Abstract
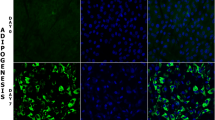
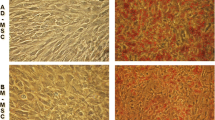
Alterations in the nuclear positioning of chromosomes and specific genes during differentiation and development have suggested strongly the existence of a relationship between non-random organization of the genome and its function. In this study, we have examined the genome organization in interphase nuclei during adipogenesis, using the pig as a model organism. We hypothesized that changes in the gene expression profile and chromatin remodeling which occur during cellular differentiation would elicit repositioning of whole chromosomes, moving specific genes on them to different regions of the nucleus. We established an in vitro adipogenesis differentiation system using mesenchymal stem cells, derived from porcine bone marrow. The nuclear position of seven adipogenesis genes (PPARG, SREBF1, FABP4, CEBPA, CEBPB, CREB, and GATA2), two control genes (SOX9 and MYL1), and six chromosomes carrying these gene loci (SSC4, SSC6, SSC12, SSC13, SSC15, and SSC17) was determined. We found that during adipogenesis, using the in vitro stem cell model system, in contrast to our original hypothesis, the nuclear position of genes involved in adipogenesis was altered radically with the up-regulation of gene expression correlating with these genes becoming more internally located within nuclei. Chromosome territories, containing these genes, were also found to alter their nuclear position during the in vitro adipogenesis model, with the most dramatic repositioning being SSC4 that moved from the nuclear periphery towards the nuclear interior. We found that during in vitro adipogenesis chromosome territories decondensed and the genes were found on loops and projections of chromatin, away from the main body of the chromosomes. From our data, it appears that the temporal repositioning of genes, emanating away from chromosomes, during adipogenesis is correlated with gene activity, supporting models of the involvement of spatial genome repositioning in regulating gene expression and the nuclear interior being an important region of the nucleus for transcription.









Similar content being viewed by others
References
Anderson SI, Lopez-Corrales NL, Gorick B, Archibald AL (2000) A large-fragment porcine genomie library resource in a BAC vector. Mamm Genome 9:811–814
Ballabio E, Cantarella CD, Federico C, Di Mare P, Hall G, Harbott J, Hughes J, Saccone S, Tosi S (2009) Ectopic expression of the HLXB9 gene is associated with an altered nuclear position in t(7;12) leukaemias. Leukemia 23(6):1179–1182
Bártová E, Kozubek S, Jirsová P, Kozubek M, Gajová H, Lukásová E, Skalníková M, Ganová A, Koutná I, Hausmann M (2002) Nuclear structure and gene activity in human differentiated cells. J Struct Biol 139:76–89
Boyle S, Gilchrist S, Bridger JM, Mahy NL, Ellis JA, Bickmore WA (2001) The spatial organization of human chromosomes within the nuclei of normal and emerin-mutant cells. Hum Mol Genet 10:211–219
Brown KE, Baxter J, Graf D, Merkenschlager M, Fisher AG (1999) Dynamic repositioning of genes in the nucleus of lymphocytes preparing for cell division. Mol Cell 3:207–217
Brown KE, Amoils S, Horn JM, Buckle VJ, Higgs DR, Merkenschlager M, Fisher AG (2001) Expression of alpha- and beta-globin genes occurs within different nuclear domains in haemopoietic cells. Nat Cell Biol 3:602–606
Brown JM, Leach J, Reittie JE, Atzberger A, Lee-Prudhoe J, Wood WG, Higgs DR, Iborra FJ, Buckle VJ (2006) Coregulated human globin genes are frequently in spatial proximity when active. J Cell Biol 172:177–187
Brown JM, Green J, das Neves RP, Wallace HA, Smith AJ, Hughes J, Gray N, Taylor S, Wood WG, Higgs DR, Iborra FJ, Buckle VJ (2008) Association between active genes occurs at nuclear speckles and is modulated by chromatin environment. J Cell Biol 182:1083–1097
Bridger JM, Herrmann H, Muenkel C, Lichter P (1998) Identification of an interchromosomal compartment by polymerisation of nuclear-targeted vimentin. J Cell Sci 111:1241–1253
Chambeyron S, Bickmore WA (2004) Chromatin decondensation and nuclear reorganization of the HoxB locus upon induction of transcription. Genes Dev 18:1119–1130
