Abstract
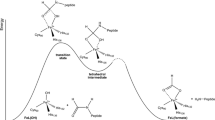
Crystal structures of polypeptide deformylase (PDF) of Escherichia coli with nickel(II) replacing the native iron(II) have been solved with chloride and formate as metal ligands. The chloro complex is a model for the correct protonation state of the hydrolytic hydroxo ligand and the protonated status of the Glu133 side chain as part of the hydrolytic mechanism. The ambiguity that recently some PDFs have been identified with Zn2+ ion as the active-site centre whereas others are only active with Fe2+ (or Co2+, Ni2+) is discussed with respect to Lewis acid criteria of the metal ion and substrate activation by the CD loop.



Similar content being viewed by others
Abbreviations
- HEPES:
-
N-(2-Hydroxyethyl)piperazine-N′-ethanesulfonic acid
- PDF:
-
Peptide deformylase
- RMSD:
-
Root mean square deviation
References
Adams JM, Capecchi MR (1966) Proc Natl Acad Sci USA 55:147–155
Meinnel T, Blanquet S (1993) J Bacteriol 175:7737–7740
Adams JM (1968) J Mol Biol 33:571–589
Livingston DM, Leder P (1969) Biochemistry 8:435–443
Mazel D, Pochet S, Marliere P (1994) EMBO J 13:914–923
Groche D, Becker A, Schlichting I, Kabsch W, Schultz S, Wagner AF (1998) Biochem Biophys Res Commun 246:342–346
Margolis P, Hackbarth C, Lopez S, Maniar M, Wang W, Yuan Z, White R, Trias J (2001) Antimicrob Agents Chemother 45:2432–2435
Meinnel T, Blanquet S (1994) J Bacteriol 176:7387–7390
Apfel CM, Locher H, Evers S, Takacs B, Hubschwerlen C, Pirson W, Page MG, Keck W (2001) Antimicrob Agents Chemother 45:1058–1064
Apfel C, Banner DW, Bur D, Dietz M, Hirata T, Hubschwerlen C, Locher H, Page MG, Pirson W, Rossé G, Specklin JL (2000) J Med Chem 43:2324–2331
Chen DZ, Patel DV, Hackbarth CJ, Wang W, Dreyer G, Young DC, Margolis PS, Wu C, Ni ZJ, Trias J, White RJ, Yuan Z (2000) Biochemistry 39:1256–1262
Clements JM, Beckett RP, Brown A, Catlin G, Lobell M, Palan S, Thomas W, Whittaker M, Wood S, Salama S, Baker PJ, Rodgers HF, Barynin V, Rice DW, Hunter MG (2001) Antimicrob Agents Chemother 45:563–570
Huntington KM, Yi T, Wei Y, Pei D (2000) Biochemistry 39:4543–4551
Azoulay-Dupuis E, Mohler J, Bedos JP (2004) Antimicrob Agents Chemother 48:80–85
Fieulaine S, Juillan-Binard C, Serero A, Dardel F, Giglione C, Meinnel T, Ferrer JL (2005) J Biol Chem 280:42315–42324
Gross M, Clements J, Beckett RP, Thomas W, Taylor S, Lofland D, Ramanathan-Girish S, Garcia M, Difuntorum S, Hoch U, Chen H, Johnson KW (2004) J Antimicrob Chemother 53:487–493
Ramanathan-Girish S, McColm J, Clements JM, Taupin P, Barrowcliffe S, Hevizi J, Safrin S, Moore C, Patou G, Moser H, Gadd A, Hoch U, Jiang V, Lofland D, Johnson KW (2004) Antimicrob Agents Chemother 48:4835–4842
Watters AA, Jones RN, Leeds JA, Denys G, Sader HS, Fritsche TR (2006) J Antimicrob Chemother 57:914–923
Becker A, Schlichting I, Kabsch W, Groche D, Schultz S, Wagner AF (1998) Nat Struct Biol 5:1053–1058
