico prediction of ligand binding energies in multiple therapeutic targets and diverse ligand sets—a case study on BACE1, TYK2, HSP90, and PERK proteins. J Phys Chem B 121:8142–8148
Furtmann N, Hu Y, Gütschow M, Bajorath J (2015) Identification of interaction hot spots in structures of drug targets on the basis of three-dimensional activity cliff information. Chem Biol Drug Des 86:1458–1465
Galatsis P, Henderson J, Kormos B, Han S, Kurumbail R, Wager T, Verhoest P, Noell G, Chen Y, Needle E, Berger Z, Steyn S, Houle C, Hirst W (2014) Kinase domain inhibition of leucine rich repeat kinase 2 (LRRK2) using a [1,2,4]triazolo[4,3-b]pyridazine scaffold. Bioorganic Med Chem Lett 24:4132–4140
Gaulton A, Hersey A, Nowotka M, Bento AP, Chambers J, Mendez D, Mutowo P, Atkinson F, Bellis LJ, Cibrián-Uhalte E, Davies M, Dedman N, Karlsson A, Magariños MP, Overington JP, Papadatos G, Smit I, Leach AR (2017) The ChEMBL database in 2017. Nucleic Acids Res 45:D945–D954
Gillett N, Chan C (2000) Applications of immunohistochemistry in the evaluation of immunosuppressive agents. Hum Exp Toxicol 19:251–254
Guha R, Medina-Franco JL (2014) On the validity versus utility of activity landscapes: are all activity cliffs statistically significant? J Cheminform 6:11
Hamaguchi H, Amano Y, Moritomo A, Shirakami S, Nakajima Y, Nakai K, Nomura N, Ito M, Higashi Y, Inoue T (2018) Discovery and structural characterization of peficitinib (ASP015K) as a novel and potent JAK inhibitor. Bioorganic Med Chem 26:4971–4983
Hernández-Flórez D, Valor L (2016) Protein-kinase inhibitors: a new treatment pathway for autoimmune and inflammatory diseases? Reumatol Clin 12:91–99
Homans SW (2007) Water, water everywhere—except where it matters? Drug Discov Today 12:534–539
Husby J, Bottegoni G, Kufareva I, Abagyan R, Cavalli A (2015) Structure-based predictions of activity cliffs. J Chem Inf Model 55:1062–1076
Joris L, Rague Schleyer P, Gleiter R (1968) Cyclopropane rings as proton-acceptor groups in hydrogen bonding. J Am Chem Soc 90:327–336
Kettle JG, Åstrand A, Catley M, Grimster NP, Nilsson M, Su Q, Woessner R (2017) Inhibitors of JAK-family kinases: an update on the patent literature 2013-2015, Part 1. Expert Opin Ther Pat 27:127–143
Khanfar MA, Taha MO (2013) Elaborate ligand-based modeling coupled with multiple linear regression and k nearest neighbor QSAR analyses unveiled new nanomolar mTOR inhibitors. J Chem Inf Model 53:2587–2612
Klebe G (2006) Virtual ligand screening: strategies, perspectives and limitations. Drug Discov Today 11:580–594
Kulagowski JJ, Blair W, Bull RJ, Chang C, Deshmukh G, Dyke HJ, Eigenbrot C, Ghilardi N, Gibbons P, Harrison TK (2012) Identification of imidazo-pyrrolopyridines as novel and potent JAK1 inhibitors. J Med Chem 55:5901–5921
Lees C, Barrett J, Parkes M, Satsangi J (2011) New IBD genetics: common pathways with other diseases. Gut 60:1739–1753
Lerner A, Jeremias P, Matthias T (2015) The world incidence and prevalence of autoimmune diseases is increasing. Int J Celiac Dis 3:151–155
Liang J, Tsui V, Van Abbema A, Bao L, Barrett K, Beresini M, Berezhkovskiy L, Blair WS, Chang C, Driscoll J, Eigenbrot C, Ghilardi N, Gibbons P, Halladay J, Johnson A, Kohli PB, Lai Y, Liimatta M, Mantik P, Menghrajani K, Murray J, Sambrone A, Xiao Y, Shia S, Shin Y, Smith J, Sohn S, Stanley M, Ultsch M, Zhang B, Wu LC, Magnuson S (2013a) Lead identification of novel and selective TYK2 inhibitors. Eur J Med Chem 67:175–187
Liang J, Van Abbema A, Balazs M, Barrett K, Berezhkovsky L, Blair W, Chang C, Delarosa D, Devoss J, Driscoll J (2013b) Lead optimization of a 4-aminopyridine benzamide scaffold to identify potent, selective, and orally bioavailable TYK2 inhibitors. J Med Chem 56:4521–4536
