Abstract
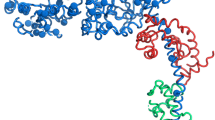
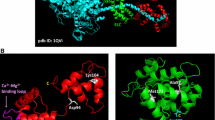
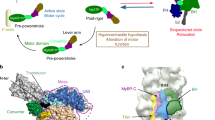
Mutations in the gene coding for cardiac myosin binding protein-C (cMyBP-C), a multi-domain (C0-C10) protein, are a major causative factor for inherited hypertrophic cardiomyopathy. Patients carrying mutations in this gene have an extremely heterogeneous clinical course, with some progressing to end-stage heart failure. The cause of this variability is unknown. We here describe molecular modeling of a double mutation in domains C1 (E258K) and C2 (E441K) in a patient with severe HCM phenotype. The three-dimensional structure for the C1-motif-C2 complex was constructed with double and single mutations being introduced. Molecular dynamic simulations were performed for 10 ns under physiological conditions. The results showed that both E258K and E441K in isolation can predominantly affect the native domain as well as the nearby motif via conformational changes and result in an additive effect when they coexist. These changes involve important regions of the motif such as phosphorylation and potential actin-binding sites. Moreover, the charge reversal mutations altered the surface electrostatic properties of the complex. In addition, we studied protein expression, which showed that the mutant proteins were expressed and we can suppose that the severe phenotype was not due to haploinsufficiency. However, additional studies on human gene expression will need to confirm this hypothesis. The double mutation affecting the regulatory N-terminal of cMyBP-C have the potential of synergistically interfering with the binding to neighbouring domains and other sarcomeric proteins. These effects may account for the severe phenotype and clinical course observed in the complex cMyBP-C genotypes.







Similar content being viewed by others
References
Richard, P., Charron, P., Carrier, L., Ledeuil, C., Cheav, T., Pichereau, C., Benaiche, A., Isnard, R., Dubourg, O., Burban, M., Gueffet, J. P., Millaire, A., Desnos, M., Schwartz, K., Hainque, B., & Komajda, M. (2003). EUROGENE Heart Failure Project. Hypertrophic cardiomyopathy: distribution of disease genes, spectrum of mutations, and implications for a molecular diagnosis strategy. Circulation, 107, 2227–2232. doi:10.1161/01.CIR.0000066323.15244.54.
Maron, B. J., Gardin, J. M., Flack, J. M., Gidding, S. S., Kurosaki, T. T., & Bild, D. E. (1995). Prevalence of hypertrophic cardiomyopathy in a general population of young adults. Echocardiographic analysis of 4111 subjects in the CARDIA study. Coronary artery risk development in (Young) adults. Circulation, 92, 785–789. doi:10.1161/01.CIR.92.4.785.
Maron, B. J., Shirani, J., Poliac, L. C., Mathenge, R., Roberts, W. C., & Mueller, F. O. (1996). Sudden death in young competitive athletes. Clinical, demographic, and pathological profiles. JAMA, 276, 199–204. doi:10.1001/jama.1996.03540030033028.
Cecchi, F., Yacoub, M. H., & Olivotto, I. (2005). Hypertrophic cardiomyopathy in the community: why we should care. Nature Clinical Practice Cardiovascular Medicine, 2, 324–325. doi:10.1038/ncpcardio0248.
Olivotto, I., Cecchi, F., Poggesi, C., & Yacoub, M. H. (2012). Patterns of disease progression in hypertrophic cardiomyopathy: an individualized approach to clinical staging. Circulation Heart Failure, 5, 535–546. doi:10.1161/CIRCHEARTFAILURE.112.967026.
Yacoub, M. H., Olivotto, I., & Cecchi, F. (2007). ‘End–stage’ hypertrophic cardiomyopathy: from mystery to model. Nature Clinical Practice Cardiovascular Medicine, 4, 232–233. doi:10.1038/ncpcardio0859.
Frey, N., Luedde, M., & Katus, H. A. (2011). Mechanisms of disease: hypertrophic cardiomyopathy. Nature Reviews Cardiology, 9, 91–100. doi:10.1038/nrcardio.2011.159.
Kelly, M., & Semsarian, C. (2009). Multiple mutations in genetic cardiovascular disease: a marker of disease severity? Circulation Cardiovascular Genetics, 2, 182–190. doi:10.1161/CIRCGENETICS.108.836478.
De Lange, W. J., Grimes, A. C., Hegge, L. F., Spring, A. M., Brost, T. M., & Ralphe, J. C. (2013). E258K HCM-causing mutation in cardiac MyBP-C reduces contractile force and accelerates twitch kinetics by disrupting the cMyBP-C and myosin S2 interaction. Journal of General Physiology, 142, 241–255. doi:10.1085/jgp.201311018.
