Abstract
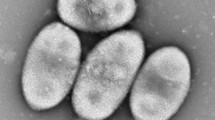
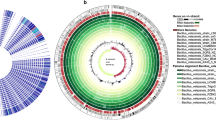
Microbially synthesized nanoparticles has received increasing attentions owing to the broad applications in biology and medicine. In this study, we report a novel bacterium that biologically generates silver nanoparticles (AgNPs). This bacterium, designated strain F202Z8T, was isolated from a rusty iron plate found in the intertidal region of Taean, South Korea. The morphological, biochemical and molecular characteristics predicted that strain F202Z8T belongs to the family Flavobacteriaceae. Phylogenetic analysis based on 16S rRNA gene sequence revealed that strain F202Z8T forms a distinct lineage with closely related genera Maribacter, Pelagihabitans, Pseudozobellia, Zobellia, Pricia, and Costertonia and showed the highest similarity to Maribacter aestuarii GY20T (94.5%). The digital DNA–DNA hybridization and average nucleotide identity values calculated from the whole genome-sequence comparison between strain F202Z8T and other members of the family Flavobacteriaceae were in the ranges of 12.7%–16.9% and 70.3%–74.4%, respectively, suggesting that strain F202Z8T represented a novel genus. The complete genome sequence of strain F202Z8T is 4,723,614 bp, with 43.26% G + C content. Based on the COG, GO, KEGG, NR, and Swiss-Prot databases, the genomic analysis of F202Z8T revealed the presence of 17 putative genes responsible for the synthesis of AgNPs. Our polyphasic taxonomic results suggested that this strain represents a novel species of a novel genus in the family Flavobacteriaceae, for which the name Aggregatimonas sangjinii gen. nov., sp. nov. is proposed. The type strain of Aggregatimonas sangjinii is F202Z8T (= KCCM 43411T = LMG 31494T). Overall, our data provide fundamental information to potentially utilize this novel bacterium for synthesis of nanoparticles.

Similar content being viewed by others
References
Ashburner M, Ball CA, Blake JA et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium Nat Genet 25:25–29. https://doi.org/10.1038/75556
Bairoch A, Apweiler R (2000) The SWISS-PROT protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res 28:45–48. https://doi.org/10.1093/nar/28.1.45
Barbeyron T, L’Haridon S, Corre E, Kloareg B, Potin P (2001) Zobellia galactanovorans gen. nov., sp. nov., a marine species of Flavobacteriaceae isolated from a red alga, and classification of [Cytophaga] uliginosa (ZoBell and Upham 1944) Reichenbach 1989 as Zobellia uliginosa gen. nov., comb. nov. Int J Syst Evol Microbiol 51:985–997. https://doi.org/10.1099/00207713-51-3-985
Bernardet JF, Nakagawa Y, Holmes B (2002) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52:1049-1070. https://doi.org/10.1099/00207713-52-3-1049
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:W81–W87. https://doi.org/10.1093/nar/gkz310
Bowman JP (2006) The marine clade of the family Flavobacteriaceae: the genera Aequorivita, Arenibacter, Cellulophaga, Croceibacter, Formosa, Gelidibacter, Gillisia, Maribacter, Mesonia, Muricauda, Polaribacter, Psychroflexus, Psychroserpens, Robiginitalea, Salegentibacter, Tenacibaculum, Ulvibacter, Vitellibacter, and Zobellia. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, 3rd edn. Springer, New York, pp 677–694
Collins MD (1985) Isoprenoid quinone analysis in bacterial classification and identification. In: Minnikin DE (ed) Goodfellow M. Chemical methods in bacterial systematics Academic Press, London, pp 267–287
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Galperin MY, Makarova KS, Wolf YI, Koonin EV (2015) Expanded microbial genome coverage and improved protein family annotation in the COG database. Nucleic Acids Res 43:D261–D269. https://doi.org/10.1093/nar/gku1223
Garrity GM, Holt JG (2001) The road map to the manual. In: Garrity GM, Boone DR, Castenholz RW (eds) Bergey’s manual of systematic bacteriology, 2nd edn. Springer, New York, pp 119–166
