Abstract
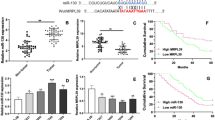
MicroRNAs (miRNAs) are a class of short, noncoding RNAs that act a crucial role in tumor development. Previous studies showed that miR-448 expression was deregulated in many tumors. However, the role of miR-448 in gastric cancer (GC) remains unknown. In our study, we demonstrated that miR-448 expression was downregulated in GC tissues compared with the corresponding nontumor tissues. We also showed that miR-448 expression was downregulated in GC cell lines. Ectopic expression of miR-448 suppressed GC cell proliferation, colony formation, and invasion. Moreover, we identified A Disintegrin And Metalloproteinases 10 (ADAM10) as a direct target gene of miR-448 in GC cell. ADAM10 expression was upregulated in GC tissues and cells. Furthermore, the expression level of miR-448 was negatively correlated with the expression level of ADAM10 in GC tissues. Moreover, ADAM10 overexpression rescued the effect of miR-448-mediated GC cell proliferation, colony formation, and invasion. These results demonstrated that miR-448 might play as a tumor suppressor miRNA partly through targeting ADAM10 expression.





Similar content being viewed by others
References
Liang J, Liu X, Xue H, Qiu B, Wei B, Sun K. Microrna-103a inhibits gastric cancer cell proliferation, migration and invasion by targeting c-myb. Cell Prolif. 2015;48:78–85.
Duan JH, Fang L. MicroRNA-92 promotes gastric cancer cell proliferation and invasion through targeting FXR. Tumour Biol. 2014;35(11):11013–9.
Liu G, Jiang C, Li D, Wang R, Wang W. MiRNA-34a inhibits EGFR-signaling-dependent MMP7 activation in gastric cancer. Tumour Biol. 2014;35(10):9801–6.
Wan X, Ding X, Chen S, Song H, Jiang H, Fang Y, et al. The functional sites of miRNAs and lncRNAs in gastric carcinogenesis. Tumour Biol. 2015;36:521–32.
Wu Y, Tao Y, Chen Y, Xu W. RhoC regulates the proliferation of gastric cancer cells through interaction with IQGAP1. PLoS One. 2012;7:e48917.
Shen J, Niu W, Zhou M, Zhang H, Ma J, Wang L. MicroRNA-410 suppresses migration and invasion by targeting MDM2 in gastric cancer. PLoS One. 2014;9:e104510.
Otsubo T, Akiyama Y, Hashimoto Y, Shimada S, Goto K, Yuasa Y. MicroRNA-126 inhibits SOX2 expression and contributes to gastric carcinogenesis. PLoS One. 2011;6:e16617.
Yasui W, Sentani K, Sakamoto N, Anami K, Naito Y, Oue N. Molecular pathology of gastric cancer: research and practice. Pathol Res Pract. 2011;207:608–12.
Li X, Luo F, Li Q, Xu M, Feng D, Zhang G, et al. Identification of new aberrantly expressed miRNAs in intestinal-type gastric cancer and its clinical significance. Oncol Rep. 2011;26:1431–9.
Inoue T, Iinuma H, Ogawa E, Inaba T, Fukushima R. Clinicopathological and prognostic significance of microRNA-107 and its relationship to DICER1 mRNA expression in gastric cancer. Oncol Rep. 2012;27:1759–64.
Fesler A, Zhai H, Ju J. Mir-129 as a novel therapeutic target and biomarker in gastrointestinal cancer. OncoTargets Ther. 2014;7:1481–5.
Wang M, Gu H, Wang S, Qian H, Zhu W, Zhang L, et al. Circulating miR-17-5p and miR-20a: molecular markers for gastric cancer. Mol Med Report. 2012;5:1514–20.
Zhang X, Tang J, Zhi X, Xie K, Wang W, Li Z, et al. miR-874 functions as a tumor suppressor by inhibiting angiogenesis through STAT3/VEGF-A pathway in gastric cancer. Oncotarget. 2015;6:1605–17.
Li Z, Yu X, Wang Y, Shen J, Wu WK, Liang J, et al. By downregulating TIAM1 expression, microRNA-329 suppresses gastric cancer invasion and growth. Oncotarget. 2015;6:17559–69.
Li Z, Lei H, Luo M, Wang Y, Dong L, Ma Y, et al. DNA methylation downregulated miR-10b acts as a tumor suppressor in gastric cancer. Gastric Cancer. 2015;18:43–54.
Yu X, Li Z, Chen G, Wu WK. MicroRNA-10b induces vascular muscle cell proliferation through Akt pathway by targeting TIP30. Curr Vasc Pharmacol. 2015;13:679–86.
Yu X, Li Z, Shen J, Wu WK, Liang J, Weng X, et al. MicroRNA-10b promotes nucleus pulposus cell proliferation through RhoC-Akt pathway by targeting HOXD10 in intervetebral disc degeneration. PLoS One. 2013;8:e83080.
Song JH, Meltzer SJ. MicroRNAs in pathogenesis, diagnosis, and treatment of gastroesophageal cancers. Gastroenterology. 2012;143:35–47. e32.
