Summary
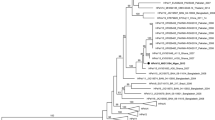
Comparisons of genome and polyprotein sequences of hepatitis C virus (HCV) isolates world-wide has led to the identification of nine major genotypes and many subtypes. This classification is based on either complete genome/polyprotein sequences or sequence data from the 5′ noncoding region, core, E1, NS3 or NS5B genes. The relative merit of different gene segments as taxonomic markers and the validity of the resulting assignments is not clear at this stage. To resolve the taxonomy of HCV genotypes and subtypes, we have compared the complete genome and polyprotein sequences of 19 HCV isolates available in the databases as well as sequences of individual genes and gene products of these isolates. Based on the correlation between sequence relationships and taxonomic assignments of other RNA viruses, we show that the nine major genotypes of HCV represent nine distinct virus species and their subtypes subspecies. Our sequence comparison of the 5′ noncoding regions and the individual gene products suggests that E2, NS2, NS5B, E1, NS4A, NS4B and NS5A (in that order) are the most appropriate regions for the discrimination between species, subspecies and strains of HCV. The 5′ noncoding, core and NS3 regions are less effective in distinguishing between species, subspecies and strains. Based on a comparison of the polymerase sequence identities of HCVs, pestiviruses and flaviviruses as well as the recent information on the size and morphology of HCV virions, we propose that HCVs, pestiviruses and flaviviruses should be classified into three separate families, namedHepciviridae, Pestiviridae andFlaviviridae, respectively rather than three genera of theFlaviviridae as currently classified. We also propose “Hepcivirus” as the genus name for HCVs.
Similar content being viewed by others
References
Baker JC (1987) Bovine viral diarrhoea virus: a review. J Am Vet Med Assoc 190: 1449–1458
Brechot C, Kremsdorf D (1993) Genetic variation of hepatitis C virus (HCV) genome: random events or a clinically relevant issue? J Hepatol 17: 285–268
Buch J, Purcell RH, Miller RH (1992) Sequence analysis of the 5′ noncoding region of hepatitis C virus. Proc Natl Acad Sci USA 89: 4942–4946
Buch J, Purcell RH, Miller RH (1993) At least 12 genotypes of hepatitis C virus predicted by sequence analysis of the putative E1 gene of isolates collected worldwide. Proc Natl Acad Sci USA 90: 8234–8238
Buch J, Purcell RH, Miller RH (1994) Sequence analysis of the core gene of 14 hepatitis C virus genotypes. Proc Natl Acad Sci USA 91: 8239–8243
Cha T-A, Beall E, Irvine B, Kolberg J, Chien D, Kuo G, Ureda MS (1992) At least five related, but distinct, hepatitis C viral genotypes exist. Proc Natl Acad Sci USA 89: 7144–7148
Chambers TJ, Han CS, Galler R, Rice CM (1990) Flavivirus genome organisation, expression and replication. Annu Rev Microbiol 44: 649–688
Chan S-W, McOmish F, Holmes EC, Dow B, Peutherer JF, Follett E, Yap PL, Simmonds P (1992) Analysis of a new hepatitis C virus type and its phylogenetic relationship to existing variants. J Gen Virol 73: 1131–1141
Choo Q-L, Richman KH, Han JH, Berger K, Lee C, Dong C, Galleogos C, Coit D, Medina Selby R, Barr PJ, Weiner AJ, Branley DW, Kuo G, Houghton M (1991) Genetic organisation and diversity of hepatitis C virus. Proc Natl Acad Sci USA 88: 2451–2455
Collett MS (1992) Molecular genetics of pestiviruses. Comp Immunol Microbiol Infect Dis 15: 145–154
Collett MS, Anderson DK, Retzel E (1988) Comparisons of the pestivirus bovine viral diarrhoea virus with members ofFlaviviridae. J Gen Virol 69: 2637–2643
Dahle J, Liess B (1992) A review on classical swine fever infection in pigs: epizootiology, clinical disease and pathology. Comp Immunol Microbiol Infect Dis 15: 203–211
