Abstract
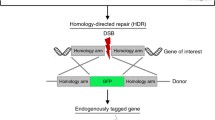
Here, we describe how to utilize CRISPR/Cas9 technology in the generation of tissue culture cells with fluorescently tagged caveolar components as well as cells deleted of endogenous caveolar components. As one example, we will describe tagging of EHD2, caveolar neck protein, with Green Fluorescent protein (eGFP) from endogenous loci (knock-in, KI). As another example, we will describe deletion (knock-out, KO) of Caveolin1 (Cav1), an essential caveolar component in NIH/3T3 cells. In both instances, the modifications were achieved by using Cas9 delivery on plasmid DNA by electroporation and by utilizing FACS cell sorting for selection or enrichment of edited population of cells. We also provide a list with tested gRNA sequences to successfully produce KI and KO of other caveolar components.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Doyon JB, Zeitler B, Cheng J, Cheng AT, Cherone JM, Santiago Y, Lee AH, Vo TD, Doyon Y, Miller JC, Paschon DE, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Drubin DG (2011) Rapid and efficient clathrin-mediated endocytosis revealed in genome-edited mammalian cells. Nat Cell Biol 13:331–337
Shvets E, Bitsikas V, Howard G, Hansen CG, Nichols BJ (2015) Dynamic caveolae exclude bulk membrane proteins and are required for sorting of excess glycosphingolipids. Nat Commun 6:6867
Hayer A, Stoeber M, Ritz D, Engel S, Meyer HH, Helenius A (2010) Caveolin-1 is ubiquitinated and targeted to intralumenal vesicles in endolysosomes for degradation. J Cell Biol 191:615–629
Parton RG, Howes MT (2010) Revisiting caveolin trafficking: the end of the caveosome. J Cell Biol 191:439–441
Yeow I, Howard G, Chadwick J, Mendoza-Topaz C, Hansen CG, Nichols BJ, Shvets E (2017) EHD proteins cooperate to generate caveolar clusters and to maintain caveolae during repeated mechanical stress. Curr Biol 27:2951–2962
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F (2013) Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8:2281–2308
Cost GJ, Cozzarelli (2007) Directed assembly of DNA molecules via simultaneous ligation and digestion. Biotechniques 42:84, 86–89
Mendoza-Topaz C, Nelson G, Howard G, Hafner S, Rademacher P, Frick M, Nichols BJ (2018) Cells respond to deletion of CAV1 by increasing synthesis of extracellular matrix. PLoS One 13:e0205306
Zhang JP, Li XL, Li GH, Chen W, Arakaki C, Botimer GD, Baylink D, Zhang L, Wen W, Fu YW, Xu J, Chun N, Yuan W, Cheng T, Zhang XB (2017) Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol 18:35
de Kreuk BJ, Nethe M, Fernandez-Borja M, Anthony EC, Hensbergen PJ, Deelder AM, Plomann M, Hordijk PL (2011) The F-BAR domain protein PACSIN2 associates with Rac1 and regulates cell spreading and migration. J Cell Sci 124:2375–2388
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Shvets, E., Mendoza-Topaz, C. (2020). Tagging and Deleting of Endogenous Caveolar Components Using CRISPR/Cas9 Technology. In: Blouin, C. (eds) Caveolae. Methods in Molecular Biology, vol 2169. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0732-9_14
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0732-9_14
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0731-2
Online ISBN: 978-1-0716-0732-9
eBook Packages: Springer Protocols