Abstract
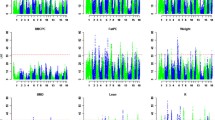
Serum glucose and lipid levels are associated with diabetes mellitus and cardiovascular disease. The purpose of this study was to identify quantitative trait loci (QTL) for serum glucose and lipids in a White Duroc × Erhualian resource population. Serum glucose, glycosylated serum proteins (GSP), and serum lipid levels were measured in a total of 760 F2 animals at 240 days. Strong positive correlations were observed between total cholesterol (TC) and low-density-lipoprotein cholesterol (LDL-C)/high-density-lipoprotein cholesterol (HDL-C). A whole-genome scan was performed with 194 microsatellites covering the pig genome across the entire resource population, revealing 2 QTL for serum glucose and 15 QTL for serum lipids. Of them, three 1% genome-wide significant QTL were identified for LDL-C, TC, and triglycerides (TG) in an adjacent region (67–73 cM) on chromosome 2 (SSC2), and the QTL for LDL-C showed the largest effect with a 95% confidence interval of 5 cM. Another 1% genome-wide significant QTL was found for LDL-C at 87 cM on SSC8. Other QTL showed 5% genome-wide significant or suggestive effects on SSC4, 5, 7, 9, 11, 14, and 15. In total, five significant QTL for serum lipids and a suggestive QTL for GSP on SSC4 were identified for the first time in pigs. Most of the identified QTL are homologous to the previously reported QTL for serum lipids in humans and mice. As correlated traits, QTL for TC and LDL-C were always located in the same genomic regions. The results shed new light on studies of human atherosclerosis and cardiovascular-related diseases.

Similar content being viewed by others
References
Aberg K, Sun G, Smelser D, Indugula SR, Tsai HJ et al (2008) Applying novel genome-wide linkage strategies to search for loci influencing type 2 diabetes and adult height in American Samoa. Hum Biol 80:99–123
Anunciado RV, Nishimura M, Mori M, Ishikawa A, Tanaka S et al (2003) Quantitative trait locus analysis of serum insulin, triglyceride, total cholesterol and phospholipid levels in the (SM/J × A/J) F2 mice. Exp Anim 52:37–42
Cefalu WT, Bell-Farrow AD, Petty M, Izlar C, Smith JA (1991) Clinical validation of a second-generation fructosamine assay. Clin Chem 37:1252–1256
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
de Koning DJ, Janss LL, Rattink AP, van Oers PA, de Vries BJ et al (1999) Detection of quantitative trait loci for backfat thickness and intramuscular fat content in pigs (Sus scrofa). Genetics 152:1679–1690
Feitosa MF, Rice T, Borecki IB, Rankinen T, Leon AS et al (2006) Pleiotropic QTL on chromosome 12q23–q24 influences triglyceride and high-density lipoprotein cholesterol levels: the HERITAGE family study. Hum Biol 3:317–327
Fox CS, Cupples LA, Chazaro I, Polak JF, Wolf PA et al (2004) Genomewide linkage analysis for internal carotid artery intimal medial thickness: evidence for linkage to chromosome 12. Am J Hum Genet 74:253–261
Franceschini G (2001) Epidemiologic evidence for high-density lipoprotein cholesterol as a risk factor for coronary artery disease. Am J Cardiol 88:9N–13N
Gallardo D, Pena RN, Amills M, Varona L, Ramirez O et al (2008) Mapping of quantitative trait loci for cholesterol, LDL, HDL and triglyceride serum concentrations in pigs. Physiol Genomics 35:199–209
Green P, Falls K, Crooks S (1990) Documentation for CRIMAP, version 2.4. Washington University School of Medicine, St. Louis, MO
