Summary
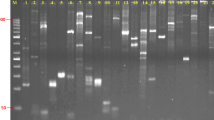
We have used random amplified polymorphic DNA (RAPD) analysis to characterize eleven cultivars of the five economically most important yam species grown in Jamaica (Dioscorea alata, D. cayenensis, D. rotundata, D. trifida and D. esculenta). Amplification of genomic DNA samples with nine different arbitrary 10mer primers revealed a total of 338 different band positions, ranging in size from 0.3 to 2.5 kb. RAPD patterns proved to be highly reproducible and somatically stable. While no variation was observed among plants belonging to the same cultivar, a large number of intervarietal and interspecific polymorphisms enabled us to reliably discriminate between all Jamaican cultivars investigated.
Similar content being viewed by others
References
Adams, R.P. & T., Demeke, 1993. Systematic relationships in Juniperus based on random amplified polymorphic DNAs (RAPDs). Taxon 42: 553–571.
Asemota, H.N., 1995. A fast, simple, and efficient miniscale method for the preparation of DNA from tissues of yam (Dioscorea spp.). Plant Mol Biol Rep 13: 214–218.
Asemota, H.N., M., Wellington, A.A., Odutuga & M.H., Ahmad, 1992. Effect of short-term storage on phenolic content, odiphenolase and peroxidase activitess of cut yam tubers (Dioscorea spp.). J Sci Food Agric 60: 309–312.
Bailey, D.C., 1983. Isozymic variation and plant breeders' rights. In: S.D., Tanksley & J.T., Orton (Eds.), Isoenzymes in Plant Genetics and Breeding, part A, pp. 425–441. Elsevier, Amsterdam.
Castiglione, S., G., Wang, G., Damiani, C., Bandi, S., Bisoffi & F., Sala, 1993. RAPD fingerprints for identification and for taxonomic studies of elite poplar (Populus spec.) clone. Theor Appl Genet 87: 54–59.
Dellaporta, S.L., J., Wood & J.B., Hicks, 1983. A plant DNA minipreparation: Version II. Plant Mol Biol Rep 1: 9–21.
Demeke, T., R.P., Adams & R., Chibbar, 1992. Potential use of random amplified polymorphic DNA (RAPD): A case study with Brassica. Theor Appl Genet 84: 990–994.
Ellsworth, D.L., K.D., Rittenhouse & R.L., Honeycutt, 1993. Artifactual variation in randomly amplified polymorphic DNA banding patterns. Bio Techniques 14: 214–217.
Felsenstein, J., 1985. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39: 783–791.
Hamon, P. & B., Touré, 1990a. Characterization of traditional yam varieties belonging to the Dioscorea cayenensis-rotundata complex by their isoenzymic patterns. Euphytica 46: 101–107.
Hamon, P. & B., Touré, 1990b. The classification of the cultivated yams (Dioscorea cayenensis-rotundata complex) of West Africa. Euphytica 47: 179–187.
Hoepfner, A.S., H., Nybom, U., Carlson & R., Franzen, 1993. DNA fingerprinting useful for monitoring cell line identity in micropropagated raspberries. Acta Agric Scand Sect B, Soil and Plant Sci 43: 53–57.
Hu, J. & C.F., Quiros, 1991. Identification of broccoli and cauliflower cultivars with RAPD markers. Plant Cell Rep 10: 505–511.
Ikediobi, C.O. & L.C., Igboanusi, 1983. Identification of yam, Dioscorea spp. cultivars by use of electrophoretic patterns of soluble tuber proteins. Biotropica 15: 65–67.
Iqbal, M.J. & A.L., Rayburn, 1994. Stability of RAPD markers for determining cultivar specific DNA profiles in rye (Secale cereale L.). Euphytica 75: 215–220.
Isabel, N., J., Beaulieu & J., Bousquet, 1995. Complete congruence between gene diversity estimates derived from genotypic data at enzyme and random amplified polymorphic DNA loci in black spruce. Proc Natl Acad Sci USA 92: 6369–6373.
Kaemmer, D., R., Afza, K., Weising, G., Kahl & F.J., Novak, 1992. Oligonucleotide and amplification fingerprinting of wild species and cultivars of banana (Musa spp.). Bio/Technology 10: 1030–1035.
