Abstract
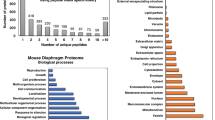
Following large-scale protein separation by two-dimensional gel electrophoresis or liquid chromatography, mass spectrometry–based proteomics can be used for the swift identification and characterization of cardiac proteins and their various proteoforms. Comparative cardiac proteomics has been widely applied for the systematic analysis of heart disease and the establishment of novel diagnostic protein biomarkers. The X-linked neuromuscular disorder Duchenne muscular dystrophy is a multisystemic disease that is characterized by late-onset cardiomyopathy. This chapter outlines the bioinformatic analysis of the subproteomic profile of cardiac tissue from wild-type versus the dystrophic mdx-4cv mouse model of dystrophinopathy.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Matkovich SJ (2019) Multiomic approaches to delineate the pathogenesis of cardiac disease. Curr Opin Cardiol 34:246–253
Sohag MMH, Raqib SM, Akhmad SA (2021) OMICS approaches in cardiovascular diseases: a mini review. Genomics Inform 19:e13
Sarhene M, Wang Y, Wei J, Huang Y, Li M, Li L, Acheampong E, Zhengcan Z, Xiaoyan Q, Yunsheng X, Jingyuan M, Xiumei G, Guanwei F (2019) Biomarkers in heart failure: the past, current and future. Heart Fail Rev 24:867–903
Joshi A, Rienks M, Theofilatos K, Mayr M (2021) Systems biology in cardiovascular disease: a multiomics approach. Nat Rev Cardiol 18:313–330
Gómez-Mendoza DP, Lara-Ribeiro AC, Verano-Braga T (2021) Pathological cardiac remodeling seen by the eyes of proteomics. Biochim Biophys Acta Proteins Proteom 1869:140622
Padula MP, Berry IJ, Rourke O, MB, Raymond BB, Santos J, Djordjevic SP (2017) A comprehensive guide for performing sample preparation and top-down protein analysis. Proteomes 5:11
Dowling P, Zweyer M, Swandulla D, Ohlendieck K (2019) Characterization of contractile proteins from skeletal muscle using gel-based top-down proteomics. Proteomes 7:25
Cupp-Sutton KA, Wu S (2020) High-throughput quantitative top-down proteomics. Mol Omics 16:91–99
Carbonara K, Andonovski M, Coorssen JR (2021) Proteomes are of Proteoforms: embracing the complexity. Proteomes 9:38
Dupree EJ, Jayathirtha M, Yorkey H, Mihasan M, Petre BA, Darie CC (2020) A critical review of bottom-up proteomics: the good, the bad, and the future of this field. Proteomes 8:14
Dowling P, Gargan S, Zweyer M, Henry M, Meleady P, Swandulla D, Ohlendieck K (2020) Protocol for the bottom-up proteomic analysis of mouse spleen. STAR Protoc 1:100196
Révész Á, Hevér H, Steckel A, Schlosser G, Szabó D, Vékey K, Drahos L (2021) Collision energies: optimization strategies for bottom-up proteomics. Mass Spectrom Rev 2:e21763
Meyers TA, Townsend D (2019) Cardiac pathophysiology and the future of cardiac therapies in Duchenne muscular dystrophy. Int J Mol Sci 20:4098
Shih JA, Folch A, Wong BL (2020) Duchenne muscular dystrophy: the heart of the matter. Curr Heart Fail Rep 17:57–66
Duan D, Goemans N, Takeda S, Mercuri E, Aartsma-Rus A (2021) Duchenne muscular dystrophy. Nat Rev Dis Primers 7:13
Ohlendieck K, Swandulla D (2021) Complexity of skeletal muscle degeneration: multi-systems pathophysiology and organ crosstalk in dystrophinopathy. Pflugers Arch 73:1813–1839
da Silva TD, Massetti T, Crocetta TB, de Mello Monteiro CB, Carll A, Vanderlei LCM, Arbaugh C, Oliveira FR, de Abreu LC, Ferreira Filho C, Godleski J, Ferreira C (2018) Heart rate variability and cardiopulmonary dysfunction in patients with Duchenne muscular dystrophy: a systematic review. Pediatr Cardiol 39:869–883
Łoboda A, Dulak J (2020) Muscle and cardiac therapeutic strategies for Duchenne muscular dystrophy: past, present, and future. Pharmacol Rep 72:1227–1263
Kipke J, Birnkrant DJ, Jin JB, Aneja A, Bahler RC (2021) A systematic review of pharmacologic therapies for the cardiomyopathy of Duchenne muscular dystrophy. Pediatr Pulmonol 56:782–795
Holland A, Ohlendieck K (2014) Proteomic profiling of the dystrophin-deficient mdx phenocopy of dystrophinopathy-associated cardiomyopathy. Biomed Res Int 2014:246195
Gowran A, Brioschi M, Rovina D, Chiesa M, Piacentini L, Mallia S, Banfi C, Pompilio G, Santoro R (2021) Multiomic approaches to uncover the complexities of dystrophin-associated cardiomyopathy. Int J Mol Sci 22:8954
Gulston MK, Rubtsov DV, Atherton HJ, Clarke K, Davies KE, Lilley KS, Griffin JL (2008) A combined metabolomic and proteomic investigation of the effects of a failure to express dystrophin in the mouse heart. J Proteome Res 7:2069–2077
Lewis C, Jockusch H, Ohlendieck K (2010) Proteomic profiling of the dystrophin-deficient MDX heart reveals drastically altered levels of key metabolic and contractile proteins. J Biomed Biotechnol 2010:648501