Chambeyron S, Da Silva NR, Lawson KA, Bickmore WA (2005) Nuclear re-organisation of the Hoxb complex during mouse embryonic development. Development 132:2215–2223
Chen K, Baxter T, Muir WM, Groenen MA, Schook LB (2007) Genetic resources, genome mapping and evolutionary genomics of the pig (Sus scrofa). Int J Biol Sci 3:153–165
Chubb JR, Bickmore WA (2003) Considering nuclear compartmentalization in the light of nuclear dynamics. Cell 112:403–406
Cremer T, Cremer M, Dietzel S, Müller S, Solovei I, Fakan S (2006) Chromosome territories—a functional nuclear landscape. Curr Opin Cell Biol 18:307–316
Croft JA, Bridger JM, Boyle S, Perry P, Teague P, Bickmore WA (1999) Differences in the localization and morphology of chromosomes in the human nucleus. J Cell Biol 145:1119–1131
Ehlers JP, Worley L, Onken MD, Harbour JW (2005) DDEF1 is located in an amplified region of chromosome 8q and is overexpressed in uveal melanoma. Clin Cancer Res 11:3609–3613
Farmer SR (2006) Transcriptional control of adipocyte formation. Cell Metab 4:263–273
Federico C, Cantarella CD, Di Mare P, Tosi S, Saccone S (2008) The radial arrangement of the human chromosome 7 in the lymphocyte cell nucleus is associated with chromosomal band gene density. Chromosoma 117:399–410
Finlan LE, Sproul D, Thomson I, Boyle S, Kerr E, Perry P, Ylstra B, Chubb JR, Bickmore WA (2008) Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet 4:e1000039
Formiguera X, Cantón A (2004) Obesity: epidemiology and clinical aspects. Best Pract Res Clin Gastroenterol 18:1125–1146
Foster HA, Bridger JM (2005) The genome and the nucleus: a marriage made by evolution. Genome organisation and nuclear architecture. Chromosoma 114:212–229
Foster HA, Abeydeera LR, Griffin DK, Bridger JM (2005) Non-random chromosome positioning in mammalian sperm nuclei, with migration of the sex chromosomes during late spermatogenesis. J Cell Sci 118:1811–1820
Goetze S, Mateos-Langerak J, Gierman HJ, de Leeuw W, Giromus O, Indemans MH, Koster J, Ondrej V, Versteeg R, van Driel R (2007) The three-dimensional structure of human interphase chromosomes is related to the transcriptome map. Mol Cell Biol 27:4475–4487
Grasser F, Neusser M, Fiegler H, Thormeyer T, Cremer M, Carter NP, Cremer T, Müller S (2008) Replication-timing-correlated spatial chromatin arrangements in cancer and in primate interphase nuclei. J Cell Sci 121:1876–1886
Guo X, Liao K (2000) Analysis of gene expression profile during 3T3-L1 preadipocyte differentiation. Gene 251:45–53
Habermann FA, Cremer M, Walter J, Kreth G, von Hase J, Bauer K, Wienberg J, Cremer C, Cremer T, Solovei I (2001) Arrangements of macro- and microchromosomes in chicken cells. Chromosome Res 9:569–584
Hepperger C, Mannes A, Merz J, Peters J, Dietzel S (2008) Three-dimensional positioning of genes in mouse cell nuclei. Chromosoma . doi:10.1007/s00412-008-0168-2
Hofmann WA, Johnson T, Klapczynski M, Fan JL, de Lanerolle P (2006) From transcription to transport: emerging roles for nuclear myosin I. Biochem Cell Biol 84:418–426
Hu Q, Kwon YS, Nunez E, Cardamone MD, Hutt KR, Ohgi KA, Garcia-Bassets I, Rose DW, Glass CK, Rosenfeld MG, Fu XD (2008) Enhancing nuclear receptor-induced transcription requires nuclear motor and LSD1-dependent gene networking in interchromatin granules. Proc Natl Acad Sci U S A 105:19199–19204
Kosak ST, Skok JA, Medina KL, Riblet R, Le Beau MM, Fisher AG, Singh H (2002) Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296:158–162
Kouadjo KE, Nishida Y, Cadrin-Girard JF, Yoshioka M, St-Amand J (2007) Housekeeping and tissue-specific genes in mouse tissues. BMC Genomics 8:127
Küpper K, Kölbl A, Biener D, Dittrich S, von Hase J, Thormeyer T, Fiegler H, Carter NP, Speicher MR, Cremer T, Cremer M (2007) Radial chromatin positioning is shaped by local gene density, not by gene expression. Chromosoma 116:285–306
Kuroda M, Tanabe H, Yoshida K, Oikawa K, Saito A, Kiyuna T, Mizusawa H, Mukai K (2004) Alteration of chromosome positioning during adipocyte differentiation. J Cell Sci 117:5897–5903
Lanctôt C, Cheutin T, Cremer M, Cavalli G, Cremer T (2007) Dynamic genome architecture in the nuclear space: regulation of gene expression in three dimensions. Nat Rev Genet 8:104–115