Olivotto, I., Girolami, F., Ackerman, M. J., Nistri, S., Bos, J. M., Zachara, E., Ommen, S. R., Theis, J. L., Vaubel, R. A., Re, F., Armentano, C., Poggesi, C., Torricelli, F., & Cecchi, F. (2008). Myofilament protein gene mutation screening and outcome of patients with hypertrophic cardiomyopathy. Mayo Clinic Proceedings, 83, 630–638. doi:10.4065/83.6.630.
Mearini, G., Stimpel, D., Krämer, E., Geertz, B., Braren, I., Gedicke-Hornung, C., Précigout, G., Müller, O. J., Katus, H. A., Eschenhagen, T., Voit, T., Garcia, L., Lorain, S., & Carrier, L. (2013). Repair of Mybpc3 mRNA by 5′-trans-splicing in a mouse model of hypertrophic cardiomyopathy. Molecular Therapy Nucleic Acids, 2, e102. doi:10.1038/mtna.2013.31.
Merk, S. (2005). Disease-causing mutations in the human cardiac myosin binding protein C gene. Genomics of cardiovascular development, adaptation, and remodeling. NHLBI Program for Genomic Applications, Harvard Medical School. Available at: http://www.cardiogenomics.org [June 2010].
Marsiglia, J. D., Batitucci Mdo, C., Fd, P., Barbirato, C., Arteaga, E., & Araújo, A. Q. (2010). Study of mutations causing hypertrophic cardiomyopathy in a group of patients from Espirito Santo, Brazil. Arquivos Brasileiros de Cardiologia, 94, 10–17.
Einheber, S., & Fischman, D. A. (1990). Isolation and characterization of a cDNA clone encoding avian skeletal muscle C-protein: an intracellular member of the immunoglobulin superfamily. Proceedings of the National Academy of Sciences of the United States of America, 87, 2157–2161. doi:10.1073/pnas.87.6.2157.
Howarth, J. W., Ramisetti, S., Nolan, K., Sadayappan, S., & Rosevear, P. R. (2012). Structural insight into unique cardiac myosin-binding protein-C motif: a partially folded domain. Journal of Biological Chemistry, 287, 8254–8262. doi:10.1074/jbc.M111.309591.
Oakley, C. E., Chamoun, J., Brown, L. J., & Hambly, B. D. (2007). Myosin binding protein-C: enigmatic regulator of cardiac contraction. International Journal of Biochemistry and Cell Biology, 39, 2161–2166. doi:10.1016/j.biocel.2006.12.008.
Govada, L., Carpenter, L., da Fonseca, P. C., Helliwell, J. R., Rizkallah, P., Flashman, E., Chayen, N. E., Redwood, C., & Squire, J. M. (2008). Crystal structure of the C1 domain of cardiac myosin binding protein-C: implications for hypertrophic cardiomyopathy. Journal of Molecular Biology, 378, 387–397. doi:10.1016/j.jmb.2008.02.044.
Ababou, A., Gautel, M., & Pfuhl, M. (2007). Dissecting the N-terminal myosin binding site of human cardiac myosin-binding protein C. Structure and myosin binding of domain C2. Journal of Biological Chemistry, 282, 9204–9215. doi:10.1074/jbc.M610899200.
Roy, A., Kucukural, A., & Zhang, Y. (2010). I-TASSER: a unified platform for automated protein structure and function prediction. Nature Protocols, 5, 725–738. doi:10.1038/nprot.2010.5.
Comeau, S. R., Gatchell, D. W., Vajda, S., & Camacho, C. J. (2004). ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics, 20, 45–50. doi:10.1093/bioinformatics/btg371.
Discovery Studio. (2012). 3.5 User Guide. Accelrys Inc., San Diego, CA, USA.
Krishnamoorthy, N., Yacoub, M. H., & Yaliraki, S. N. (2011). A computational modeling approach for enhancing self-assembly and biofunctionalisation of collagen biomimetic peptides. Biomaterials, 32, 7275–7285. doi:10.1016/j.biomaterials.2011.06.074.
Van Der Spoel, D., Lindahl, E., Hess, B., Groenhof, G., Mark, A. E., & Berendsen, H. J. (2005). GROMACS: fast, flexible, and free. Journal of Computational Chemistry, 26, 1701–1718. doi:10.1002/jcc.20291.
Hess, B., Kutzner, C., van der Spoel, D., & Lindahl, E. (2008). GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. Journal of Chemical Theory and Computation, 4, 435–447. doi:10.1021/ct700301q.