Deng H, Guo Y, Song H, Xiao B, Sun W, Liu Z, et al. MicroRNA-195 and microRNAa-378 mediate tumor growth suppression by epigenetical regulation in gastric cancer. Gene. 2013;518:351–9.
Bin Z, Dedong H, Xiangjie F, Hongwei X, Qinghui Y. The microRNA-367 inhibits the invasion and metastasis of gastric cancer by directly repressing Rab23. Genet Test Mol Biomarkers. 2015;19:69–74.
Yu X, Li Z: The role of mirnas in cutaneous squamous cell carcinoma. Journal of cellular and molecular medicine 2016;20:3–9.
Li Z, Yu X, Shen J, Law PT, Chan MT, Wu WK. MicroRNA expression and its implications for diagnosis and therapy of gallbladder cancer. Oncotarget. 2015;6:13914–24.
Yu X, Li Z. The role of microRNAs expression in laryngeal cancer. Oncotarget. 2015;6:23297–305.
Li Z, Yu X, Shen J, Liu Y, Chan MT, Wu WK. MicroRNA dysregulation in rhabdomyosarcoma: a new player enters the game. Cell Prolif. 2015;48:511–6.
Itesako T, Seki N, Yoshino H, Chiyomaru T, Yamasaki T, Hidaka H, et al. The microRNA expression signature of bladder cancer by deep sequencing: the functional significance of the miR-195/497 cluster. PLoS One. 2014;9:e84311.
Vrba L, Munoz-Rodriguez JL, Stampfer MR, Futscher BW. Mirna gene promoters are frequent targets of aberrant DNA methylation in human breast cancer. PLoS One. 2013;8:e54398.
Xu X, Wang W, Su N, Zhu X, Yao J, Gao W, et al. miR-374a promotes cell proliferation, migration and invasion by targeting SRCIN1 in gastric cancer. FEBS Lett. 2015;589:407–13.
Kong WQ, Bai R, Liu T, Cai CL, Liu M, Li X, et al. MicroRNA-182 targets camp-responsive element-binding protein 1 and suppresses cell growth in human gastric adenocarcinoma. FEBS J. 2012;279:1252–60.
Han TS, Hur K, Xu G, Choi B, Okugawa Y, Toiyama Y, et al. MicroRNA-29c mediates initiation of gastric carcinogenesis by directly targeting ITGB1. Gut. 2015;64:203–14.
Lv Y, Lei Y, Hu Y, Ding W, Zhang C, Fang C. miR-448 negatively regulates ovarian cancer cell growth and metastasis by targeting CXCL12. Clin Transl Oncol. 2015;17:903–9.
Zhu H, Zhou X, Ma C, Chang H, Li H, Liu F, et al. Low expression of miR-448 induces EMT and promotes invasion by regulating ROCK2 in hepatocellular carcinoma. Cell Physiol Biochem. 2015;36:487–98.
Li QQ, Chen ZQ, Cao XX, Xu JD, Xu JW, Chen YY, et al. Involvement of Nf-kappab/miR-448 regulatory feedback loop in chemotherapy-induced epithelial-mesenchymal transition of breast cancer cells. Cell Death Differ. 2011;18:16–25.
Mullooly M, McGowan PM, Kennedy SA, Madden SF, Crown J, O’ Donovan N, et al. Adam10: a new player in breast cancer progression? Br J Cancer. 2015;113:945–51.
Jing P, Sa N, Liu X, Xu W. MicroR-140-5p suppresses tumor cell migration and invasion by targeting ADAM10-mediated Notch1 signaling pathway in hypopharyngeal squamous cell carcinoma. Exp Mol Pathol. 2015;100:132–8.
Saftig P, Lichtenthaler SF. The alpha secretase ADAM10: a metalloprotease with multiple functions in the brain. Prog Neurobiol. 2015;135:1–20.
Chang YC, Jan CI, Peng CY, Lai YC, Hu FW, Yu CC. Activation of microRNA-494-targeting Bmi1 and ADAM10 by silibinin ablates cancer stemness and predicts favourable prognostic value in head and neck squamous cell carcinomas. Oncotarget. 2015;6:24002–16.
Yoshimura T, Tomita T, Dixon MF, Axon AT, Robinson PA, Crabtree JE. ADAMs (a disintegrin and metalloproteinase) messenger RNA expression in helicobacter pylori-infected, normal, and neoplastic gastric mucosa. J Infect Dis. 2002;185:332–40.
Tanida S, Joh T, Itoh K, Kataoka H, Sasaki M, Ohara H, et al. The mechanism of cleavage of EGFR ligands induced by inflammatory cytokines in gastric cancer cells. Gastroenterology. 2004;127:559–69.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Xuesong Wu and Haoran Tang are co-first authors.
Rights and permissions
About this article
Cite this article
Wu, X., Tang, H., Liu, G. et al. miR-448 suppressed gastric cancer proliferation and invasion by regulating ADAM10. Tumor Biol. 37, 10545–10551 (2016). https://doi.org/10.1007/s13277-016-4942-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-4942-0