Abstract
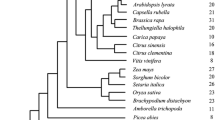
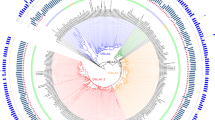
Gene duplication provides the key materials for new genes and novel functions. However, the mechanism underlying functional innovation remains unknown. In this study, we revealed the evolutionary pattern of the prenyltransferases of the UbiA gene family in 15 higher plants. Prenyltransferases of the UbiA gene family are involved in many important biological processes of both primary and secondary metabolism. Based on the phylogenetic relationships of the UbiA genes, seven subfamilies are classified. Confirming this classification, genes within each subfamily are characterized by similar exon numbers, exon lengths and patterns of motif combinations. Similar numbers of UbiA genes are found in different species within each subfamily except for Subfamily I, in which a Phaseoleae-specific expansion is detected in clade I-A. Homologous genes in clade I-A evolve rapidly, exchange sequences frequently and experience positive selection. Genes in clade I-A function as flavonoid prenyltransferase synthesis secondary compounds, while other genes from Subfamily I encode homogentisate phytyltransferase, which plays a role in primary metabolism. Thus, our results suggest that the secondary metabolism genes acquire new functions from those of primary metabolism through gene duplication and neofunctionalization driven by positive selection.




Similar content being viewed by others
References
Akashi T, Sasaki K, Aoki T, Ayabe S, Yazaki K (2009) Molecular cloning and characterization of a cDNA for pterocarpan 4-dimethylallyltransferase catalyzing the key prenylation step in the biosynthesis of glyceollin, a soybean phytoalexin. Plant Physiol 149:683–693
Assis R, Bachtrog D (2013) Neofunctionalization of young duplicate genes in Drosophila. Proc Natl Acad Sci U S A 110:17409–17414
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208
Bonitz T, Alva V, Saleh O, Lupas AN, Heide L (2011) Evolutionary relationships of microbial aromatic prenyltransferases. PLoS ONE 6:e27336
Botta B, Monache GD, Menendez P, Boffi A (2005) Novel prenyltransferase enzymes as a tool for flavonoid prenylation. Trends Pharmacol Sci 26:606–608
Eckhardt U, Grimm B, Hortensteiner S (2004) Recent advances in chlorophyll biosynthesis and breakdown in higher plants. Plant Mol Biol 56:1–14
Gaubier P, Wu HJ, Laudie M, Delseny M, Grellet F (1995) A chlorophyll synthetase gene from Arabidopsis thaliana. Mol Gen Genet 249:58–64
Guo AY, Zhu QH, Chen X, Luo JC (2007) GSDS: a gene structure display server. Yi Chuan 29:1023–1026
Han MV, Demuth JP, McGrath CL, Casola C, Hahn MW (2009) Adaptive evolution of young gene duplicates in mammals. Genome Res 19:859–867
Heide L (2009) Prenyl transfer to aromatic substrates: genetics and enzymology. Curr Opin Chem Biol 13:171–179
Holub EB (2001) The arms race is ancient history in Arabidopsis, the wildflower. Nat Rev Genet 2:516–527
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator v3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinform 2008:420747
Kondrashov FA (2012) Gene duplication as a mechanism of genomic adaptation to a changing environment. Proc Biol Sci 279:5048–5057
Lavin M, Herendeen PS, Wojciechowski MF (2005) Evolutionary rates analysis of Leguminosae implicates a rapid diversification of lineages during the tertiary. Syst Biol 54:575–594
Li C, Zhang YM (2011) Molecular evolution of glycinin and beta-conglycinin gene families in soybean (Glycine max L. Merr.). Heredity 106:633–641
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Lynch M (2000) The evolutionary fate and consequences of duplicate genes. Science 290:1151–1155
Lynch M, Crease TJ (1990) The analysis of population survey data on DNA sequence variation. Mol Biol Evol 7:377–394
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Ober D (2005) Seeing double: gene duplication and diversification in plant secondary metabolism. Trends Plant Sci 10:444–449
Ohara K, Yamamoto K, Hamamoto M, Sasaki K, Yazaki K (2006) Functional characterization of OsPPT1, which encodes p-hydroxybenzoate polyprenyltransferase involved in ubiquinone biosynthesis in Oryza sativa. Plant Cell Physiol 47:581–590
Okada K, Ohara K, Yazaki K, Nozaki K, Uchida N, Kawamukai M, Nojiri H, Yamane H (2004) The AtPPT1 gene encoding 4-hydroxybenzoate polyprenyl diphosphate transferase in ubiquinone biosynthesis is required for embryo development in Arabidopsis thaliana. Plant Mol Biol 55:567–577