Berendsen, H. J. C, Postma J. P. M, van Gunsteren W. F, Hermans J. (1981). Interaction models for water in relation to protein hydration. In: Intermolecular forces, edited by Pullman B, D. Reidel. Publishing Company. pp 331–342.
van Gunsteren, W. F., Billeter, S. R., Eising, A. A., Hünenberger, P. H., Krüger, P., Mark, A. E., Scott, W. R. P., & Tironi, I. G. (1996). Biomolecular simulation: the GROMOS96 manual and user guide (pp. 1–1042). Zürich: Vdf Hochschulverlag AG an der ETH Zürich.
Hess, B., Bekker, H., Berendsen, H. J. C., & Fraaije, J. G. E. M. (1997). LINCS: a linear constraint solver for molecular simulations. Journal of Computational Chemistry, 18, 1463–1472. doi:10.1002/(SICI)1096-987X(199709)18.
Miyamoto, S., & Kollman, P. A. (1992). SETTLE: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. Journal of Computational Chemistry, 13, 952–962. doi:10.1002/jcc.540130805.
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F., DiNola, A., DiNola, A., & Haak, J. R. (1984). Molecular dynamics with coupling to an external bath. Journal of Chemical Physics, 81, 3684–3691.
Wallace, A. C., Laskowski, R. A., & Thornton, J. M. (1995). LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Engineering, 8, 127–134. doi:10.1093/protein/8.2.127.
Sinha, N., & Smith-Gill, S. J. (2002). Electrostatics in protein binding and function. Current Protein and Peptide Science, 3, 601–614. doi:10.2174/1389203023380431.
Krishnamoorthy, N., Gajendrarao, P., Eom, S. H., Kwon, Y. J., Cheong, G. W., & Lee, K. W. (2008). Molecular modeling study of CodX reveals importance of N-terminal and C-terminal domain in the CodWX complex structure of Bacillus subtilis. Journal of Molecular Graphics and Modelling, 27, 1–12. doi:10.1016/j.jmgm.2008.01.009.
Helms, A. S., Davis, F. M., Coleman, D., Bartolone, S. N., Glazier, A. A., Pagani, F., Yob, J. M., Sadayappan, S., Pedersen, E., Lyons, R., Westfall, M. V., Jones, R., Russell, M. W., & Day, S. M. (2014). Sarcomere mutation-specific expression patterns in human hypertrophic cardiomyopathy. Circulation Cardiovascular Genetics, 7(4), 434–443.
Gautel, M., Zuffardi, O., Freiburg, A., & Labeit, S. (1995). Phosphorylation switches specific for the cardiac isoform of myosin binding protein-C: a modulator of cardiac contraction? EMBO Journal, 14, 1952–1960.
Bahrudin, U., Morisaki, H., Morisaki, T., Ninomiya, H., Higaki, K., Nanba, E., Igawa, O., Takashima, S., Mizuta, E., Miake, J., Yamamoto, Y., Shirayoshi, Y., Kitakaze, M., Carrier, L., & Hisatome, I. (2008). Ubiquitin-proteasome system impairment caused by a missense cardiac myosin-binding protein C mutation and associated with cardiac dysfunction in hypertrophic cardiomyopathy. Journal of Molecular Biology, 384(4), 896–907. doi:10.1016/j.jmb.2008.09.070.
Marston, S., Copeland, O., Jacques, A., Livesey, K., Tsang, V., McKenna, W. J., Jalilzadeh, S., Carballo, S., Redwood, C., & Watkins, H. (2009). Evidence from human myectomy samples that MYBPC3 mutations cause hypertrophic cardiomyopathy through haploinsufficiency. Circulation Research, 105, 219–222. doi:10.1161/CIRCRESAHA.109.202440.
Marston, S., Copeland, O., Gehmlich, K., Schlossarek, S., & Carrier, L. (2012). How do MYBPC3 mutations cause hypertrophic cardiomyopathy? Journal of Muscle Research and Cell Motility, 33, 75–80. doi:10.1007/s10974-011-9268-3.
van Dijk, S. J., Dooijes, D., dos Remedios, C., Michels, M., Lamers, J. M., Winegrad, S., Schlossarek, S., Carrier, L., ten Cate, F. J., Stienen, G. J., & van der Velden, J. (2009). Cardiac myosin-binding protein C mutations and hypertrophic cardiomyopathy: haploinsufficiency, deranged phosphorylation, and cardiomyocyte dysfunction. Circulation, 119, 1473–1483. doi:10.1161/CIRCULATIONAHA.108.838672.
Sadayappan, S., Gulick, J., Osinska, H., Martin, L. A., Hahn, H. S., Dorn, G. W., 2nd, Klevitsky, R., Seidman, C. E., Seidman, J. G., & Robbins, J. (2005). Cardiac myosin binding protein-C phosphorylation and cardiac function. Circulation Research, 97, 1156–1163. doi:10.1161/01.RES.0000190605.79013.4d.
