Abstract
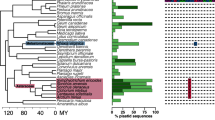
High-throughput sequencing of universal bacterial 16S rRNA gene (16S rDNA) amplicons is a routine method for characterizing bacterial diversity in a range of environments. For eukaryotic host-associated communities, however, plastid and mitochondrial genes are often co-amplified with, and greatly outnumber, bacterial 16S rDNA. This makes it difficult to obtain sufficient numbers of target 16S rDNA sequences to characterize the diversity of endophytic bacterial communities. This chapter describes a method that improves the amplification of bacterial 16S rDNA from plant tissues by using a peptide nucleic acid (PNA) PCR clamp. The PNA clamp selectively binds to a targeted region of the plant genome and inhibits its amplification during PCR. PNA clamps are especially useful for characterizing bacterial communities on plant tissues with lower levels of microbial colonization such as the root tips and leaves.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Donn S, Kirkegaard JA, Perera G et al (2015) Evolution of bacterial communities in the wheat crop rhizosphere. Environ Microbiol 17:610–621. https://doi.org/10.1111/1462-2920.12452
Kawasaki A, Donn S, Ryan PR et al (2016) Microbiome and exudates of the root and rhizosphere of Brachypodium distachyon, a model for wheat. PLoS One 11:e0164533. https://doi.org/10.1371/journal.pone.0164533
Rho H, Hsieh M, Kandel SL et al (2018) Do endophytes promote growth of host plants under stress? A meta-analysis on plant stress mitigation by endophytes. Microbial Ecol 75:407–418. https://doi.org/10.1007/s00248-017-1054-3
Rastogi G, Tech JJ, Coaker GL et al (2010) A PCR-based toolbox for the culture-independent quantification of total bacterial abundances in plant environments. J Microbiol Methods 83:127–132. https://doi.org/10.1016/j.mimet.2010.08.006
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1. https://doi.org/10.1093/nar/gks808
Chelius MK, Triplett EW (2001) The diversity of archaea and bacteria in association with the roots of Zea mays L. Microbial Ecol 41:252–263. https://doi.org/10.1007/s002480000087
Hanshew AS, Mason CJ, Raffa KF et al (2013) Minimization of chloroplast contamination in 16S rRNA gene pyrosequencing of insect herbivore bacterial communities. J Microbiol Methods 95:149–155. https://doi.org/10.1016/j.mimet.2013.08.007
Lundberg DS, Yourstone S, Mieczkowski P et al (2013) Practical innovations for high-throughput amplicon sequencing. Nat Methods 10:999–1002. https://doi.org/10.1038/nmeth.2634
Sakai M, Ikenaga M (2013) Application of peptide nucleic acid (PNA)-PCR clamping technique to investigate the community structures of rhizobacteria associated with plant roots. J Microbiol Methods 92:281–288. https://doi.org/10.1016/j.mimet.2012.09.036
Nielsen PE, Egholm M (1999) An introduction to peptide nucleic acid. Curr Issues Mol Biol 1:89–104. https://doi.org/10.21775/cimb.001.089
Ørum H, Nielsen PE, Egholm M et al (1993) Single base pair mutation analysis by PNA directed PCR clamping. Nucleic Acids Res 21:5332–5336. https://doi.org/10.1093/nar/21.23.5332
Beckers B, De Beeck MO, Thijs S et al (2016) Performance of 16s rDNA primer pairs in the study of rhizosphere and endosphere bacterial microbiomes in metabarcoding studies. Front Microbiol 7. https://doi.org/10.3389/fmicb.2016.00650
Cole JR, Wang Q, Fish JA et al (2014) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633–D642. https://doi.org/10.1093/nar/gkt1244
Terahara T, Chow S, Kurogi H et al (2011) Efficiency of peptide nucleic acid-directed PCR clamping and its application in the investigation of natural diets of the Japanese eel leptocephali. PLoS One 6:e25715. https://doi.org/10.1371/journal.pone.0025715
Kawasaki A, Okada S, Zhang C et al (2018) A sterile hydroponic system for characterising root exudates from specific root types and whole-root systems of large crop plants. Plant Methods 14:114. https://doi.org/10.1186/s13007-018-0380-x
Acknowledgments
A.K. is funded by CSIRO OCE Postdoctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Kawasaki, A., Ryan, P.R. (2021). Peptide Nucleic Acid (PNA) Clamps to Reduce Co-amplification of Plant DNA During PCR Amplification of 16S rRNA Genes from Endophytic Bacteria. In: Carvalhais, L.C., Dennis, P.G. (eds) The Plant Microbiome. Methods in Molecular Biology, vol 2232. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1040-4_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1040-4_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1039-8
Online ISBN: 978-1-0716-1040-4
eBook Packages: Springer Protocols