Abstract
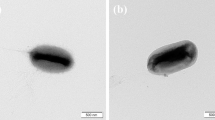
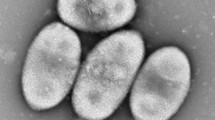
A taxonomic study was carried out on a novel aerobic bacterial strain (designated CC-LY736T) isolated from a fermentor in Taiwan. Cells of strain CC-LY736T were Gram-stain negative, spiral-shaped and motile by means of a monopolar flagellum. Strain CC-LY736T shared the greatest degree of 16S rRNA gene sequence similarity to Azospirillum irakense DSM 11586T (97.2 %), Rhodocista centenaria JCM 21060T (96.3 %) and Rhodocista pekingensis JCM 11669T (96.1 %). The major fatty acids were C16:0, C16:1 ω5c, C19:0 cyclo ω8c, C18:1 ω7c/C18:1 ω6c, C16:0 3-OH and C18:1 2-OH. The predominant polar lipids included phosphatidylcholine, phosphatidylglycerol, diphosphatidylglycerol, phosphatidylethanolamine, phosphatidylmethylethanolamine, phosphatidyldimethylethanolamine and two unidentified glycolipids. The common major respiratory quinone was ubiquinone Q-10 and predominant polyamines were sym-homospermidine and putrescine. The DNA G+C content of strain CC-LY736T was 67.6 ± 0.1 mol %. During phylogenetic analysis, strain CC-LY736T formed a unique phyletic lineage associated with Rhodocista species. However, the combination of genetic, chemotaxonomic and physiological data clearly indicated that strain CC-LY736T was a novel representative of the family Rhodospirillaceae. Based on the polyphasic comparison, the name Niveispirillum fermenti gen. nov., sp. nov. is proposed; the type strain of the type species is CC-LY736T (= BCRC 80504T = LMG 27263T). In addition, the reclassifications of Azospirillum irakense as Niveispirillum irakense comb. nov. (type strain KBC1T = ATCC 51182T = BCRC 15764T = CIP 103311T), and Azospirillum amazonense as Nitrospirillum amazonense gen. nov., sp. nov. (type strain Am14T = ATCC 35119T = BCRC 14279T = DSM 3787T) are proposed based on the polyphasic taxonomic data obtained in this study.

Similar content being viewed by others
References
Bally R, Thomas-Bauzon D, Heulin T, Balandreau J, Richard C, De Ley J (1983) Determination of the most frequent N2-fixing bacteria in a rice rhizosphere. Can J Microbiol 29:881–887
Coenye T, Goris J, Spilker T (2002) Characterization of unusual bacteria isolated from respiratory secretions of cystic fibrosis patients and description of Inquilinus limosus gen. nov., sp. nov. J Clin Microbiol 40:2062–2069
Collins MD (1985) Isoprenoid quinone analysis in classification and identification. In: Goodfellow M, Minnikin DE (eds) Chemical methods in bacterial systematics. Academic Press, London, pp 267–287
Döbereiner J, Day JM (1976) Associative symbioses in tropical grasses: characterization of microorganisms and dinitrogen-fixing sites. In: Newton WE, Nyman CJ (eds) Proceedings of the first international symposium on N2 fixation. Washington State University Press, Pullman, pp 518–538
Eckert B, Weber OB, Kirchhof G, Halbritter A, Stoffels M, Hartmann A (2001) Azospirillum doebereinerae sp. nov., a nitrogen-fixing bacterium associated with the C(4)-grass Miscanthus. Int J Sys Evol Microbiol 51:17–26
Edwards U, Rogall T, Blocker H, Emde M, Bottger EC (1989) Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res 17:7843–7853
Euzéby JP, Kudo T (2001) Corrigenda to the validation lists. Int J Sys Evol Microbiol 51:1933–1938
Falk EC, Döbereiner J, Johnson JL, Krieg NR et al (1985) Deoxyribonucleic acid homology of Azospirillum amazonense Magalhães, 1984 and emendation of the description of the genus Azospirillum. Int J Sys Bacteriol 35:117–118
Falk EC, Johnson JL, Baldani VLD, Döbereiner J, Krieg NR (1986) Deoxyribonucleic and ribonucleic acid homology studies of the genera Azospirillum and Conglomeromonas. Int J Sys Bacteriol 36:80–85
Favinger J, Stadtwald R, Gest H (1989) Rhodospirillum centenum, sp. nov., a thermotolerant cyst-forming anoxygenic photosynthetic bacterium. Antonie Van Leeuwenhoek 55:291–296
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Sys Zool 20:406–416
GCG (1995). Wisconsin package version 8.1 program manual. Genetics Computer Group, Madison
Gest H, Favinger J (2001) Taxonomic ambiguities: a case history. Int J Sys Evol Microbiol 51:707–710