Koller, B., B., Lehmann, J.M., McDermott & C., Gessler, 1993. Identification of apple cultivars using RAPD markers. Theor Appl Genet 85: 901–904.
Lanham, P.G., R.M., Brenna, C., Hackett & R.J., McNicol, 1995. RAPD fingerprinting of blackcurrant (Ribes nigrum L.) cultivars. Theor Appl Genet 90: 166–172.
Mailer, R.J., R., Scarth & B., Fristensky, 1994. Discrimination among cultivars of rapeseed (Brassica napus L.) using DNA polymorphisms amplified from arbitrary primers. Theor Appl Genet 87: 697–704.
Martin, F.W. & A.M., Rhodes, 1978. The relationship of Dioscorea cayenensis and D. rotundata. Trop Agric (Trinidad) 55: 193–206.
Muralidharan, K. & E.K., Wakeland, 1993. Concentration of primer and template qualitatively affects products in random-amplified polymorphic DNA PCR. Bio Techniques 14: 362–264.
Muzac-Tucker, I., H.N., Asemota & M.H., Ahmad, 1993. Biochemical composition and storage of Jamaican yams (Dioscorea spp.). J Sci Food Agric 62: 219–224.
Nei, M. & W.H., Li, 1979. Mathematical model for studying genetic variation in terms of restriction endonuclease.. Proc Natl Acad Sci USA 76: 5269–5273.
Novy, R.G. & N., Vorsa, 1995. Identification of intracultivar genetic heterogeneity in cranberry using silver-stained RAPDs. HortScience 30: 600–604.
Novy, R.G., C., Kobak, J., Goffreda & N., Vorsa, 1994. RAPDs identify varietal misclassification and regional divergence in cranberry (Vaccinium macrocarpon (Ait.) Pursh). Theor Appl Genet 88: 1004–1010.
Ramser, J., C., Lopez-Peralta, R., Wetzel, K., Weising & G., Kahl, 1996. Genomic variation and relationships in aerial yam (Dioscorea bulbifera L.) detected by random amplified polymorphic DNA. Genome 39: 17–25.
Retter, R.S., J.G.K., Williams, K.A., Feldmann, J.A., Rafalski, S.V., Tingey & P.A., Scolnik, 1992. Global and local genome mapping in Arabidopsis thaliana by using recombinant inbred lines and random amplified polymorphic DNA. Proc Natl Acad Sci USA 89: 1477–1481.
Saitou, N. & M., Nei, 1987. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4: 406–425.
Sambrook, J., E.F., Fritsch & T., Maniatis, 1989. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
Van de, Peer, Y. & R.De, Wachter, 1993. TREECON: A software package for the construction and drawing of evolutionary trees. Comput Applic Biosci 9: 177–182.
Virk, P.S., H.J., Newbury, M.T., Jackson & B.V., Ford-Lloyd, 1995a. The identification of duplicate accessions within a rice germplasm collection using RAPD analysis. theor Appl Genet 90: 1049–1055.
Virk, P.S., B.V., Ford-Lloyd, M.T., Jackson & H.J., Newbury, 1995b. Use of RAPD for the study of diversity within plant germplasm collections. Heredity 74: 170–179.
Wetton, J.H., R.E., Carter, D.T., Parkin & D., Walters, 1987. Demographic study of a wild house sparrow population by DNA fingerprinting. Nature (London) 327: 147–149.
Williams, J.G.K., A.R., Kubelik, K.J., Livak, J.A., Rafalski & S.V., Tingey, 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleid Acids Res 18: 6531–6535.
Wolff, K., E., Zietkiewicz & H., Hofstra, 1995. Identification of chrysanthemum cultivars and stability of DNA fingerprint patterns. Theor Appl Genet 91: 439–447.
Yang, X. & C., Quiros, 1993. Identification and classification of celery with RAPD markers. Theor Appl Genet 86: 205–212.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Asemota, H.N., Ramser, J., Lopéz-Peralta, C. et al. Genetic variation and cultivar identification of Jamaican yam germplasm by random amplified polymorphic DNA analysis. Euphytica 92, 341–351 (1995). https://doi.org/10.1007/BF00037118
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00037118