Devereux J, Haeberli P, Mithies O (1984) A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res 12: 387–395
Dolja VV, Carrington JC (1992) Evolution of positive strand RNA viruses. Semin Virol 3: 315–326
Dusheiko G, Alberti A, Columbo M, Craxi A, Prieto J, Sanchez-Tapias J, Simmonds P (1994) The influence of sėrotypes of hepatitis C virus on sustained response to alpha interferon treatment. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego, p 224
Dusheiko G, Schmilovitz-Weiss H, Brown D, McOmish F, Yap P-L, Sherlock S, McIntyre N, Simmonds P (1994) Hepatitis C virus genotypes: an investigation of type-specific differences in geographical origin and disease. Hepatology 19: 13–18
Enomoto N, Takada A, Nakao T, Date T (1990) There are two major types of hepatitis C virus in Japan. Biochem Biophys Res Commun 170: 1021–1025
Francki RIB, Fauquet CM, Knudson DL, Brown F (1991) Classification and Nomenclature of Viruses. Fifth Report of the International Committee on Taxonomy of Viruses. Springer, Wien New York (Arch Virol [Suppl] 2)
Fu J, Tan B-H, Yap E-H, Chan Y-C, Tan YH (1992) Full-length cDNA sequence of dengue type 1 virus (Singapore strain S275/90). Virology 188: 953–958
Goldbach R, Wellink J (1988) Evolution of plus strand RNA viruses. Intervirology 29: 260–268
Gorman BM (1984) Application of evolutionary species concepts to viruses. Abstract, Sixth International Congress of Virology, Sendai, p 388
Habili N, Symons RH (1989) Evolutionary relationship between luteoviruses and other RNA plant viruses based on sequence motifs in their putative RNA polymerases and nucleic acid helicases. Nucleic Acids Res 17: 9543–9555
Han JH, Shyamala V, Richman KH, Brauer MJ, Irvine B, Ureda MS, Tekamp-Olson P, Kuo G, Choo Q-L, Houghton M (1991) Characterisation of the terminal regions of hepatitis C virus viral RNA: identification of conserved sequences in the 5′ untranslated region and poly (A) tails at the 3′ end. Proc Natl Acad Sci USA 88: 1711–1715
Hijikata M, Kato N, Ootsuyama Y, Nakagawa M, Ohkoshi S, Shimotohno K (1991) Hypervariable regions in the putative glycoprotein of hepatitis C virus. Biochem Biophys Res Commun 175: 220–228
Hijikata M, Mizushima H, Tanji Y, Komoda Y, Hirowatari Y, Akagi T, Kato N, Kimura K, Shimotohno K (1993) Proteolytic processing and membrane association of putative nonstructural proteins of hepatitis C virus. Proc Natl Acad Sci USA 90: 10773–10777
Houghton M, Richman K, Han J, Berger K, Lee C, Dong C, Overby L, Weiner A, Bradley D, Kuo G, Choo Q-L (1991) Hepatitis C virus (HCV): a relative of the pestiviruses and flaviviruses. In: Hollinger EB, Lemon SM, Margolis HS (eds) Viral hepatitis and liver disease. Williams & Wilkins, Baltimore, pp 328–333
Kaito M, Watanabe S, Tsukiyama-Kohara K, Yamaguchi K, Kobayashi Y, Konishi M, Yokoi M, Ishida S, Suzuki S, Kohara M (1994) Hepatitis C virus particle detected by immunoelectron microscopic study. J Gen Virol 75: 1755–1760
Kanai K, Kako M, Okamoto H (1992) HCV genotypes in chronic hepatitis C and response to interferon. Lancet 339: 1543
Kato M, Hijikata M, Nakagawa M, Oostuyama Y, Muraiso K, Ohkoshi S, Shimotohno K (1991) Molecular structure of the Japanese hepatitis C virus genome. FEBS Lett 280: 325–328
Kingsbury DW (1985) Species classification problems in virus taxonomy. Intervirology 24: 62–70
Koonin EV (1991) The phylogeny of RNA-dependent RNA polymerases of positive-strand RNA viruses. J Gen Virol 72: 2197–2206
Koonin EV, Dolja VV (1993) Evolution and taxonomy of positive-strand RNA viruses: implications of comparative analysis of amino acid sequences. Crit Rev Biochem Mol Biol 28: 375–430
Li XM, Jeffers LJ, Shao LJ, DeMedina M, Reddy KR, Schiff ER (1994) Hepatitis C virus (HCV) characteristic ofTogaviridae family. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego, p 24
