Abstract
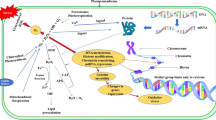
Ma-LMM01 is a lytic phage infecting the toxic cyanobacterium Microcystis aeruginosa. We have investigated the transcription dynamics of host genes during phage infection using quantitative reverse transcriptase-PCR. No significant shutdown of host transcription occurred. There was no change in the transcript levels of the psbA genes [photosystem II D1 (PSII D1)], but the transcript levels of the stress response genes and the alternative σ factor gene sigB were upregulated. The transcript levels of the Calvin cycle genes and pentose phosphate pathway genes did not change or only slightly decreased, suggesting that sufficient amounts of NADPH and nucleic acid were available without redirection of the carbon flux. These results suggest that Ma-LMM01 infection induces protection of the host’s photosynthetic apparatus, conserves the host’s PSII through phycobilisome degradation using its own NblA and provides protection to the photosystem using host stress response genes. This protection of the host’s photosystem without extensive genetic manipulation may have some benefits for viruses infecting cyanobacteria that inhabit surface waters and may also be advantageous for Ma-LMM01 to avoid the host defense systems.





Similar content being viewed by others
References
Thompson LR, Zeng Q, Kelly L, Huang KH, Singer AU, Stubbe J, Chisholm SW (2011) Phage auxiliary metabolic genes and the redirection of cyanobacterial host carbon metabolism. Proc Natl Acad Sci USA 108:E757–E764
Lavigne R, Darius P, Summer EJ, Seto D, Mahadevan P, Nilsson AS, Ackermann HW, Kropinski AM (2009) Classification of Myoviridae bacteriophages using protein sequence similarity. BMC Microbiol 9:224
Carstens EB, Ball LA (2009) Ratification vote on taxonomic proposals to the International Committee on Taxonomy of Viruses (2008). Arch Virol 154:1181–1188
Yoshida T, Nagasaki K, Takashima Y, Shirai Y, Tomaru Y, Takao Y, Sakamoto S, Hiroishi S, Ogata H (2008) Ma-LMM01 infecting toxic Microcystis aeruginosa illuminates diverse cyanophage genome strategies. J Bacteriol 190:1762–1772
Yoshida T, Takashima Y, Tomaru Y, Shirai Y, Takao Y, Hiroishi S, Nagasaki K (2006) Isolation and characterization of a cyanophage infecting the toxic cyanobacterium Microcystis aeruginosa. Appl Environ Microbiol 72:1239–1247
Yoshida-Takashima Y, Yoshida M, Ogata H, Nagasaki K, Hiroishi S, Yoshida T (2012) Cyanophage infection in the bloom-forming cyanobacteria Microcystis aeruginosa in surface freshwater. Microbes Environ 27:350–355
Mlouka A, Comte K, Castets AM, Bouchier C, Tandeau de Marsac N (2004) The gas vesicle gene cluster from Microcystis aeruginosa and DNA rearrangements that lead to loss of cell buoyancy. J Bacteriol 186:2355–2365
Gao EB, Gui JF, Zhang QY (2012) A novel cyanophage with a cyanobacterial nonbleaching protein A gene in the genome. J Virol 86:236–245
Kasai F, Kawachi M, Erata M, Watanabe MM (2004) List of strains, microalgae and protozoa. National Institute for Environmental Studies, Tsukuba
Yoshida T, Maki M, Okamoto H, Hiroishi S (2005) Coordination of DNA replication and cell division in cyanobacteria Microcystis aeruginosa. FEMS Microbiol Lett 251:149–154
Yagi O, Hagiwara T, Takamura Y, Sudo R (1984) Growth characteristics of axenic and unialgal Microcystis isolated from Lake Kasumigaura. Jpn J Water Poll Res 7:496–503
Sato N (1995) A family of cold-regulated RNA-binding protein genes in the cyanobacterium Anabaena variabilis M3. Nucleic Acids Res 23:2161–2167
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134
Yoshida M, Yoshida T, Yoshida-Takashima Y, Kashima A, Hiroishi S (2010) Real-time PCR detection of host-mediated cyanophage gene transcripts during infection of a natural Microcystis aeruginosa population. Microbes Environ 25:211–215
Kimura S, Yoshida T, Hosoda N, Honda T, Kuno S, Kamiji R, Hashimoto R, Sako Y (2012) Diurnal infection patterns and impact of Microcystis cyanophages in a Japanese pond. Appl Environ Microbiol 78:5805–5811
Lonetto M, Gribskov M, Gross CA (1992) The sigma 70 family: sequence conservation and evolutionary relationships. J Bacteriol 174:3843–3849
Mizuuchi K, O’Dea MH, Gellert M (1978) DNA gyrase: subunit structure and ATPase activity of the purified enzyme. Proc Natl Acad Sci USA 75:5960–5963
Radding CM (1981) Recombination activities of E. coli RecA protein. Cell 25:3–4
