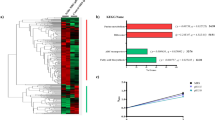
Abstract
Effect of acidic condition on the fermentation characteristics was investigated by the continuous culture of Escherichia coli. In accordance with down-regulation of crp gene transcript level as well as up-regulation of arcA, the expressions of the TCA cycle genes were down-regulated, which caused more acetate formation at acidic condition under aerobic condition. It was also shown that yfiD was up-regulated in accordance with up-regulation of fnr, and the respiratory pathway genes were up-regulated under acidic condition. The effect of phoB gene knockout on fermentation characteristics was also investigated. Under micro-aerobic condition, the fermentation pattern changed in such a way that formate and lactate were more produced at lower pH due to up-regulations of pflA, yfiD and ldhA genes, whereas ethanol was less produced as compared to the case at neutral pH. The overall regulation mechanism under acidic condition was clarified based on fermentation characteristics and gene transcript levels.



Similar content being viewed by others
References
Arnold KW, Kaspar CW (1995) Starvation- and stationary-phase induced acid tolerance in Escherichia coli O157:H7. Appl Environ Microbiol 61:2037–2039
Aronis HR (2002) Signal transduction and regulatory mechanisms involved in control of the σS (RpoS) subunit of RNA polymerase. Microbiol Mol Microbiol Rev 66:373–395
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita T, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:0008
Bearson BL, Wilson L, Foster JW (1998) A low pH-inducible, PhoPQ-dependent acid tolerance response protects Salmonella typhimurium against inorganic acid Stress. J Bacteriol 180:2409–2417
Blankenhorn D, Phillips J, Slonczewski J (1999) Acid- and base-induced proteins during aerobic and anaerobic growth of Escherichia coli revealed by two-dimensional gel electrophoresis. J Bacteriol 181:2209–2216
Bunch PK, Mat-Jan F, Lee N, Clark DP (1997) The ldhA gene encoding the fermentative lactate dehydrogenase of Escherichia coli. Microbiol 143:187–195
Castanie-Cornet MP, Foster JW (2001) Escherichia coli acid resistance: cAMP receptor protein and a 20 bp cis-acting sequence control pH and stationary phase expression of the gadA and gadBC glutamate decarboxylase genes. Microbiology 147:709–715
Castanié-Cornet MP, Treffandier H, Francez-Charlot A, Gutierrez C, Cam K (2007) The glutamate-dependent acid resistance system in Escherichia coli: essential and dual role of the His-Asp phosphorelay RcsCDB/AF. Microbiology 153:238–246
Castanié-Cornet MP, Cam K, Bastiat B, Cros A, Bordes P, Gutierrez C (2010) Acid stress response in Escherichia coli: mechanism of regulation of gadA transcription by RcsB and GadE. Nucleic Acids Res 38(11):3546–3554
Cheville AM, Arnold KW, Buchrieser C, Cheng CM, Kaspa CW (1996) RpoS regulation of acid, heat, and salt tolerance in Escherichia coli O157:H7. Appl Environ Microbiol 62:1822–1824
Dukan S, Dadon S, Smulski DR, Belkin S (1996) Hypochlorous acid activates the heat shock and soxRS systems of Escherichia coli. Appl Environ Microbiol 2(11):4003–4008
Eguchi Y, Itou J, Yamane M, Demizu R, Yamato F, Okada A, Mori H, Kato A, Utsumi R (2007) B1500, a small membrane protein, connects the two-component systems EvgS/EvgA and PhoQ/PhoP in Escherichia coli. Proc Natl Acad Sci USA 104:18712–18717
Fukuda M, Yasukochi Y, Kikuchi Y, Nagata Y, Kimbara K, Horiuchi H, Takagi M, Yano K (1994) Identification of the bphA and bphB genes of Pseudomonas sp. strain KKS102 involved in degradation of biphenyl and polychlorinated biphenyls. Biochem Biophys Res Commun 202:850–856