Lin C, Lindenbach BD, Pragai BM, McCourt DW, Rice CM (1994) Processing of the hepatitis C virus E2-NS2 region: identification of p7 and two distinct E2-specific products with different C termini. J Virol 68: 5063–5073
Mandl CW, Heinz FX, Stockl E, Kunz C (1989) Genome sequence of tick-borne encephalitis virus (Western subtype) and comparative analysis of nonstructural proteins with other flaviviruses. Virology 173: 291–301
Martell M, Esteban JI, Quer J, Genesca J, Weiner A, Esteban R, Guarda J, Gomez J (1992) Hepatitis C virus (HCV) circulates as a population of different but closely related genomes: quasispecies nature of HCV genome distribution. J Virol 66: 3225–3229
McOmish F, Yap PL, Dow BC, Follett EAC, Seed C, Keller AJ, Cobain TJ, Krusius T, Kolho E, Naukkarinen R, Lin C, Lai C, Leong S, Medgyesi GA, Hejjas M, Kiyokawa H, Fukuda K, Cuypers T, Saeed AA, Al-Rasheed AM, Lin M, Simmonds P (1994) Geographical distribution of hepatitis C virus genotypes in blood donors: an international collaborative study. J Clin Microbiol 32: 884–892
Matsuura Y, Miyamura T (1993) The molecular biology of hepatitis C virus. Semin Virol 4: 297–304
Mellor J, Simmonds P, Yap PL, Holmes EC (1994) Extraordinary diversity of HCV variants within genotypes 3 and 4: Relationships to routes of transmission. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego, p 161
Miller RH, Purcell RH (1990) Hepatitis C virus shares amino acid sequence similarity with pestiviruses and flaviviruses as well as members of two plant virus super groups. Proc Natl Acad Sci USA 87: 2057–2061
Monath TP (1986) Pathology of flaviviruses. In: Schlesinger S, Schlesinger MJ (eds) The togaviridae and flaviviridae. Plenum Press, New York, pp 375–440
Mori S, Kato N, Yagyu A, Tanaka T, Ikeda Y, Petchclai B, Chiewsilp P, Kurimura T, Shimotohno K (1992) A new type of hepatitis C virus in patients in Thailand. Biochem Biophys Res Commun 183: 334–342
Okamoto H, Okada S, Sugiyama Y, Kurai K, Iizuka H, Machida A, Miyakawa Y, Mayumi M (1991) Nucleotide sequence of the genomic RNA of hepatitis C virus isolated from a human carrier: comparison with reported isolates for conserved and divergent regions. J Gen Virol 72: 2697–2704
Okamoto H, Kurai K, Okada S, Yamamoto K, Iizuka H, Tanaka T, Fukuda S, Tsuda F, Mishiro S (1992) Full length sequence of hepatitis C virus genome having poor homology to reported isolates: comparative study of four distinct genotypes. Virology 188: 331–341
Okamoto H, Sugiyama Y, Okada S, Kurai K, Akahane Y, Sugai Y, Tanaka T, Sato K, Tsuda F, Miyakawa Y, Mayumi M (1992) Typing hepatitis C virus by polymerase chain reaction with type-specific primers: application to clinical surveys and tracing infection sources. J Gen Virol 73: 673–679
Pontisso P, Ruvoletto MG, Gerotto M, Nicoletti M, Tisminetzky S, Baralle F, Alberti A (1994) A multicentric study on distribution and clinical features of HCV genotypes in chronic hepatitis C in Italy. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego, p 163
Pozzato G, Moretti M, Franzin F, Croce LS, Tiribelli C, Masayu T, Kaneko S, Unoura M, Kobayashi K (1991) Severity of liver disease with different hepatitis C virus clones. Lancet 338: 509
Prescott LE, Yap PL, Pike I, Parker D, Simmonds P (1994) A serological assay to detect six major HCV genotypes. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego, p 264
Prince AM, Parker T, Levine D, Huima T, Kimball LF (1994) Use of HCV association with VLDL visualisation of HCV and putative defective interfering particles. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego
Rybicki E (1990) The classification of organisms at the edge of life or problems with virus systematics. S Afr J Sci 86: 182–186
Shukla DD, Hoyne P, Ward CW (1994) A revised taxonomy of hepatitis C virus. Abstract, Second International Meeting on Hepatitis C and Related Viruses, San Diego, p 167