Hardy R, Burns RC, Holsten RD (1973) Application of the acetylene-ethylene assay for measurement of nitrogen fixation. Soil Biol Biochem 5:47–81
Heiner CR, Hunkapiller LK, Chen SM, Glass JI, Chen EY (1998) Sequencing multimegabase-template DNA using BigDye terminator chemistry. Genome Res 8:557–561
Kawasaki H, Hoshino Y, Kuraiski Y, Yamasato K (1992) Rhodocista centenaria gen. nov., sp. nov., a cyst-forming anoxygenic photosynthetic bacterium and its phylogenetic position in the Proteobacteria alpha group. J Gen Appl Microbiol 38:541–551
Kawasaki H, Hoshino Y, Yamasato K (1993) Phylogenetic diversity of phototrophic purple non-sulfur bacteria in the Proteobacteria group. FEMS Microbiol Lett 112:61–66
Khammas KM, Ageron E, Grimont PAD, Kaiser P (1989) Azospirillum irakense sp. nov., a new nitrogen-fixing bacterium associated with rice roots and rhizosphere soil. Res Microbiol 140:679–693
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Sys Evol Microbiol 62:716–721
Kirchhof G, Reis VM, Baldan JI, Eckert B, Döbereiner J, Hartmann A (1997) Occurrence, physiological and molecular analysis of endophytic diazotrophic bacteria in gramineous energy plants. Plant Soil 194:45–55
Koch B, Evans HJ (1966) Reduction of acetylene to ethylene by soybean root nodules. Plant Physiol 41:1748–1750
Ladha JK, So RB, Watanabe I (1987) Composition of Azospirillum species associated with wetland rice plants grown in different soils. Plant Soil 102:127–129
Lavrinenko K, Chernousova E, Gridneva E, Dubinina G, Akimov V, Kuever J, Lysenko A, Grabovich M (2010) Azospirillum thiophilum sp. nov., a novel diazotrophic bacterium isolated from a sulfide spring. Int J Sys Evol Microbiol 60:2832–2837
Lin S-Y, Young C-C, Hupfer H, Siering C, Arun AB, Chen W-M, Lai W-A, Shen F-T, Rekha PD, Yassin AF (2009) Azospirillum picis sp. nov., isolated from discarded tar. Int J Sys Evol Microbiol 59:761–765
Lin S-Y, Shen F-T, Young C-C (2011) Rapid detection and identification of the free-living nitrogen fixing genus Azospirillum by 16S rRNA-gene-targeted genus-specific primers. Antonie Van Leeuwenhoek 99:837–844
Lin S-Y, Shen F-T, Young L-S, Zhu Z-L, Chen W-M, Young C-C (2012) Azospirillum formosense sp. nov., a novel diazotrophic bacterium isolated from agricultural soil. Int J Sys Evol Microbiol 62:1185–1190
Lin S-Y, Liu Y-C, Hameed A, Hsu Y-H, Lai W-A, Shen F-T, Young C–C (2013) Azospirillum fermentarium sp. nov., a nitrogen-fixing species isolated from a fermenter. Int J Sys Evol Microbiol 63:3762–3768
Magalhães FM, Baldani JL, Souto SM, Kuykendall JR, Döbereiner J (1983) A new acid-tolerant Azospirillum species. An Acad Bras Cien 55:417–430
Mehnaz S, Weselowski B, Lazarovits G (2007a) Azospirillum canadense sp. nov., a nitrogen-fixing bacterium isolated from corn rhizosphere. Int J Sys Evol Microbiol 57:620–624
Mehnaz S, Weselowski B, Lazarovits G (2007b) Azospirillum zeae sp. nov., a diazotrophic bacterium isolated from rhizosphere soil of Zea mays. Int J Sys Evol Microbiol 57:2805–2809
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G+C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Sys Bacteriol 39:159–167
Miller LT (1982) Single derivatization method for routine analysis of bacterial whole-cell fatty acid methyl esters, including hydroxyl acids. J Clin Microbiol 16:584–586
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal K, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Murray RGE, Doetsch RN, Robinow CF (1994) Methods for general and molecular bacteriology. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Determination and cytological light microscopy. American Society for Microbiology, Washington, DC, pp 31–32
Ostle AG, Holt JG (1982) Nile blue A as a fluorescent stain for poly-β-hydroxybutyrate. Appl Environ Microbiol 44:238–241
Paisley R (1996) MIS whole cell fatty acid analysis by gas chromatography training manual. MIDI, Newark
Patriquin DG, Döbereiner J, Jain DK (1983) Sites and processes of association between diazotrophs and grasses. Can J Microbiol 29:900–915
Pfennig N, Lünsdorf H, Süling J, Imhoff JF (1997) Rhodospira trueperi, gen. nov., sp. nov., a new phototrophic Proteobacterium of the alpha group. Arch Microbiol 168:39–45