Liang J, Van Abbema A, Balazs M, Barrett K, Berezhkovsky L, Blair W, Chang C, Delarosa D, Devoss J, Driscoll J, Eigenbrot C, Goodacre S, Ghilardi N, Macleod C, Johnson A, Bir Kohli P, Lai Y, Lin Z, Mantik P, Menghrajani K, Nguyen H, Peng I, Sambrone A, Shia S, Smith J, Sohn S, Tsui V, Ultsch M, Williams K, Wu L, Yang W, Zhang B, Magnuson S (2017) Identification of an imidazopyridine scaffold to generate potent and selective TYK2 inhibitors that demonstrate activity in an in vivo psoriasis model. Bioorganic Med Chem Lett 27:4370–4376
Lipinski CA, Lombardo F, Dominy BW, Feeney PJ (2012) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 64:4–17
Lupardus P, Ultsch M, Wallweber H, Bir Kohli P, Johnson A, Eigenbrot C (2014) Structure of the pseudokinase-kinase domains from protein kinase TYK2 reveals a mechanism for Janus kinase (JAK) autoinhibition. Proc Natl Acad Sci USA 111:8025–8030
Ma H, Deacon S, Horiuchi K (2008) The challenge of selecting protein kinase assays for lead discovery optimization. Expert Opin Drug Discov 3:607–621
Ma H-L, Liang S, Li J, Napierata L, Brown T, Benoit S, Senices M, Gill D, Dunussi-Joannopoulos K, Collins M (2008) IL-22 is required for Th17 cell–mediated pathology in a mouse model of psoriasis-like skin inflammation. J Clin Invest 118:597–607
Matusevicius D, Kivisäkk P, He B, Kostulas N, Özenci V, Fredrikson S, Link H (1999) Interleukin-17 mRNA expression in blood and CSF mononuclear cells is augmented in multiple sclerosis. Mult Scler 5:101–104
Moslin R, Gardner D, Santella J, Zhang Y, Duncia J, Liu C, Lin J, Tokarski J, Strnad J, Pedicord D, Chen J, Blat Y, Zupa-Fernandez A, Cheng L, Sun H, Chaudhry C, Huang C, D'arienzo C, Sack J, Muckelbauer J, Chang C, Tredup J, Xie D, Aranibar N, Burke J, Carter P, Weinstein D (2017) Identification of imidazo[1,2-b]pyridazine TYK2 pseudokinase ligands as potent and selective allosteric inhibitors of TYK2 signalling. MedChemComm 8:700–712
Norman P (2012) Selective JAK1 inhibitor and selective TYK2 inhibitor patents. Expert Opin Ther Pat 22:1233–1249
Patterson H, Nibbs R, Mcinnes I, Siebert S (2014) Protein kinase inhibitors in the treatment of inflammatory and autoimmune diseases. Clin Exp Immunol 176:1–10
Roskoski R (2016) Janus kinase (JAK) inhibitors in the treatment of inflammatory and neoplastic diseases. Pharmacol Res 111:784–803
Shahin R, AlQtaishat S, Taha MO (2012) Elaborate ligand-based modeling reveal new submicromolar Rho kinase inhibitors. J Comput Aided Mol Des 26:249–266
Sigurdsson S, Nordmark G, Göring HH, Lindroos K, Wiman A-C, Sturfelt G, Jönsen A, Rantapää-Dahlqvist S, Möller B, Kere J (2005) Polymorphisms in the tyrosine kinase 2 and interferon regulatory factor 5 genes are associated with systemic lupus erythematosus. Am J Hum Genet 76:528–537
Sigurdsson S, Padyukov L, Kurreeman FA, Liljedahl U, Wiman AC, Alfredsson L, Toes R, Rönnelid J, Klareskog L, Huizinga TW (2007) Association of a haplotype in the promoter region of the interferon regulatory factor 5 gene with rheumatoid arthritis. Arthritis Rheum 56:2202–2210
Srivastava A, Nagai T, Srivastava A, Miyashita O, Tama F (2018) Role of COmputational Methods in Going Beyond X-ray Crystallography to Explore Protein Structure and Dynamics. Int J Mol Sci 19:3401
Stark GR, Kerr IM, Williams BR, Silverman RH, Schreiber RD (1998) How cells respond to interferons. Annu Rev Biochem 67:227–264
Steuber H, Zentgraf M, Gerlach C, Sotriffer C, Heine A, Klebe G (2006) Expect the unexpected or caveat for drug designers: multiple structure determinations using aldose reductase crystals treated under varying soaking and co-crystallisation conditions. J Mol Biol 363:174–187