Johnson EK, Zhang L, Adams ME, Phillips A, Freitas MA, Froehner SC, Green-Church KB, Montanaro F (2012) Proteomic analysis reveals new cardiac-specific dystrophin-associated proteins. PLoS One 7:e43515
Holland A, Dowling P, Zweyer M, Swandulla D, Henry M, Clynes M, Ohlendieck K (2013) Proteomic profiling of cardiomyopathic tissue from the aged mdx model of Duchenne muscular dystrophy reveals a drastic decrease in laminin, nidogen and annexin. Proteomics 13:2312–2323
Murphy S, Dowling P, Zweyer M, Mundegar RR, Henry M, Meleady P, Swandulla D, Ohlendieck K (2016) Proteomic analysis of dystrophin deficiency and associated changes in the aged mdx-4cv heart model of dystrophinopathy-related cardiomyopathy. J Proteome 145:24–36
Chung HS, Kim GE, Holewinski RJ, Venkatraman V, Zhu G, Bedja D, Kass DA, Van Eyk JE (2017) Transient receptor potential channel 6 regulates abnormal cardiac S-nitrosylation in Duchenne muscular dystrophy. Proc Natl Acad Sci U S A 114:E10763–E10771
Tamiyakul H, Kemter E, Kösters M, Ebner S, Blutke A, Klymiuk N, Flenkenthaler F, Wolf E, Arnold GJ, Fröhlich T (2020) Progressive proteome changes in the myocardium of a Pig Model for Duchenne muscular dystrophy. iScience 23:101516
Blundon M, Ganesan V, Redler B, Van PT, Minden JS (2019) Two-dimensional difference gel electrophoresis. Methods Mol Biol 1855:229–247
Partridge TA (2013) The mdx mouse model as a surrogate for Duchenne muscular dystrophy. FEBS J 280:4177–4186
Bengtsson NE, Hall JK, Odom GL, Phelps MP, Andrus CR, Hawkins RD, Hauschka SD, Chamberlain JR, Chamberlain JS (2017) Muscle-specific CRISPR/Cas9 dystrophin gene editing ameliorates pathophysiology in a mouse model for Duchenne muscular dystrophy. Nat Commun 8:14454
Mi H, Ebert D, Muruganujan A, Mills C, Albou LP, Mushayamaha T, Thomas PD (2021) PANTHER version 16: a revised family classification, tree-based classification tool, enhancer regions and extensive API. Nucleic Acids Res 49(D1):D394–D403
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C (2021) The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49(D1):D605–D612
Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X, Liu S, Bo X, Yu G (2021) clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. The. Innovations 2021:100141
Grantham J (2020) The molecular Chaperone CCT/TRiC: an essential component of Proteostasis and a potential modulator of protein aggregation. Front Genet 11:172
Lin BL, Song T, Sadayappan S (2017) Myofilaments: movers and rulers of the sarcomere. Compr Physiol 7:675–692
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kanehisa M, Sato Y, Furumichi M, Morishima K, Tanabe M (2019) New approach for understanding genome variations in KEGG. Nucleic Acids Res 7(D1):D590–D595
Kanehisa M, Sato Y, Kawashima M (2022) KEGG mapping tools for uncovering hidden features in biological data. Protein Sci 31:47–53
Tyanova S, Temu T, Cox J (2016) The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat Protoc 11:2301–2319
Wichmann C, Meier F, Virreira Winter S, Brunner AD, Cox J, Mann M (2019) MaxQuant.Live enables global targeting of more than 25,000 peptides. Mol Cell Proteomics 18:982–994
Tyanova S, Cox J (2018) Perseus: a bioinformatics platform for integrative analysis of proteomics data in cancer research. Methods Mol Biol 1711:133–148
Mi H, Muruganujan A, Huang X, Ebert D, Mills C, Guo X, Thomas PD (2019) Protocol update for large-scale genome and gene function analysis with the PANTHER classification system (v.14.0). Nat Protoc 14:703–721
Murphy S, Henry M, Meleady P, Zweyer M, Mundegar RR, Swandulla D, Ohlendieck K (2015) Simultaneous Pathoproteomic evaluation of the dystrophin-glycoprotein complex and secondary changes in the mdx-4cv Mouse Model of Duchenne muscular dystrophy. Biology (Basel) 4:397–423
Zweyer M, Sabir H, Dowling P, Gargan S, Murphy S, Swandulla D, Ohlendieck K (2022) Histopathology of Duchenne muscular dystrophy in correlation with changes in proteomic biomarkers. Histol Histopathol 37:101–116
Acknowledgments
Research in the author’s laboratory has been supported by a project grant from the Kathleen Lonsdale Institute for Human Health Research, Maynooth University and equipment funding under the Research Infrastructure Call 2012 by Science Foundation Ireland (SFI-12/RI/2346/3).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Murphy, S., Zweyer, M., Swandulla, D., Ohlendieck, K. (2023). Bioinformatic Analysis of the Subproteomic Profile of Cardiomyopathic Tissue. In: Ohlendieck, K. (eds) Difference Gel Electrophoresis. Methods in Molecular Biology, vol 2596. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2831-7_26
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2831-7_26
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2830-0
Online ISBN: 978-1-0716-2831-7
eBook Packages: Springer Protocols