Abstract
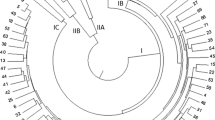
In recent years, Pongamia has been considered as important renewable source of biodiesel, however not much molecular information is available in this species. Molecular characterization of this legume tree will enhance our understanding in improving the optimal yields of oil through breeding and enable us to meet the future demands for biodiesel. To assess the molecular genetic diversity in 46 Pongamia pinnata accessions collected from six different states of India, amplified fragment length polymorphism (AFLP) marker system was employed. Five AFLP primer combinations produced 520 discernible fragments, of which 502 (96.5%) were polymorphic. AFLP primer informativeness was estimated evaluating four parameters namely polymorphism information content (PIC), effective multiplex ratio (EMR), marker index (MI) and resolving power (RP). In total, 51 unique fragments were detected of which 19 unique fragments were observed with primer combination E-ACG / M-CTA. Although neighbour joining (NJ) method did not group accessions strictly according to their region of collection, a good level of genetic diversity was observed in examined germplasm. However, accessions collected from Karnataka showed comparatively higher diversity than accessions from other states. The diverse accessions identified in this study may be useful in Pongamia pinnata improvement to meet the future demands of biodiesel.
Similar content being viewed by others
Abbreviations
- AFLP:
-
Amplified fragment length polymorphism
- EMR:
-
Effective multiplex ratio
- FAME:
-
Fatty acid methyl esters
- MI:
-
Marker index
- PIC:
-
Polymorphism information content
- RP:
-
Resolving power
- SVO:
-
Straight vegetable oil
References
Wani SP, Osman M, Emmanuel D'Silva & Sreedevi TK, Asian Biotechnol Deve Rev, 6 (2006) 11.
Scott PT, Pregelj L, Chen N & Gresshoff PM, Bioenergy Research, 1 (2008) 2.
Atchison E, Am J Bot, 36 (1951) 538.
Sarbhoy RK, Cytologia, 42 (1977) 415.
Kumar V, Chandrashekar K & Sidhu OP, J Entomol Res, 30 (2006) 103.
Khurma UR & Mangotra A, S Pac J Nat Sci, 22 (2004) 50.
Yadav YS, Siddiqui AU & Parihar A, J Phytol Res, 16 (2005) 263.
Elanchezhiyan M, Rajarajan S, Rajendran P, Subramanian S & Thyagarajan SP, J Mad Microbiol, 36 (1993) 262.
D'sllva E, Wani SP & Nagnath B, Patancheru 502 324, Global theme on Agroecosystems, Report No. 11 (2004) 28.
Azam MM, Waris A & Nahar NM, Biomass and Bioenergy, 29 (2005) 293.
Karmee SJ & Chadha A, Biores Technol, 96 (2005) 1425.
Nagaraj G & Mulda N, J Oilseeds Res, 21 (2004) 117.
Natanam R, Kadirvel R & Chandrasekaran D, Animal Feed Sci Technol, 25 (1989) 201.
Rathore V & Madras G, Fuel, 66 (2007) 2650.
Bena G, Jubler MF, Olivieri I & Lejeune B, J Mot Evol, 46 (1998) 299.
Parani M, Lakshmi M, Kumar SP & Parida A, Curr Sci, 76 (2000) 1235.
Casiva PV, Saidman BO, Vilardi VC & Cialdella AM, Am J Bot, 69 (2002) 843.
Vos P, Hogers R, Bleaker M, Reijans M, Lee T, Homes M, Friters A, Pot J, Paleman J, Kulper M & Zabeau M, Nucleic Acids Res, 23 (1995) 4407.
Tatikonda L, Wani SP, Kannan S, Beerelli N, Holsington DA, Devi P & Varshney RK, Plant Sci, 176 (2009) 505.
Roldan-Rulz J, Dendauw E, VanBockstaele A, Depicker M & Loose De, Mot Breed, 6 (2000) 125.
Varshney RK, Chabane K, Hendre PS & Aggarwal RK, Plant Sci, 73 (2007) 638.
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S & Rafalski A, Mot Breed, 2 (1996) 225.
Prevost A & Wilkinson MJ, Theor Appl Genet, 96 (1999) 107.
Nei M, American Naturalist, 106 (1972) 283.
Swofford DL, Phylogenetic analysis using Parsimony (*and Other Methods) Version 4, Sinauer Associates, Sunderland, Massachusetts, (2003).
Hudson DH, Richter DC, Rausch C, Dezulian T, Franz M & Rupp R, BMC Biointormatics, 6 (2007) 460.
Milbourne D, Meyer R, Bradshaw J, Baird E, Boner N, Proven J, Powell W & Waugh R, Mot Breed, 3 (1997) 127.
Russell JR, Fuller JD, Macaulay M, Hatz BG, Jahoor A, Powell W & Waugh R, Theor Appl Genet, 95 (1997) 714.
Sun Q-B, Li L-F, Li Y, Wu G-J & Ge X-J, Crop Sci, 46 (2008) 1865.
Fu Yong-Bi, Gugel RK & Katepa-Mupondwa F, Plant Gen Res, 4 (2006) 87.
Rosales-Serna R, Herna'ndez-Delgado S, Gonza’ lez-Paz M, Acosta-Gallegos JA & Mayek-Pe'rez N, Crop Sci, 45 (2005) 1951.
Fernandez M, Figueiras A & Benito C, Theor Appl Genet, 104 (2002) 845.
Laurentin H & Karlovsky P, Genet Resour Crop Evol, 54 (2007) 1437.
Gupta S, Srivastava M, Mishra G.P, Naik PK, Chauhan RS, Tiwari SK, Kumar M & Singh R, Atr J Biotechnol, 7 (2008) 4230.
Bohn M & Utz H, Crop Sci, 39 (1999) 228.
Srivastava A, Gupta V, Pental D & Pradhan AK, Theor Appl Genet, 102 (2001) 193.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Thudi, M., Manthena, R., Wani, S.P. et al. Analysis of Genetic Diversity in Pongamia [Pongamia pinnata (L) Pierrre] using AFLP Markers. J. Plant Biochem. Biotechnol. 19, 209–216 (2010). https://doi.org/10.1007/BF03263342
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF03263342