Poly F, Monrozier LJ, Bally R (2001) Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil. Res Microbiol 152:95–103
Reinhold B, Hurek T, Fendrik I, Pot B, Gillis M, Kersters K, Thielemans S, Ley JD (1987) Azospirillum halopraeferens sp. nov., a nitrogen-fixing organism associated with roots of Kallar grass (Leptochloa fusca (L.) Kunth). Int J Sys Bacteriol 37:43–51
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical note 101, MIDI Inc, Newark
Scherer P, Kneifel H (1983) Distribution of polyamines in methanogenic bacteria. J Bacteriol 154:1315–1322
Schlegel HG, Lafferty R, Krauss I (1970) The isolation of mutants not accumulating poly-β-hydroxybutyric acid. Arch Microbiol 71:283–294
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tarrand JJ, Krieg NR, Döbereiner J (1978) A taxonomic study of the Spirillum lipoferum group, with descriptions of a new genus, Azospirillum gen. nov., and two species, Azospirillum lipoferum (Beijerinck) comb. nov. and Azospirillum brasilense sp. nov. Can J Microbiol 24:967–980
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Trüper HG, Imhoff JF (1989) Bergey’s manual of systematic bacteriology. In: Staley JT, Bryant MP, Pfennig N, Holt JG (eds) Genus Rhodospirillum. Williams and Wilkins, Baltimore, pp 1662–1666
Watts D, MacBeath JR (2001) Automated fluorescent DNA sequencing on the ABI PRISM 310 genetic analyzer. Meth Mol Biol 167:153–170
Wayne LG, Brenner DJ, Colwell RR, Grimont PAD, Kandler O, Krichevsky MI, Moore LH, Moore WEC, Murray RGE et al (1987) International committee on systematic bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int J Sys Bacteriol 37:463–464
Weon H-Y, Kim B-Y, Hong S-B, Joa J-H, Nam S-S, Lee K-H, Kwon S-W (2007) Skermanella aerolata sp. nov., isolated from air, and emended description of the genus Skermanella. Int J Sys Evol Microbiol 57:1539–1542
Xie CH, Yokota A (2005) Azospirillum oryzae sp. nov., a nitrogen-fixing bacterium isolated from the roots of the rice plant Oryza sativa. Int J Sys Evol Microbiol 55:1435–1438
Young C-C, Hupfer H, Siering C, Ho M-J, Arun AB, Lai W-A, Rekha PD, Shen F-T, Hung M-H, Chen W-M, Yassin AF (2008) Azospirillum rugosum sp. nov., isolated from oil-contaminated soil. Int J Sys Evol Microbiol 58:959–963
Zehr JP, McReynolds LA (1989) Use of degenerate oligonucleotides for amplification of the nifH gene from the marine cyanobacterium Trichodesmium thiebautii. Appl Environ Microbiol 55:2522–2526
Zhang D, Yang H, Zhang W, Huang Z, Liu S-J (2003) Rhodocista pekingensis sp. nov., a cyst-forming phototrophic bacterium from a municipal wastewater treatment plant. Int J Sys Evol Microbiol 53:1111–1114
Acknowledgments
Authors would like to thank Mr. Wen-Shao Yen for the isolation of this novel bacterium. This research work was kindly supported by grants from the National Science Council, the Council of Agriculture, Executive Yuan and in part by the Ministry of Education, Taiwan, ROC under the ATU plan.
Author information
Authors and Affiliations
Corresponding author
Additional information
The GenBank/EMBL/DDBJ accession number for the 16S rRNA sequence of strain CC-LY736T is JX843283. In this study, the GenBank/EMBL/DDBJ accession number for the 16S rRNA sequence of strains Azospirillum amazonense BCRC 14279T, A. halopraeferens DSM 3675T, A. irakense DSM 11586T, A. lipoferum DSM 1691T, A. canadense LMG 23617T were GU256437, GU256439, GU256440, GU256441 and HM636056, respectively. A transmission electron micrograph of cells of strain CC-LY736T, polar lipid profile and polyamine compositions of all strains used in this study are available as supplementary material with the online version of this paper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lin, SY., Hameed, A., Shen, FT. et al. Description of Niveispirillum fermenti gen. nov., sp. nov., isolated from a fermentor in Taiwan, transfer of Azospirillum irakense (1989) as Niveispirillum irakense comb. nov., and reclassification of Azospirillum amazonense (1983) as Nitrospirillum amazonense gen. nov. Antonie van Leeuwenhoek 105, 1149–1162 (2014). https://doi.org/10.1007/s10482-014-0176-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-014-0176-6