Gajiwala KS, Burley SK (2000) HDEA, a periplasmic protein that supports acid resistance in pathogenic enteric bacteria. J Mol Biol 295:605–612
Gallegos M, Schleif R, Bairoch A, Hofmann K, Ramos JL (1997) AraC/XylS family of transcriptional regulators. Microbiol Mol Biol Rev 61:393–410
Green J, Baldwin ML (1997) HlyX, the FNR homologue of Actinobacillus pleuropneumoniae, is a [4Fe-4S]-containing oxygen-responsive transcription regulator that anaerobically activates FNR-dependent class I promoters via an enhanced AR1-contact. Mol Microbiol 24:593–605
Groisman EA (2001) The pleiotropic two-component regulatory system PhoP-PhoQ. J Bacteriol 183:1835–1842
Hersh BM, Farooq FT, Barstad DN, Blankenhorn DL, Slonczewski JL (1996) A glutamate-dependent acid resistance gene in Escherichia coli. J Bacteriol 178:3978–3981
Heyde M, Laloi P, Portalier R (2000) Involvement of carbon source and acetyl phosphate in the external-pH-dependent expression of porin genes in Escherichia coli. J Bacteriol 182:198–202
Jung IL, Kim IG (2003) Polyamines and glutamate decarboxylase-based acid resistance in Escherichia coli. J Biol Chem 278(6):22846–22852
Kabir MM, Shimizu K (2003) Gene expression patterns for metabolic pathway in pgi knockout Escherichia coli with and without phb genes based on RT-PCR. J Biotechnol 105:11–31
Kato A, Ohnishi H, Yamamoto K, Furuta E, Tanabe H, Utsumi R (2000) Transcription of emrKY is regulated by the EvgA-EvgS two-component system in Escherichia coli K-12. Biosci Biotechnol Biochem 64:1203–1209
Kier LD, Weppelman RM, Ames BN (1979) Regulation of nonspecific acid phosphatase in Salmonella: phoN and phoP genes. J Bacteriol 138(1):155–161
Kihara M, Macnab R (1981) Cytoplasmic pH mediates pH taxis and weak-acid repellent taxis of bacteria. J Bacteriol 145:1209–1221
Kirkpatrick C, Maurer L, Oyelakin N, Yoncheva Y (2001) Acetate and formate stress: opposite responses in the proteome of Escherichia coli. J Bacteriol 183:6466–6477
Lange R, Hengge-Aronis R (1994) The cellular concentration of σS subunit of RNA polymerase in Escherichia coli is controlled at the levels of transcription, translation, and protein stability. Genes Dev 8:1600–1612
Lazar SW, Almirón M, Tormo A, Kolter R (1998) Role of the Escherichia coli SurA protein in stationary-phase survival. J Bacteriol 180:5704–5711
Lesley JA, Waldburger CD (2003) Repression of Escherichia coli PhoP-PhoQ signaling by acetate reveals a regulatory role for acetyl coenzyme A. J Bacteriol 185(8):2563–2570
Liochev SI, Hausladen A, Beyer WF, Fridorich I (1994) NADPH: ferredoxin oxidoreductase acts as a paraquat diaphorase and is a member of the sox regulon. PNAS USA 91:1328–1331
Ma D, Cook DN, Alberti M, Pon NG, Nikaido H, Hearst JE (1995) Genes acrA and acrB encode a stress induced efflux system of Escherichia coli. Mol Microbiol 16:45–55
Ma Z, Gong S, Richard H, Tucker DL, Conway T, Foster JW (2003) GadE (YhiE) activates glutamate decarboxylase-dependent acid resistance in Escherichia coli K-12. Mol Microbiol 49:1309–1320
Marzan LW, Shimizu K (2011) Metabolic regulation of Escherichia coli and its phoB and phoR genes knockout mutants under phosphate and nitrogen limitations as well as at acidic condition. Microb Cell Fact 10:39
Masuda N, Church GM (2002) Escherichia coli gene expression responsive to levels of response regulator EvgA. J Bacteriol 184:6225–6234
Masuda N, Church GM (2003) Regulatory network of acid resistance genes in Escherichia coli. Mol Microbiol 48(3):699–712
Neil RW, Sarah LM, Robin AH, Gary S, Ruth ER, Margaret MA, Jeffrey G (2002) Expression of the Escherichia coli yfiD gene responds to intracellular pH and reduces the accumulation of acidic metabolic end products. Microbiology 148:1015–1026