Kaneko T, Nakajima N, Okamoto S, Suzuki I, Tanabe Y, Tamaoki M, Nakamura Y, Kasai F, Watanabe A, Kawashima K, Kishida Y, Ono A, Shimizu Y, Takahashi C, Minami C, Fujishiro T, Kohara M, Katoh M, Nakazaki N, Nakayama S, Yamada M, Tabata S, Watanabe MM (2007) Complete genomic structure of the bloom-forming toxic cyanobacterium Microcystis aeruginosa NIES-843. DNA Res 14:247–256
Hohn T, Hohn B, Engel A, Wurtz M, Smith PR (1979) Isolation and characterization of the host protein groE involved in bacteriophage lambda assembly. J Mol Biol 129:359–373
Vierling E (1991) The roles of heat shock proteins in plants. Annu Rev Plant Physiol Plant Mol Biol 42:579–620
Sato T, Minagawa S, Kojima E, Okamoto N, Nakamoto H (2010) HtpG, the prokaryotic homologue of Hsp90, stabilizes a phycobilisome protein in the cyanobacterium Synechococcus elongatus PCC 7942. Mol Microbiol 76:576–589
Tanaka N, Nakamoto H (1999) HtpG is essential for the thermal stress management in cyanobacteria. FEBS Lett 458:117–123
Barker M, de Vries R, Nield J, Komenda J, Nixon PJ (2006) The deg proteases protect Synechocystis sp. PCC 6803 during heat and light stresses but are not essential for removal of damaged D1 protein during the photosystem two repair cycle. J Biol Chem 281:30347–30355
Clausen T, Kaiser M, Huber R, Ehrmann M (2011) HTRA proteases: regulated proteolysis in protein quality control. Nat Rev Mol Cell Biol 12:152–162
Imamura S, Asayama M, Shirai M (2004) In vitro transcription analysis by reconstituted cyanobacterial RNA polymerase: roles of group 1 and 2 sigma factors and a core subunit, RpoC2. Genes Cells 9:1175–1187
Asayama M, Kato H, Shibato J, Shirai M, Ohyama T (2002) The curved DNA structure in the 5′-upstream region of the light-responsive genes: its universality, binding factor and function for cyanobacterial psbA transcription. Nucleic Acids Res 30:4658–4666
Koerner JF, Snustad DP (1979) Shutoff of host macromolecular synthesis after T-even bacteriophage infection. Microbiol Rev 43:199–223
Pineda M, Gregory BD, Szczypinski B, Baxter KR, Hochschild A, Miller ES, Hinton DM (2004) A family of anti-sigma70 proteins in T4-type phages and bacteria that are similar to AsiA, a transcription inhibitor and co-activator of bacteriophage T4. J Mol Biol 344:1183–1197
Lindell D, Jaffe JD, Coleman ML, Futschik ME, Axmann IM, Rector T, Kettler G, Sullivan MB, Steen R, Hess WR, Church GM, Chisholm SW (2007) Genome-wide expression dynamics of a marine virus and host reveal features of co-evolution. Nature 449:83–86
Clokie MR, Shan J, Bailey S, Jia Y, Krisch HM, West S, Mann NH (2006) Transcription of a ‘photosynthetic’ T4-type phage during infection of a marine cyanobacterium. Environ Microbiol 8:827–835
Wikner J, Vallino JJ, Steward GF, Smith DC, Azam F (1993) Nucleic acids from the host bacterium as a major source of nucleotides for three marine bacteriophages. FEMS Microbiol Ecol 12:237–248
Wu J, Sunda W, Boyle EA, Karl DM (2000) Phosphate depletion in the Western North Atlantic Ocean. Science 289:759–762
Poranen MM, Ravantti JJ, Grahn AM, Gupta R, Auvinen P, Bamford DH (2006) Global changes in cellular gene expression during bacteriophage PRD1 infection. J Virol 80:8081–8088
Makarova KS, Wolf YI, Snir S, Koonin EV (2011) Defense islands in bacterial and archaeal genomes and prediction of novel defense systems. J Bacteriol 193:6039–6056
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Kuno S, Yoshida T, Kaneko T, Sako Y (2012) Intricate interactions between the bloom-forming cyanobacterium Microcystis aeruginosa and foreign genetic elements, revealed by diversified clustered regularly interspaced short palindromic repeat (CRISPR) signatures. Appl Environ Microbiol 78:5353–5360
Fineran PC, Blower TR, Foulds IJ, Humphreys DP, Lilley KS, Salmond GP (2009) The phage abortive infection system, ToxIN, functions as a protein-RNA toxin-antitoxin pair. Proc Natl Acad Sci USA 106:894–899
Engelberg-Kulka H, Amitai S, Kolodkin-Gal I, Hazan R (2006) Bacterial programmed cell death and multicellular behavior in bacteria. PLoS Genet 2:e135
Imamura S, Asayama M (2009) Sigma factors for cyanobacterial transcription. Gene Regul Syst Bio 3:65–87
Acknowledgments
This study was partially supported by a Grant-in-Aid for Scientific Research (B) (no. 20310045).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Honda, T., Takahashi, H., Sako, Y. et al. Gene expression of Microcystis aeruginosa during infection of cyanomyovirus Ma-LMM01. Fish Sci 80, 83–91 (2014). https://doi.org/10.1007/s12562-013-0685-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12562-013-0685-7