Shukla DD, Ward CW, Brunt AA (1994) The potyviridae. CAB International, Wallingford
Simmonds P, Holmes EC, Cha T-A, Chan S-W, McOmish F, Irvine B, Beall E, Yap PL, Kolberg J, Ureda MS (1993) Classification of hepatitis C virus into six major genotypes and a series of subtypes by phylogenetic analysis of the NS5 region. J Gen Virol 74: 2391–2399
Simmonds P, McOmish F, Yap PL, Chan S-W, Lin CK, Dusheiko G, Saeed AA, Holmes EC (1993) Sequence variability in the 5′ noncoding region of hepatitis C virus: identification of a new virus type and restrictions on sequence diversity. J Gen Virol 74: 661–668
Simmonds P, Alberti A, Alter HJ, Bonino F, Bradley DW, Brechot C, Brouwer JT, Chan SW, Chayama K, Chen DS, Choo QL, Colombo M, Cuypers HTM, Date T, Dusheiko GM, Esteban JI, Fay O, Hadziyannis SJ, Han J, Hatzakis A, Holmes EC, Hotta H, Houghton M, Irvine B, Kohara M, Kolberg JA, Kuo G, Lau JYN, Lelie PN, Maertens G, McOmish F, Miyamura T, Mizokami M, Nomoto A, Prince AM, Reesink HW, Rice C, Roggendorf M, Schalm SW, Shikata T, Shimotohno K, Stuyver L, Trepo C, Weiner A, Yap PL, Ureda MS (1994) A proposed system for the nomenclature of hepatitis C viral genotypes. Hepatology 19: 1321–1324
Simmonds P, Smith DB, McOmish F, Yap PL, Kolberg J, Ureda MS, Holmes EC (1994) Identification of genotypes of hepatitis C virus by sequence comparisons in the core, E1 and NS-5 regions. J Gen Virol 75: 1053–1061
Strauss JH, Preugschat F, Strauss EG (1991) Structure and function of the flavivirus and pestivirus genomes. In: Hollinger EB, Lemon SM, Margolis HS (eds) Viral hepatitis and liver disease. Williams & Wilkins, Baltimore, pp 333–344
Suzich JA, Tamura JK, Palmer-Hill F, Warrener P, Grakoui A, Rice CM, Feinstone SM, Collett MS (1993) Hepatitis C virus NS3 protein polynucleotide-stimulated nucleotide triphosphatase and comparison with the related pestivirus and flavivirus enzymes. J Virol 67: 6152–6158
Takada N, Takase S, Enomoto N, Takada A, Date T (1992) Clinical backgrounds of the patients having different types of hepatitis C virus genomes. J Hepatol 14: 35–40
Takamizawa A, Mori C, Fuke I, Manabe S, Murakami S, Fujita J, Onishi E, Andoh T, Yoshida I, Okayama H (1991) Structure and organisation of the hepatitis C virus genome isolated from human carriers. J Virol 65: 1105–1113
Tokita H, Okamoto H, Tsuda F, Song P, Nakata S, Chosa T, Iizuka H, Mishiro S, Miyakawa Y, Mayumi M (1994) Hepatitis C virus variants from Vietnam are classifiable into the seventh, eighth and ninth major genetic groups. Proc Natl Acad Sci USA 91: 11022–11026
Van Regenmortel MHV (1989) Applying the species concept to plant viruses. Arch Virol 104: 1–17
Ward CW (1993) Progress towards a higher taxonomy of viruses. Res Virol 144: 419–453
Weiner AJ, Brauer MJ, Rosenblatt J, Richman KH, Tung J, Crawford K, Bonino F, Saracco G, Choo Q-L, Houghton M, Han JH (1991) Variable and hypervariable domains are found in the regions of HCV corresponding to the flavivirus envelope and NS1 proteins and the pestivirus envelope glycoproteins. Virology 180: 842–848
Westaway EG, Brinton MA, Gaidamovich SY, Horzinek MC, Igarashi A, Kaariainen L, Lvov DK, Porterfield JS, Russell PK, Trent DW (1985) Flaviviridae. Intervirology 24: 183–192
Yoshioka K, Kakumu S, Wakita T, Ishikawa T, Itoh Y, Takayanagi M, Higashi Y, Shibata M, Morishima T (1992) Detection of hepatitis C virus by polymerase chain reaction and response to interferon-alpha therapy: relationship to genotypes of hepatitis C virus. Hepatology 16: 293–299
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Shukla, D.D., Hoyne, P.A. & Ward, C.W. Evaluation of complete genome sequences and sequences of individual gene products for the classification of hepatitis C viruses. Archives of Virology 140, 1747–1761 (1995). https://doi.org/10.1007/BF01384339
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01384339