Abstract
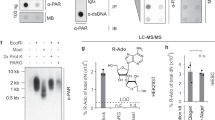
We describe a method for analyzing multiple products of PARylation by PARP1 and/or PARP2 using high-pressure liquid chromatography. The method quantitates the small molecules NAD+ (the substrate), nicotinamide (the byproduct of PARylation or hydrolysis of NAD+), and ADPR, the product of NAD+ hydrolysis. The method also quantitates the products of PARylation following digestion of the PAR chains into “ends,” “middles,” and “branches.” This method is useful for dissecting both the activity and the partitioning of PARylation products between different outcomes (i.e., long chains vs. short chains, PARylation vs. hydrolysis).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Gibson BA, Kraus WL (2012) New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol 13:411–424
Bai P (2015) Biology of poly(ADP-ribose) polymerases: the factotums of cell maintenance. Mol Cell 58:947–958
Langelier MF, Eisemann T, Riccio AA et al (2018) PARP family enzymes: regulation and catalysis of the poly(ADP-ribose) posttranslational modification. Curr Opin Struct Biol 53:187–198
Suskiewicz MJ, Palazzo L, Hughes R et al (2020) Progress and outlook in studying the substrate specificities of PARPs and related enzymes. FEBS J:1–12
Gibbs-Seymour I, Fontana P, Rack JGM et al (2016) HPF1/C4orf27 is a PARP-1-interacting protein that regulates PARP-1 ADP-ribosylation activity. Mol Cell 62:432–442
Bonfiglio JJ, Fontana P, Zhang Q et al (2017) Serine ADP-ribosylation depends on HPF1. Mol Cell 65:932–940.e6
Palazzo L, Leidecker O, Prokhorova E et al (2018) Serine is the major residue for ADP- ribosylation upon DNA damage. Elife 7:e34334
Rudolph J, Roberts G, Muthurajan UM et al (2020) HPF1 and nucleosomes mediate a dramatic switch in activity of PARP1 from polymerase to hydrolase. Elife 10:e65773
Suskiewicz MJ, Zobel F, Ogden TEH et al (2020) HPF1 completes the PARP active site for DNA damage-induced ADP-ribosylation. Nature 579:598–602
Sun F, Zhao P, Zhang N et al (2021) HPF1 remodels the active site of PARP1 to enable the serine ADP-ribosylation of histones. Nat Commun 12:1028
Dasovich M, Beckett MQ, Bailey S, et al (2021) Identifying poly(ADP-ribose)-binding proteins with photoaffinity-based proteomics. J Am Chem Soc 143:3037–3042
Lam AT, Zhang XN, Courouble V et al (2021) A bifunctional NAD+for profiling poly-ADP-ribosylation-dependent interacting proteins. ACS Chem Biol 16:389–396
Kliza KW, Liu Q, Roosenboom LWM et al (2021) Reading ADP-ribosylation signaling using chemical biology and interaction proteomics. Mol Cell:1–16
Farmer H, McCabe H, Lord CJ et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434:917–921
Bryant HE, Schultz N, Thomas HD et al (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434:913–917
Curtin NJ, Szabo C (2020) Poly(ADP-ribose) polymerase inhibition: past, present and future. Nat Rev Drug Discov 19:711–736
Langelier M-F, Planck JL, Servent KM et al (2011) Purification of human PARP-1 and PARP-1 domains from Escherichia coli for structural and biochemical analysis. Methods in molecular biology, In, pp 209–226
Muthurajan U, Mattiroli F, Bergeron S et al (2016) In vitro chromatin assembly: strategies and quality control. Methods Enzymol 573:3
Acknowledgments
Funding was provided by the National Cancer Institute R01 CA218255 and by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Rudolph, J., Luger, K. (2023). Analyzing PARP1 Activity: Small Molecule Reactants and Attached Chains of Poly (ADP-Ribose). In: Tulin, A.V. (eds) Poly(ADP-Ribose) Polymerase. Methods in Molecular Biology, vol 2609. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2891-1_4
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2891-1_4
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2890-4
Online ISBN: 978-1-0716-2891-1
eBook Packages: Springer Protocols