Tong, C. W., Stelzer, J. E., Greaser, M. L., Powers, P. A., & Moss, R. L. (2008). Acceleration of crossbridge kinetics by protein kinase A phosphorylation of cardiac myosin binding protein C modulates cardiac function. Circulation Research, 103, 974–982. doi:10.1161/CIRCRESAHA.108.177683.
Sadayappan, S., Gulick, J., Osinska, H., Barefield, D., Cuello, F., Avkiran, M., Lasko, V. M., Lorenz, J. N., Maillet, M., Martin, J. L., Brown, J. H., Bers, D. M., Molkentin, J. D., James, J., & Robbins, J. (2011). A critical function for Ser-282 in cardiac Myosin binding protein-C phosphorylation and cardiac function. Circulation Research, 109, 141–150. doi:10.1161/CIRCRESAHA.111.242560.
Moos, C., Mason, C. M., Besterman, J. M., Feng, I. N., & Dubin, J. H. (1978). The binding of skeletal muscle C-protein to F-actin, and its relation to the interaction of actin with myosin subfragment-1. Journal of Molecular Biology, 124, 571–586.
Squire, J. M., Luther, P. K., & Knupp, C. (2003). Structural evidence for the interaction of C-protein (MyBP-C) with actin and sequence identification of a possible actin-binding domain. Journal of Molecular Biology, 331, 713–724.
Razumova, M. V., Shaffer, J. F., Tu, A. Y., Flint, G. V., Regnier, M., & Harris, S. P. (2006). Effects of the N-terminal domains of myosin binding protein-C in an in vitro motility assay: evidence for long-lived cross-bridges. Journal of Biological Chemistry, 281, 35846–35854. doi:10.1074/jbc.M606949200.
Bhuiyan, M. S., Gulick, J., Osinska, H., Gupta, M., & Robbins, J. (2012). Determination of the critical residues responsible for cardiac myosin binding protein C’s interactions. Journal of Molecular and Cellular Cardiology, 53, 838–847. doi:10.1016/j.yjmcc.2012.08.028.
Craig, R., Lee, K. H., Mun, J. Y., Torre, I., & Luther, P. K. (2014). Structure, sarcomeric organization, and thin filament binding of cardiac myosin-binding protein-C. Pflügers Archiv, 466, 425–431. doi:10.1007/s00424-013-1426-6.
Luther, P. K., Winkler, H., Taylor, K., Zoghbi, M. E., Craig, R., Padrón, R., Squire, J. M., & Liu, J. (2011). Direct visualization of myosin-binding protein C bridging myosin and actin filaments in intact muscle. Proceedings of the National Academy of Sciences of the United States of America, 108, 11423–11428. doi:10.1073/pnas.1103216108.
Acknowledgments
We would like to thank Dr. Othmane Bouhali, Director of the Research Computing and Mr. Faisal Chaudhry, Senior Lead Systems Engineer, Texas A&M University in Qatar for providing supercomputing facility. We also extend our thanks to Dr. Brian P. Mitchelson, QCRC and Mr. Mark Radford for their suggestions. No human or animal studies were carried out by the authors of this article.
Funding
This work was supported by Qatar Foundation through Qatar Cardiovascular Research Center, Doha, Qatar, and it was also supported by Magdi Yacoub Research Network, London, UK. IO and FG are supported by the Italian Ministry of Health (RF 2010 – 2313451 “Hypertrophic cardiomyopathy: new insights from deep sequencing and psychosocial evaluation”) and NET-2011-02347173 (Mechanisms and treatment of coronary microvascular dysfunction in patients with genetic or secondary left ventricular hypertrophy). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of Interest
None declared.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Daniel P. Judge oversaw the review of this article
Poornima Gajendrarao and Navaneethakrishnan Krishnamoorthy contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Figure 1
The effect of MYBPC3 variants on protein stability in H9C2 cells. H9C2 cells were either transfected with wild type (WT) or MYBPC3 variants or untransfected (control). After 36h of transfection, cells were lysed and immunoblotted with MYBPC3 antibody (DOCX 263 kb)
Rights and permissions
About this article
Cite this article
Gajendrarao, P., Krishnamoorthy, N., Selvaraj, S. et al. An Investigation of the Molecular Mechanism of Double cMyBP-C Mutation in a Patient with End-Stage Hypertrophic Cardiomyopathy. J. of Cardiovasc. Trans. Res. 8, 232–243 (2015). https://doi.org/10.1007/s12265-015-9624-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-015-9624-6