Grubbs KJ, Bleich RM, Santa Maria KC, Allen SE, Farag S, AgBiome Team, Shank EA, Bowers AA (2017) Large-scale bioinformatics analysis of Bacillus genomes uncovers conserved roles of natural products in Bacterial physiology. mSystems 2 e00040–17. https://doi.org/10.1128/mSystems.00040-17
Hu J, Yang QQ, Ren Y, Zhang WW, Zheng G, Sun C, Pan J, Zhu X-F, Zhang X-Q, Wu M (2015) Maribacter thermophilus sp. nov., isolated from an algal bloom in an intertidal zone, and emended description of the genus Maribacter. Int J Syst Evol Microbiol 65:36–41. https://doi.org/10.1099/ijs.0.064774-0
Kanehisa M, Sato Y, Morishima K (2016) BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 428:726–731. https://doi.org/10.1016/j.jmb.2015.11.006
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Kluge AG, Farris JS (1969) Quantitative phyletics and the evolution of anurans. Syst Biol 18:1–32. https://doi.org/10.1093/sysbio/18.1.1
Kumar A, Vemula PK, Ajayan PM, John G (2008) Silver-nanoparticle-embedded antimicrobial paints based on vegetable oil. Nat Mater 7:236–241. https://doi.org/10.1038/nmat2099
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kwon KK, Lee YK, Lee HK (2006) Costertonia aggregata gen. nov., sp. nov., a mesophilic marine bacterium of the family Flavobacteriaceae, isolated from a mature biofilm. Int J Syst Evol Microbiol 56:1349–1353. https://doi.org/10.1099/ijs.0.64168-0
Kwon YM, Kim KW, Kim JYH, Choi T-Y, Yang S-H, Oh CH, Kwon KK, Kim S-J (2018) Euzebyella algicola sp. nov., a marine bacterium of the family Flavobacteriaceae, isolated from green algae. Int J Syst Evol Microbiol 68:764–768. https://doi.org/10.1099/ijsem.0.002581
Kwon YM, Yang SH, Kwon KK, Kim SJ (2014) Nonlabens antarcticus sp. nov., a psychrophilic bacterium isolated from glacier ice, and emended descriptions of Nonlabens marinus Park et al 2012 and Nonlabens agnitus Yi and Chun 2012. Int J Syst Evol Microbiol 64:400–405. https://doi.org/10.1099/ijs.0.056606-0
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Li S, Niu Y, Chen H, He P (2021) Complete genome sequence of an Arctic Ocean bacterium Shewanella sp. Arc9-LZ with capacity of synthesizing silver nanoparticles in darkness. Mar Genomics 56:100808. https://doi.org/10.1016/j.margen.2020.100808
Li S, Zhang Z, Gu X, Lin X (2020) Complete genome sequence of paracoccus sp. Arc7-R13, A silver nanoparticles synthesizing bacterium isolated from arctic ocean sediments. Mar Genomics 50:100694. https://doi.org/10.1016/j.margen.2019.100694
Li W, Jaroszewski L, Godzik A (2002) Tolerating some redundancy significantly speeds up clustering of large protein databases. Bioinformatics 18:77–82. https://doi.org/10.1093/bioinformatics/18.1.77
Lü JM, Wang X, Marin-Muller C, Wang H, Lin PH, Yao Q, Chen C (2009) Current advances in research and clinical applications of PLGA-based nanotechnology. Expert Rev Mol Diagn 9:325–341. https://doi.org/10.1586/erm.09.15
McBride MJ (2014) The family Flavobacteriaceae. In: Rosenberg E, Delong EF, Lory S, Stackebrandnt E, Thonpson F (eds) The prokaryotes, 4th edn. Springer, Berlin, pp 643–676
Meier-Kolthoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60. https://doi.org/10.1186/1471-2105-14-60
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaala A, Parletta JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Na S, Kim YO, Yoon S, Ha S, Baek I, Chun J (2018) UBCG: Up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:280–285. https://doi.org/10.1007/s12275-018-8014-6
Nedashkovskaya OI, Suzuki M, Lee JS, Lee KC, Shevchenko LS, Mikhailov VV (2009) Pseudozobellia thermophila gen. nov., sp. nov., a bacterium of the family Flavobacteriaceae, isolated from the green alga Ulva fenestrata. Int J Syst Evol Microbiol 59:806–810. https://doi.org/10.1099/ijs.0.004143-0
Nedashkovskaya OI, Suzuki M, Vancanneyt M, Cleenwerck I, Lysenko AM, Mikhailov VV, Swings J (2004) Zobellia amurskyensis sp. nov., Zobellia laminariae sp. nov. and Zobellia russellii sp. nov., novel marine bacteria of the family Flavobacteriaceae. Int J Syst Evol Microbiol 59:806–810. https://doi.org/10.1099/ijs.0.63091-0
