Summary
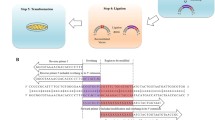
When plasmid pC194-1 is ligated to pBR322 to generate plasmid pHV15-1, deletions occur with high frequency within the joined pBR322 DNA. Generation of deletions is recE4 independent, and occurs in B. subtilis with a 1,000-fold higher frequency than in Escherichia coli. In the hybrid plasmid pVH15-1, deletion end-points are not at random, but at defined locations within pBR322. We propose that the base alteration, characterizing pC194-1, has stabilized within the plasmid a stem/loop structure, which acts as a deletion generator.
Similar content being viewed by others
References
Albertini AM, Hofer M, Calos MP, Miller JH (1982) On the formation of spontaneous deletions: The importance of short sequence homologies in the generation of large deletions. Cell 29:319–328
Alonso JC, Trautner TA (1985a) A gene controlling segregation of the Bacillus subtilis plasmid pC194. Mol Gen Genet 198:427–431
Alonso JC, Trautner TA (1985c) Cold sensitivity in the transfer of a plasmid with a deletion hot spot into recombination deficient B. subtilis cells. Mol Gen Genet 198:437–440
Campbell A (1962) Episomes. Adv Genet 11:101–145
Canosi U, Morelli G, Trautner TA (1978) The relationship between molecular structure and transformation efficiency of some S. aureus plasmids isolated from B. subtilis. Mol Gen Genet 166:259–267
Canosi U, Mazza G, Tailor R (1982) Identification of a mutation in a Bacillus subtilis plasmid affecting host sensitivity to methyl methanesulfonate. Genetic Exchange: A Celebration and a New Generation. Genetic and Cellular Technology, Vol. 1, Ed. UN Streips, SH Goodgal, WR Guild, GA Wilson, pp. 107–116, Marcel Dekker Inc., New York and Basel
Chang ACY, Cohen SN (1979) High frequency transformation of Bacillus subtilis protoplasts by plasmid DNA. Mol Gen Genet 168:111–115
Horinouchi S, Weisblum B (1982) Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance. J Bact 150:815–825
Ikeda H, Aoki K, Naito A (1982) Illegitimate recombination mediated in vitro by DNA gyrase of Escherichia coli: Structure of recombinant DNA molecules. Proc Natl Acad Sci USA 79:3724–3728
Jones IM, Primrose SB, Ehrlich SD (1982) Recombination between short direct repeats in a recA host. Mol Gen Genet 188:486–489
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular Cloning. A Laboratory Manual. Cold Spring Harb Laboratory
Marvo SL, King SR, Jasjunas SR (1983) The role of short regions of homology in intermolecular illegitimate recombinations. Proc Natl Acad Sci USA 80:2452–2456
Reif HJ, Saedler H (1975) IS1 is involved in deletion formation in the gal region of E. coli K12. Mol Gen Genet 137:17–28
Streisinger G, Okada Y, Emrich J, Newton J, Tsugita A, Terzaghi E, Inouye M (1966) Frameshift mutations and the genetic code. Cold Spring Harb Symp Quant Biol 31:77–84
Sutcliffe JG (1979) Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol 43:300–307
Vieira J, Messing J (1982) The pUC plasmid, an M13mp7 derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19:259–268
Author information
Authors and Affiliations
Additional information
Communicated by H. Saedler
Rights and permissions
About this article
Cite this article
Alonso, J.C., Trautner, T.A. Generation of deletions through a cis-acting mutation in plasmid pC194. Mol Gen Genet 198, 432–436 (1985). https://doi.org/10.1007/BF00332935
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00332935