Becker A, Schlichting I, Kabsch W, Schultz S, Wagner AF (1998) J Biol Chem 273:11413–11416
Jain R, Hao B, Liu RP, Chan MK (2005) J Am Chem Soc 127:4558–4559
Smith KJ, Petit CM, Aubart K, Smyth M, McManus E, Jones J, Fosberry A, Lewis C, Lonetto M, Christensen SB (2003) Protein Sci 12:349–360
Leopoldini M, Russo N, Toscano M (2006) J Phys Chem B 110:1063–1072
Madison V, Duca J, Bennett F, Bohanon S, Cooper A, Chu M, Desai J, Girijavallabhan V, Hare R, Hruza A, Hendrata S, Huang Y, Kravec C, Malcolm B, McCormick J, Miesel L, Ramanathan L, Reichert P, Saksena A, Wang J, Weber PC, Zhu H, Fischmann T (2002) Biophys Chem 101–102:239–247
Li Y, Ren S, Gong W (2002) Acta Crystallogr D Biol Crystallogr 58:846–848
Zhou Z, Song X, Li Y, Gong W (2004) J Mol Biol 339:207–215
Deng H, Callender R, Zhu J, Nguyen KT, Pei D (2002) Biochemistry 41:10563–10569
Wu XH, Quan JM, Wu YD (2007) J Phys Chem B 111:6236–6244
Xiao C, Zhang Y (2007) J Phys Chem B 111:6229–6235
Lazennec C, Meinnel T (1997) Anal Biochem 244:180–182
Pflugrath JW (1999) Acta Crystallogr D Biol Crystallogr 55:1718–1725
McCoy AJ (2007) Acta Crystallogr D Biol Crystallogr 63:32–41
Murshudov GN, Vagin AA, Dodson EJ (1997) Acta Crystallogr D Biol Crystallogr 53:240–255
Winn MD, Murshudov GN, Papiz MZ (2003) Meth Enzymol 374:300–321
Collaborative Computational Project Number 4 (1994) Acta Crystallogr D Biol Crystallogr 50:760–763
Emsley P, Cowtan K (2004) Acta Crystallogr D Biol Crystallogr 60:2126–2132
DeLano WL (2002) The PyMOL molecular graphics system. DeLano Scientific, San Carlos
Chan MK, Gong W, Rajagopalan PT, Hao B, Tsai CM, Pei D (1997) Biochemistry 36:13904–13909
Ragusa S, Blanquet S, Meinnel T (1998) J Mol Biol 280:515–523
Meinnel T, Blanquet S (1995) J Bacteriol 177:1883–1887
Palm GJ, Lederer T, Orth P, Saenger W, Takahashi M, Hillen W, Hinrichs W (2008) J Biol Inorg Chem 13:1097–1110
Rajagopalan PT, Grimme S, Pei D (2000) Biochemistry 39:779–790
Pearson RG (1963) J Am Chem Soc 85:3533–3539
Pearson RG (1966) Science 151:172–177
Parr RG, Pearson RG (1983) J Am Chem Soc 105:7512–7516
Corpet F (1988) Nucleic Acids Res 16:10881–10890
Gouet P, Courcelle E, Stuart DI, Metoz F (1999) Bioinformatics 15:305–308
Nguyen KT, Wu JC, Boylan JA, Gherardini FC, Pei D (2007) Arch Biochem Biophys 468(2):217–225
Acknowledgments
We thank Thomas Schweder (University of Greifswald) and his group for help in cloning E. coli PDF. Y.T.H.N. is grateful for a personal grant from the MOET, Vietnam, and financial support by DAAD, Germany.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yen, N.T.H., Bogdanović, X., Palm, G.J. et al. Structure of the Ni(II) complex of Escherichia coli peptide deformylase and suggestions on deformylase activities depending on different metal(II) centres. J Biol Inorg Chem 15, 195–201 (2010). https://doi.org/10.1007/s00775-009-0583-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-009-0583-8