Abstract
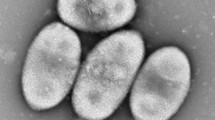
Chemo-organotrophic iodide (I−)-oxidizing bacterial strains Hi-2T and Mie-1 were isolated from iodide-rich natural gas brine water in Chiba and surface seawater in Mie, Japan, respectively. Cells of strains Hi-2T and Mie-1 were aerobic, Gram-negative and rod-shaped (0.3–0.5 µm width and 1.2–4.4 µm in length). Two isolates grew optimally at 30 °C, pH 7.5 and with 3% NaCl (w/v). Iodide oxidation to form molecular iodine (I2) was a biochemically unique trait for strains Hi-2T and Mie-1. The major cellular fatty acids are C18:1ω7c, C16:1ω5c and C18:1 2-OH. A phylogenetic analysis based on the 16S rRNA gene sequence revealed that strains Hi-2T and Mie-1 were located near Iodidimonas muriae C-3T with 99.2% sequence similarity. The calculated digital DNA–DNA hybridization (dDDH) value of 65.7–65.9% between the two isolates and I. muriae C-3T was lower than the threshold of 70%, which was used for prokaryotic species delineation. Strains Hi-2T and Mie-1 differed in the hydrolysis of aesculin, the hydrolysis of gelatin and the major cellular fatty acids composition from I. muriae C-3T. Considering these biochemical properties, the major cellular fatty acids composition and dDDH value, a novel species is proposed for strains Hi-2T (= JCM 17844T = LMG 28661T) and Mie-1 (= JCM 17845 = LMG 28662), to be named Iodidimonas gelatinilytica.

Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in this published article.
References
Amachi S, Muramatsu Y, Akiyama Y, Miyazaki K, Yoshiki S, Hanada S, Kamagata Y, Ban-nai T, Shinoyama H, Fujii T (2005) Isolation of iodide-oxidizing bacteria from iodide-rich natural gas brines and seawaters. Microb Ecol 49:547–557. https://doi.org/10.1007/s00248-004-0056-0
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Edberg SC, Pittman S, Singer JM (1977) Esculin hydrolysis by Enterobacteriaceae. J Clin Microbiol 6:111–116
Ehara A, Suzuki H, Kanesaki Y, Yoshikawa H, Amachi S (2014) Draft genome sequence of strain Q-1, an iodide-oxidizing alphaproteobacterium isolated from natural gas brine water. Genome Announc 2:e00659-e714. https://doi.org/10.1128/genomeA.00659-14
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376. https://doi.org/10.1007/BF01734359
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91. https://doi.org/10.1099/ijs.0.64483-0
Gozlan RS (1968) Isolation of iodine-producing bacteria from aquaria. Antonie Van Leeuwenhoek 34:226. https://doi.org/10.1007/BF02046433
Gozlan RS, Margalith P (1973) Iodide oxidation by a marine bacterium. J Appl Bact 36:407–417. https://doi.org/10.1111/j.1365-2672.1973.tb04122.x
Gozlan RS, Margalith P (1974) Iodide oxidation by Pseudomonas iodooxidans. J Appl Bact 37:493–499. https://doi.org/10.1111/j.1365-2672.1974.tb00474.x
Hasegawa M, Kishino H, Yano TA (1985) Dating of the human ape splitting by a molecular clock of mitochondrial-DNA. J Mol Evol 22:160–174. https://doi.org/10.1007/BF02101694
Hucker GJ, Conn HJ (1923). Methods of gram staining. New York State Agricultural Experiment Station. Technical Bulletin No. 93, Ithaca, pp 3–37
Iino T, Ohkuma M, Kamagata Y, Amachi S (2016) Iodidimonas muriae gen. nov., sp. nov., an aerobic iodide-oxidizing bacterium isolated from brine of a natural gas and iodine recovery facility, and proposals of Iodidimonadaceae fam. nov., Iodidimonadales ord. nov., Emcibacteraceae fam. nov. and Emcibacterales ord. nov. Int J Syst Evol Microbiol 66:5016–5022. https://doi.org/10.1099/ijsem.0.001462
Komagata K, Suzuki K (1987) Lipid and cell-wall analysis in bacterial systematics. Methods Microbiol 19:161–208. https://doi.org/10.1016/S0580-9517(08)70410-0