Stubbs M, Reyda S, Dullweber F, Möller M, Klebe G, Dorsch D, Mederski W, Wurziger H (2002) pH-dependent binding modes observed in trypsin crystals: lessons for structure-based drug design. Chembiochem 3:246–249
Stumpfe D, Bajorath J (2012) Exploring activity cliffs in medicinal chemistry: miniperspective. J Med Chem 55:2932–2942
Stumpfe D, Dimova D, Bajorath J (2014) Composition and topology of activity cliff clusters formed by bioactive compounds. J Chem Inf Model 54:451–461
Taha MO (2012) Mixing pharmacophore modeling and classical QSAR analysis as powerful tool for lead discovery. In: Taha MO (ed) Virtual screening. InTech, Shanghai, China
Taha MO, Atallah N, Al-Bakri AG, Paradis-Bleau C, Zalloum H, Younis KS, Levesque RC (2008a) Discovery of new MurF inhibitors via pharmacophore modeling and QSAR analysis followed by in-silico screening. Bioorg Med Chem 16:1218–1235
Taha MO, Bustanji Y, Al-Ghussein MA, Mohammad M, Zalloum H, Al-Masri IM, Atallah N (2008b) Pharmacophore modeling, quantitative structure–activity relationship analysis, and in silico screening reveal potent glycogen synthase kinase-3β inhibitory activities for cimetidine, hydroxychloroquine, and gemifloxacin. J Med Chem 51:2062–2077
Taha MO, Dahabiyeh LA, Bustanji Y, Zalloum H, Saleh S (2008c) Combining ligand-based pharmacophore modeling, quantitative structure−activity relationship analysis and in silico screening for the discovery of new potent hormone sensitive lipase inhibitors. J Med Chem 51:6478–6494
Taha MO, Qandil AM, Al-Haraznah T, Khalaf RA, Zalloum H, Al-Bakri AG (2011) Discovery of New Antifungal Leads via Pharmacophore Modeling and QSAR analysis of fungal n-myristoyl transferase inhibitors followed by in silico screening. Chem Biol Drug Des 78:391–407
Thermo Fisher Scientific (2016) SelectScreen™ Biochemical Kinase Profiling Service Z′-LYTE™ Screening Protocol and Assay Conditions. 1–38
Tokarski J, Zupa-Fernandez A, Tredup J, Pike K, Chang C, Xie D, Cheng L, Pedicord D, Muckelbauer J, Johnson S, Wu S, Edavettal S, Hong Y, Witmer M, Elkin L, Blat Y, Pitts W, Weinstein D, Burke J (2015) Tyrosine kinase 2-mediated signal transduction in t lymphocytes is blocked by pharmacological stabilization of its pseudokinase domain. J Biol Chem 290:11061–11074
Triballeau N, Acher F, Brabet I, Pin J-P, Bertrand H-O (2005) Virtual screening workflow development guided by the “receiver operating characteristic” curve approach. Application to high-throughput docking on metabotropic glutamate receptor subtype 4. J Med Chem 48:2534–2547
Tsui V, Gibbons P, Ultsch M, Mortara K, Chang C, Blair W, Pulk R, Stanley M, Starovasnik M, Williams D (2011) A new regulatory switch in a JAK protein kinase. Proteins 79:393–401
Whitley D (1994) A genetic algorithm tutorial. Stat Comput 4:65–85
Yogo T, Nagamiya H, Seto M, Sasaki S, Shih-Chung H, Ohba Y, Tokunaga N, Lee G, Rhim C, Yoon C, Cho S, Skene R, Yamamoto S, Satou Y, Kuno M, Miyazaki T, Nakagawa H, Okabe A, Marui S, Aso K, Yoshida M (2016) Structure-based design and synthesis of 3-amino-1,5-dihydro-4h-pyrazolopyridin-4-one derivatives as tyrosine kinase 2 inhibitors. J Med Chem 59:733–749
Zheng W, Tropsha A (2000) Novel variable selection quantitative structure-property relationship approach based on the k-nearest-neighbor principle. J Chem Inf Comput Sci 40:185–194
Acknowledgements
The authors thank the Deanship of Scientific Research at the University of Jordan for funding this project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Al-Barghouthy, E.Y., Abuhammad, A. & Taha, M.O. QSAR-guided pharmacophore modeling and subsequent virtual screening identify novel TYK2 inhibitor. Med Chem Res 28, 1368–1387 (2019). https://doi.org/10.1007/s00044-019-02377-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-019-02377-7