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW (2015) CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25:1043–1055. https://doi.org/10.1101/gr.186072.114
Patil MP, Kim G (2018) Marine microorganisms for synthesis of metallic nanoparticles and their biomedical applications. Colloids Surf B 172:487–495. https://doi.org/10.1016/j.colsurfb.2018.09.007
Reichenbach H (1992) The order Cytophagales. In: Trüper HG, Dworkin M, Harder W, Schleifer KH (eds) Balows A. The prokaryotes Springer, New York, pp 3631–3679
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sastry RK, Rashmi HB, Rao NH, Ilyas SM (2010) Integrating nanotechnology into agri-food systems research in India: a conceptual framework. Technol Forecast Soc Change 77:639–648. https://doi.org/10.1016/j.techfore.2009.11.008
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Murray RGE, Wood WA, Krieg NR (eds) Gerhardt P. Methods for general and molecular bacteriology American Society for Microbiology, Washington DC, pp 607–654
Srivastava SK, Constanti M (2012) Room temperature biogenic synthesis of multiple nanoparticles (Ag, Pd, Fe, Rh, Ni, Ru, Pt Co, and Li) by Pseudomonas aeruginosa SM1. J Nanopart Res 14:831. https://doi.org/10.1007/s11051-012-0831-7
Stackebrandt E, Frederiksen W, Garrity GM et al (2002) Report of the ad hoc committee for the re-evaluation of the species definition in bacteriology. Int J Syst Evol Microbiol 52:1043–1047. https://doi.org/10.1099/00207713-52-3-1043
Sun C, Yang Z, Zhang C, Liu Z, He J, Liu Q, Zhang T, Ju J, Ma J (2019) Genome mining of Streptomyces atratus SCSIO ZH16: discovery of atratumycin and identification of its biosynthetic gene cluster. Org Lett 21:1453–1457. https://doi.org/10.1021/acs.orglett.9b00208
Wang Q, Liu F, Zhang DC (2020) Pelagihabitans pacificus gen. nov., sp. nov., a member of the family Flavobacteriaceae isolated from a deep-sea seamount. Int J Syst Evol Microbiol 70:4569–4575. https://doi.org/10.1099/ijsem.0.004315
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yu J, Zhang L, Qian J, Zhuc Z, Nid S, Liu G, Xu X (2019) In situ exsolution of silver nanoparticles on AgTaO3-SrTiO3 solid solutions as efficient plasmonic photocatalysts for water splitting. Appl Catal B 256:117818. https://doi.org/10.1016/j.apcatb.2019.117818
Yu Y, Li HR, Zeng YX, Sun K, Chen B (2012) Pricia antarctica gen. nov., sp. nov., a member of the family Flavobacteriaceae, isolated from Antarctic intertidal sediment. Int J Syst Evol Microbiol 62:2218–2223. https://doi.org/10.1099/ijs.0.037515-0
Zhang D, Ma XL, Gu Y, Huang H, Zhang GW (2020) Green synthesis of metallic nanoparticles and their potential applications to treat cancer. Front Chem 8:799. https://doi.org/10.3389/fchem.2020.00799
Acknowledgements
We thank Professor Aharon Oren for his help with etymology. This work was supported by a grant from the National Marine Biodiversity Institute of Korea (2022M00300).
Funding
This work was supported by a grant from the National Marine Biodiversity Institute of Korea (2022M00300).
Author information
Authors and Affiliations
Contributions
DC and YMK performed all experiments, data analysis, and manuscript finalization. JJ performed additional phylogenomic analysis and interpretation. JYHK and KWK participated in the review of the manuscript and isolation of this strain. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Data availability
All data generated or analyzed during this study are included in this manuscript and its supplementary information files.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chung, D., Jung, J., Kim, J.Y.H. et al. Aggregatimonas sangjinii gen. nov., sp. nov., a novel silver nanoparticle synthesizing bacterium belonging to the family Flavobacteriaceae. Antonie van Leeuwenhoek 115, 325–335 (2022). https://doi.org/10.1007/s10482-021-01700-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01700-w