Lunney JK (2007) Advances in swine biomedical model genomics. Int J Biol Sci 3:179–184
Meaburn KJ, Misteli T (2007) Cell biology: chromosome territories. Nature 445:379–781
Meaburn KJ, Misteli T (2008) Locus-specific and activity-independent gene repositioning during early tumorigenesis. J Cell Biol 180:39–50
Meaburn KJ, Cabuy E, Bonne G, Levy N, Morris GE, Novelli G, Kill IR, Bridger JM (2007) Primary laminopathy fibroblasts display altered genome organization and apoptosis. Aging Cell 6:139–153
Mehta IS, Elcock LS, Amira M, Kill IR, Bridger JM (2008) Nuclear motors and nuclear structures containing A-type lamins and emerin: is there a functional link? Biochem Soc Trans 36:1384–1388
Misteli T (2007) Beyond the sequence: cellular organization of genome function. Cell 128:787–800
Morey C, Da Silva NR, Perry P, Bickmore WA (2007) Nuclear reorganisation and chromatin decondensation are conserved, but distinct, mechanisms linked to Hox gene activation. Development 134:909–919
Morey C, Kress C, Bickmore WA (2009) Lack of bystander activation shows that localization exterior to chromosome territories is not sufficient to upregulate gene expression. Genome Res (in press)
Nakamura T, Shiojima S, Hirai Y, Iwama T, Tsuruzoe N, Hirasawa A, Katsuma S, Tsujimoto G (2003) Temporal gene expression changes during adipogenesis in human mesenchymal stem cells. Biochem Biophys Res Commun 303:306–312
Osborne CS, Chakalova L, Brown KE, Carter D, Horton A, Debrand E, Goyenechea B, Mitchell JA, Lopes S, Reik W, Fraser P (2004) Active genes dynamically colocalize to shared sites of ongoing transcription. Nat Genet 36:1065–1071
Osborne CS, Chakalova L, Mitchell JA, Horton A, Wood AL, Bolland DJ, Corcoran AE, Fraser P (2007) Myc dynamically and preferentially relocates to a transcription factory occupied by Igh. PLoS Biol 5:e192
Pérez-Mancera PA, Bermejo-Rodríguez C, González-Herrero I, Herranz M, Flores T, Jiménez R, Sánchez-García I (2007) Adipose tissue mass is modulated by SLUG (SNAI2). Hum Mol Genet 16:2972–2986
Ragoczy T, Bender MA, Telling A, Byron R, Groudine M (2006) The locus control region is required for association of the murine beta-globin locus with engaged transcription factories during erythroid maturation. Genes Dev 20:1447–1457
Reddy KL, Zullo JM, Bertolino E, Singh H (2008) Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452:243–247
Rosen ED, MacDougald OA (2006) Adipocyte differentiation from the inside out. Nat Rev Mol Cell Biol 7:885–896
Saccone S, Federico C, Bernardi G (2002) Localization of the gene-richest and the gene-poorest isochores in the interphase nuclei of mammals and birds. Gene 300:169–178
Sadoni N, Targosz BS, Englmann A, Fesser S, Koch J, Schindelhauer D, Zink D (2008) Transcription-dependent spatial arrangements of CFTR and conserved adjacent loci are not conserved in human and murine nuclei. Chromosoma 117:381–397
Schöfer C, Weipoltshammer K (2008) Gene dynamics and nuclear architecture during differentiation. Differentiation 76:41–56
Schook L, Beattie C, Beever J, Donovan S, Jamison R, Zuckermann F, Niemi S, Rothschild M, Rutherford M, Smith D (2005) Swine in biomedical research: creating the building blocks of animal models. Anim Biotechnol 16:183–190
Somech R, Shaklai S, Geller O, Amariglio N, Simon AJ, Rechavi G, Gal-Yam EN (2005) The nuclear-envelope protein and transcriptional repressor LAP2beta interacts with HDAC3 at the nuclear periphery, and induces histone H4 deacetylation. J Cell Sci 118:4017–4025
Szczerbal I, Chmurzynska A (2008) Chromosomal localization of nine porcine genes encoding transcription factors involved in adipogenesis. Cytogenet Genome Res 121:50–54
Szczerbal I, Chmurzynska A, Switonski M (2007a) Cytogenetic mapping of eight genes encoding fatty acid binding proteins (FABPs) in the pig genome. Cytogenet Genome Res 118:63–66
Szczerbal I, Lin L, Stachowiak M, Chmurzynska A, Mackowski M, Winter A, Flisikowski K, Fries R, Switonski M (2007b) Cytogenetic mapping of DGAT1, PPARA, ADIPOR1 and CREB genes in the pig. J Appl Genet 48:73–76