Lechevalier MP, De Bievre C, Lechevalier H (1977) Chemotaxonomy of aerobic actinomycetes: phospholipid composition. Biochem Syst Ecol 5:249–260. https://doi.org/10.1016/0305-1978(77)90021-7
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar HM, Buchner A, Lai T, Steppi S, Jobb G, Förster W, Brettske I, Gerber S, Ginhart AW, Gross O, Grumann S, Hermann S, Jost R, König A, Liss T, Lüssmann R, May M, Nonhoff B, Reichel B, Strehlow R, Stamatakis A, Stuckmann N, Vilbig A, Lenke M, Ludwig T, Bode A, Schleifer K-H (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32:1363–1371. https://doi.org/10.1093/nar/gkh293
Meier-Kothoff JP, Auch AF, Klenk HP, Göker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. https://doi.org/10.1186/1471-2105-14-60
Meier-Kolthoff JP, Hahnke RL, Petersen J, Scheuner C, Michael V, Fiebig A, Rohde C, Rohde M, Fartmann B, Goodwin LA, Chertkov O, Reddy T, Pati A, Ivanova NN, Markowitz V, Kyrpides NC, Woyke T, Göker M, Klenk H-P (2014) Complete genome sequence of DSM 30083(T), the type strain (U5/41(T)) of Escherichia coli, and a proposal for delineating subspecies in microbial taxonomy. Stand Genomic Sci 9:2. https://doi.org/10.1186/1944-3277-9-2
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Schaal AM, A, Parlett JH, (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Moriya Y, Itho M, Okuda S, Yoshizawa AC, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:W182–W185. https://doi.org/10.1093/nar/gkm321
Pickett MJ, Greenwood JR, Harvey SM (1991) Test for detecting degradation of gelatin: comparison of five methods. J Clin Microbiol 29:2322–2325. https://doi.org/10.2527/2001.791261x
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. MIDI Inc., Newark, DE
Suzuki M, Eda Y, Ohsawa S, Kanesaki Y, Yoshikawa H, Tanaka K, Muramatsu Y, Yoshikawa J, Sato I, Fujii T, Amachi S (2012) Iodide oxidation by a novel multicopper oxidase from the alphaproteobacterium strain Q-1. Appl Environ Microbiol 78:3941–3949. https://doi.org/10.1128/AEM.00084-12
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The Clustal_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tool. Nucleic Acids Res 24:4876–4882. https://doi.org/10.1093/nar/25.24.4876
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer K-H, Whitman WB, Euzéby J, Amann R, Rosselló-Móra R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645. https://doi.org/10.1038/nrmicro3330
Acknowledgements
We thank Prof. Aahron Oren (The Hebrew University of Jerusalem) for helpful corrections of the proposed name and its etymology.
Funding
This work was supported by a Grant-in-Aid from the Japan Society for the Promotion of Science (Grant No. 19H05689).
Author information
Authors and Affiliations
Contributions
TI carried out all morphological, phenotypic and chemotaxonomic characterisation, constructed the 16S rRNA gene phylogenetic trees, did genomic analyses and drafted the original manuscript. KO and MH determined genome sequence of strains C-3T, Hi-2T and Mie-1. MO supervised the research and provided the research funding. SA isolated strains Hi-2T and Mie-1 and carried out the primal characterisation. All authors proofread the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Iino, T., Oshima, K., Hattori, M. et al. Iodidimonas gelatinilytica sp. nov., aerobic iodide-oxidizing bacteria isolated from brine water and surface seawater. Antonie van Leeuwenhoek 114, 625–631 (2021). https://doi.org/10.1007/s10482-021-01546-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01546-2