Sadre R, Gruber J, Frentzen M (2006) Characterization of homogentisate prenyltransferases involved in plastoquinone-9 and tocochromanol biosynthesis. FEBS Lett 580:5357–5362
Saiki K, Mogi T, Ogura K, Anraku Y (1993) In vitro heme O synthesis by the cyoE gene product from Escherichia coli. J Biol Chem 268:26041–26044
Saleh O, Haagen Y, Seeger K, Heide L (2009) Prenyl transfer to aromatic substrates in the biosynthesis of aminocoumarins, meroterpenoids and phenazines: the ABBA prenyltransferase family. Phytochemistry 70:1728–1738
Sasaki K, Mito K, Ohara K, Yamamoto H, Yazaki K (2008) Cloning and characterization of naringenin 8-prenyltransferase, a flavonoid-specific prenyltransferase of Sophora flavescens. Plant Physiol 146:1075–1084
Sasaki K, Tsurumaru Y, Yamamoto H, Yazaki K (2011) Molecular characterization of a membrane-bound prenyltransferase specific for isoflavone from Sophora flavescens. J Biol Chem 286:24125–24134
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Shen G, Huhman D, Lei Z, Snyder J, Sumner LW, Dixon RA (2012) Characterization of an isoflavonoid-specific prenyltransferase from Lupinus albus. Plant Physiol 159:70–80
Shimada H, Ohno R, Shibata M, Ikegami I, Onai K, Ohto MA, Takamiya K (2005) Inactivation and deficiency of core proteins of photosystems I and II caused by genetical phylloquinone and plastoquinone deficiency but retained lamellar structure in a T-DNA mutant of Arabidopsis. Plant J 41:627–637
Sun X, Zhang Y, Yang S, Chen JQ, Hohn B, Tian D (2008) Insertion DNA promotes ectopic recombination during meiosis in Arabidopsis. Mol Biol Evol 25:2079–2083
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tian L, DellaPenna D, Dixon RA (2007) The pds2 mutation is a lesion in the Arabidopsis homogentisate solanesyltransferase gene involved in plastoquinone biosynthesis. Planta 226:1067–1073
Tian D, Wang Q, Zhang P, Araki H, Yang S, Kreitman M, Nagylaki T, Hudson R, Bergelson J, Chen JQ (2008) Single-nucleotide mutation rate increases close to insertions/deletions in eukaryotes. Nature 455:105–108
Venkatesh TV, Karunanandaa B, Free DL, Rottnek JM, Baszis SR, Valentin HE (2006) Identification and characterization of an Arabidopsis homogentisate phytyltransferase paralog. Planta 223:1134–1144
Wang J, Tan S, Zhang L, Li P, Tian D (2011) Co-variation among major classes of LRR-encoding genes in two pairs of plant species. J Mol Evol 72:498–509
Xu G, Ma H, Nei M, Kong H (2009) Evolution of F-box genes in plants: different modes of sequence divergence and their relationships with functional diversification. Proc Natl Acad Sci U S A 106:835–840
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15:568–573
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Nielsen R (2002) Codon-substitution models for detecting molecular adaptation at individual sites along specific lineages. Mol Biol Evol 19:908–917
Yang S, Feng Z, Zhang X, Jiang K, Jin X, Hang Y, Chen JQ, Tian D (2006) Genome-wide investigation on the genetic variations of rice disease resistance genes. Plant Mol Biol 62:181–193
Yang S, Jiang K, Araki H, Ding J, Yang YH, Tian D (2007) A molecular isolation mechanism associated with high intra-specific diversity in rice. Gene 394:87–95
Yazaki K, Kunihisa M, Fujisaki T, Sato F (2002) Geranyl diphosphate: 4-hydroxybenzoate geranyltransferase from Lithospermum erythrorhizon. Cloning and characterization of a ket enzyme in shikonin biosynthesis. J Biol Chem 277:6240–6246
Yazaki K, Sasaki K, Tsurumaru Y (2009) Prenylation of aromatic compounds, a key diversification of plant secondary metabolites. Phytochemistry 70:1739–1745
Zhang J, Nielsen R, Yang Z (2005) Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol 22:2472–2479
Acknowledgments
This work was supported in part by the National Natural Science Foundation of China (31301342, 31370034), Key Transgenic Breeding Program of China (2013ZX08004-003), Priority Academic Program Development of Jiangsu Higher Education Institutions Jiangsu Provincial Program (PAPD) and the Nanjing Agricultural University Youth Science and Technology Innovation Fund (KJ2013002).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, J., Chu, S., Zhu, Y. et al. Positive selection drives neofunctionalization of the UbiA prenyltransferase gene family. Plant Mol Biol 87, 383–394 (2015). https://doi.org/10.1007/s11103-015-0285-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0285-2