Nishino K, Yamaguchi A (2001) Overexpression of the response regulator evgA of the two-component signal transduction system modulates multidrug resistance conferred by multidrug resistance transporters. J Bacteriol 183:1455–1458
Olofsson K, Bertilsson M, Lidén G (2008) A short review on SSF—an interesting process option for ethanol production from lignocellulosic feedstocks. Biotechnol Biofuels 7:1
Rahman M, Shimizu K (2008) Altered acetate metabolism and biomass production in several Escherichia coli mutants lacking rpoS-dependent metabolic pathway genes. Mol BioSyst 4:160–169
Rahman M, Hasan MR, Oba T, Shimizu K (2006) Effect of rpoS gene knockout on the metabolism of Escherichia coli during exponential growth phase and early stationary phase based on gene expressions, enzyme activities and intracellular metabolite concentrations. Biotechnol Bioeng 94:585–595
Rahman M, Hasa MM, Shimizu K (2008) Growth phase-dependent changes in the expression of global regulatory genes and associated metabolic pathways in Escherichia coli. Biotechnol Lett 30:853–860
Richard HT, Foster JW (2003) Acid resistance in Escherichia coli. Adv Appl Microbiol 52:167–186
Richard HT, Foster JW (2004) Escherichia coli glutamate- and arginine-dependent acid resistance systems increase internal pH and reverse transmembrane potential. J Bacteriol 86(18):6032–6041
Robison K, Church G (1994) Dpinteract: a database on DNA–protein interactions. [WWW document]. http://arep.med.harvard.edu/dpinteract
Russell JB, Diez-Gonzalez F (1998) The effects of fermentation acids on bacterial Growth. Adv Microb Physiol 39:205–234
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Sayed AK, Foster JW (2009) A 750 bp sensory integration region directs global control of the Escherichia coli GadE acid resistance regulator. Mol Microbiol 71(6):1435–1450
Schauder S, Bassler BL (2001) The languages of bacteria. Genes Dev 15:1468–1480
Small P, Blankerhorn D, Welty D, Zinser E, Slonczewski JL (1994) Acid and base resistance in Escherichia coli and Shigella flexneri: role of rpoS and growth pH. J Bacteriol 176:1729–1737
Soncini FC, Vescovi EG, Solomon F, Groisman EA (1996) Molecular basis of the magnesium deprivation response in Salmonella typhimurium: identification of PhoP-regulated genes. J Bacteriol 178:5092–5099
Stincone A, Daudi N, Rahman AS, Antczak P, Henderson I, Cole J, Johnson MD, Lund P, Falciani F (2011) A systems biology approach sheds new light on Escherichia coli acid resistance. Nucleic Acids Res 39(17):7512–7528
Suziedeliené E, Suziedélis K, Garbenciuté V, Normak S (1999) The acid-inducible asr gene in Escherichia coli: transcriptional control by the PhoBR operon. J Bacteriol 181(7):2084–2093
Tucker DL, Tucker N, Ma Z, Foster JW, Miranda RL, Cohen PS, Conway T (2003) Genes of GadX-GadW regulon in Escherichia coli. J Bacteriol 185:3190–3201
Vescovi EG, Soncini FC, Groisman EA (1996) Mg2+as an extracellular signal: environmental regulation of Salmonella virulence. Cell 84:165–174
Waterman SR, Small PLC (2003) Transcriptional expression of Escherichia coli glutamate-dependent acid resistance gene gadA and gadBC in an hns rpoS mutant. J Bacteriol 185:4644–4647
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Jorge Membrillo-Hernández.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Marzan, L.W., Hasan, C.M.M. & Shimizu, K. Effect of acidic condition on the metabolic regulation of Escherichia coli and its phoB mutant. Arch Microbiol 195, 161–171 (2013). https://doi.org/10.1007/s00203-012-0861-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-012-0861-7