Abstract
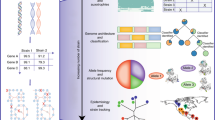
Genome-Scale metabolic models (GEMs) are a relevant tool in systems biology for in silico strain optimisation and drug discovery. An easier way to reconstruct a model is to use available GEMs as templates to create the initial draft, which can be curated up until a simulation-ready model is obtained. This approach is implemented in merlin’s BiGG Integration Tool, which reconstructs models from existing GEMs present in the BiGG Models database. This study aims to assess draft models generated using models from BiGG as templates for three distinct organisms, namely, Streptococcus thermophilus, Xylella fastidiosa and Mycobacterium tuberculosis. Several draft models were reconstructed using the BiGG Integration Tool and different templates (all, selected and random). The variability of the models was assessed using the reactions and metabolic functions associated with the model’s genes. This analysis showed that, even though the models shared a significant portion of reactions and metabolic functions, models from different organisms are still differentiated. Moreover, there also seems to be variability among the templates used to generate the draft models to a lower extent. This study concluded that the BiGG Integration Tool provides a fast and reliable alternative for draft reconstruction for bacteria.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Thiele, I., Palsson, B.Ø.: A protocol for generating a high-quality genome-scale metabolic reconstruction. Nat. Protoc. 5, 93–121 (2010)
Feist, A.M., Herrgård, M.J., Thiele, I., Reed, J.L., Palsson, B.: Reconstruction of biochemical networks in microorganisms. Nat. Rev. Microbiol. 7, 129–143 (2009)
Orth, J.D., Thiele, I., Palsson, B.Ø.: What is flux balance? Nat. Biotechnol. 28, 245–248 (2010)
O’Brien, E.J., Monk, J.M., Palsson, B.O.: Using genome-scale models to predict biological capabilities. Cell 161, 971–987 (2015)
Zhang, C., Hua, Q.: Applications of genome-scale metabolic models in biotechnology and systems medicine. Front. Physiol. 6, 1–8 (2016)
Norsigian, C.J., et al.: BiGG Models 2020: multi-strain genome-scale models and expansion across the phylogenetic tree. Nucleic Acids Res. 48, D402–D406 (2020)
Machado, D., Andrejev, S., Tramontano, M., Patil, K.R.: Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 46, 7542–7553 (2018)
Dias, O., Rocha, M., Ferreira, E.C., Rocha, I.: Reconstructing genome-scale metabolic models with merlin. Nucleic Acids Res. 43, 3899–3910 (2015)
Capela, J., et al..: merlin v4.0: an updated platform for the reconstruction of high-quality genome-scale metabolic models. bioRxiv (2021)
Sequeira, J.C., Rocha, M., Alves, M.M., Salvador, A.F.: UPIMAPI, reCOGnizer and KEGGCharter: three tools for functionalannotation. In: BOD 2021 - X Bioinformatics Open Days. Braga, Portugal, vol. 57 (2021)
Galperin, M.Y., Kristensen, D.M., Makarova, K.S., Wolf, Y.I., Koonin, E.V.: Microbial genome analysis: The COG approach. Brief. Bioinform. 20, 1063–1070 (2019)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Oliveira, A. et al. (2022). Towards a Multivariate Analysis of Genome-Scale Metabolic Models Derived from the BiGG Models Database. In: Rocha, M., Fdez-Riverola, F., Mohamad, M.S., Casado-Vara, R. (eds) Practical Applications of Computational Biology & Bioinformatics, 15th International Conference (PACBB 2021). PACBB 2021. Lecture Notes in Networks and Systems, vol 325. Springer, Cham. https://doi.org/10.1007/978-3-030-86258-9_14
Download citation
DOI: https://doi.org/10.1007/978-3-030-86258-9_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-86257-2
Online ISBN: 978-3-030-86258-9
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)