Takizawa T, Gudla PR, Guo L, Lockett S, Misteli T (2008a) Allele-specific nuclear positioning of the monoallelically expressed astrocyte marker GFAP. Genes Dev 22:489–498
Takizawa T, Meaburn KJ, Misteli T (2008b) The meaning of gene positioning. Cell 135:9–13
Tanabe H, Habermann FA, Solovei I, Cremer M, Cremer T (2002a) Non-random radial arrangements of interphase chromosome territories: evolutionary considerations and functional implications. Mutat Res 504:37–45
Tanabe H, Müller S, Neusser M, von Hase J, Calcagno E, Cremer M, Solovei I, Cremer C, Cremer T (2002b) Evolutionary conservation of chromosome territory arrangements in cell nuclei from higher primates. Proc Natl Acad Sci U S A 99:4424–4429
Tong Q, Dalgin G, Xu H, Ting CN, Leiden JM, Hotamisligil GS (2000) Function of GATA transcription factors in preadipocyte–adipocyte transition. Science 290:134–138
Volpi EV, Chevret E, Jones T, Vatcheva R, Williamson J, Beck S, Campbell RD, Goldsworthy M, Powis SH, Ragoussis J, Trowsdale J, Sheer D (2000) Large-scale chromatin organization of the major histocompatibility complex and other regions of human chromosome 6 and its response to interferon in interphase nuclei. J Cell Sci 113:1565–1576
Williams RR, Broad S, Sheer D, Ragoussis J (2002) Subchromosomal positioning of the epidermal differentiation complex (EDC) in keratinocyte and lymphoblast interphase nuclei. Exp Cell Res 272:163–175
Williams RR, Azuara V, Perry P, Sauer S, Dvorkina M, Jørgensen H, Roix J, McQueen P, Misteli T, Merkenschlager M, Fisher AG (2006) Neural induction promotes large-scale chromatin reorganisation of the Mash1 locus. J Cell Sci 119:132–140
Xu M, Cook PR (2008) The role of specialized transcription factories in chromosome pairing. Biochim Biophys Acta 1783:2155–2160
Zalenskaya IA, Zalensky AO (2004) Non-random positioning of chromosomes in human sperm nuclei. Chromosome Res 12:163–173
Zeng L, Rahrmann E, Hu Q, Lund T, Sandquist L, Felten M, O'Brien TD, Zhang J, Verfaillie C (2006) Multipotent adult progenitor cells from swine bone marrow. Stem Cells 24:2355–2366
Zink D, Amaral MD, Englmann A, Lang S, Clarke LA, Rudolph C, Alt F, Luther K, Braz C, Sadoni N, Rosenecker J, Schindelhauer D (2004) Transcription-dependent spatial arrangements of CFTR and adjacent genes in human cell nuclei. J Cell Biol 166:815–825
Zou L, Zou X, Chen L, Li H, Mygind T, Kassem M, Bünger C (2008) Multilineage differentiation of porcine bone marrow stromal cells associated with specific gene expression pattern. J Orthop Res 26:56–64
Acknowledgments
We thank Prof. Wendy Bickmore and Dr. Paul Perry (MRC HGU, Edinburgh) for erosion script, Prof. Malcom A. Ferguson-Smith (Cambridge Resource Centre for Comparative Genomics, Cambridge University, UK) for flow-sorted pig chromosome paints, and Prof. Fengtang Yang (Wellcome Trust Sanger Institute, Cambridge, UK) for paint probe for porcine chromosome 12. We also thank Dr. Maciej Szydlowski (Poznan University of Life Sciences, Poland) for helping with statistical analysis. This study was financed by the Polish Ministry of Science and Higher Education, grant N301 3381 33, fellowship 9/MOB/2007/0 and monies awarded to JMB from Brunel University.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by T. Misteli
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Table S1
(DOC 35 kb)
Table S2
(DOC 32 kb)
Table S3
(DOC 29 kb)
Table S4
(DOC 30 kb)
Fig. S1
Ideograms of porcine chromosomes (SSC4, SSC6, SSC12, SSC13, SSC15, SSC17) showing the location of selected genes. (JPG 145 kb)
Fig. S2
Relative positioning of four gene loci—PPARG, SREBF1, CEBPA, and FABP4 with nucleoli. Nucleoli were revealed with anti-fibrillarin (green) and gene probes were delineated in red. a MSC; b AD14. c The graph shows the percentage of total gene signals co-localized with nucleoli (means ± SEM). Scale bar, 10 μm (JPG 234 kb)
Rights and permissions
About this article
Cite this article
Szczerbal, I., Foster, H.A. & Bridger, J.M. The spatial repositioning of adipogenesis genes is correlated with their expression status in a porcine mesenchymal stem cell adipogenesis model system. Chromosoma 118, 647–663 (2009). https://doi.org/10.1007/s00412-009-0225-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-009-0225-5