Guo YM, Mao HR, Ren J, Yan XM, Duan YY et al (2009) A comprehensive linkage map of the porcine genome from a large scale White Duroc × Erhualian resource population and evaluation of factors affecting recombination rates. Anim Genet 40:47–52
Hasler-Rapacz J, Ellegren H, Fridolfsson AK, Kirkpatrick B, Kirk S et al (1998) Identification of a mutation in the low density lipoprotein receptor gene associated with recessive familial hypercholesterolemia in swine. Am J Med Genet 76:379–386
Hauser ER, Crossman DC, Granger CB, Haines JL, Jones CJ et al (2004) A genomewide scan for early-onset coronary artery disease in 438 families: the GENECARD Study. Am J Hum Genet 75:436–447
Heijmans BT, Beekman M, Putter H, Lakenberg N, van der Wijk HJ et al (2005) Meta-analysis of four new genome scans for lipid parameters and analysis of positional candidates in positive linkage regions. Eur J Hum Genet 13:1143–1153
Herrera VL, Didishvili T, Lopez LV, Myers RH, Ruiz-Opazo N (2004) Genome-wide scan identifies novel QTLs for cholesterol and LDL levels in F2[Dahl R×S]-intercross rats. Circ Res 94:446–452
Imperatore G, Knowler WC, Pettitt DJ, Kobes S, Fuller JH et al (2000) A locus influencing total serum cholesterol on chromosome 19p: results from an autosomal genomic scan of serum lipid concentrations in Pima Indians. Arterioscler Thromb Vasc Biol 20:2651–2656
JAMA (2001) Executive summary of the third report of the national cholesterol education program (NCEP) expert panel on detection, evaluation, and treatment of high blood cholesterol in adults (Adult Treatment Panel III). JAMA 285:2486–2497
Kathiresan S, Manning AK, Demissie S, D’Agostino RB, Surti A et al (2007) A genome-wide association study for serum lipid phenotypes in the Framingham Heart Study. BMC Med Genet 8:S17
Libby P (2001) Managing the risk of atherosclerosis: the role of high-density lipoprotein. Am J Cardiol 88:3N–8N
Libby P (2002) Inflammation in atherosclerosis. Nature 420:868–874
Mao HR, Guo YM, Yang GC, Yang B, Ren J et al (2008) A genome-wide scan for quantitative trait loci affecting limb bone lengths and areal bone mineral density of the distal femur in a large White Duroc × Erhualian F2 population. BMC Genet 9:63
Ng MC, So WY, Lam VK, Cockram CS, Bell GI et al (2004) Genome-wide scan for metabolic syndrome and related quantitative traits in Hong Kong Chinese and confirmation of a susceptibility locus on chromosome 1q21–q25. Diabetes 53:2676–2683
North KE, Goring HH, Cole SA, Diego VP, Almasy L et al (2006) Linkage analysis of LDL cholesterol in American Indian populations: the Strong Heart Family Study. J Lipid Res 47:59–66
Pollin TI, Hsueh WC, Steinle NI, Snitker S, Shuldiner AR et al (2004) A genome-wide scan of serum lipid levels in the Old Order Amish. Atherosclerosis 173:89–96
Seaton G, Haley CS, Knott SA, Kearsey M, Visscher PM (2002) QTL Express: mapping quantitative trait loci in simple and complex pedigrees. Bioinformatics 18:339–340
Stylianou IM, Langley SR, Walsh K, Chen Y, Revenu C et al (2008) Differences in DBA/1J and DBA/2J reveal lipid QTL genes. J Lipid Res 49:2402–2413
Uusitupa MI, Niskanen LK, Siitonen O, Voutilainen E, Pyorala K (1993) Ten-year cardiovascular mortality in relation to risk factors and abnormalities in lipoprotein composition in type 2 (non-insulin-dependent) diabetic and non-diabetic subjects. Diabetologia 36:1175–1184
Visscher PM, Thompson R, Haley CS (1996) Confidence intervals in QTL mapping by bootstrapping. Genetics 143:1013–1020
Wang XS, Paigen B (2005) Genetics of variation in HDL cholesterol in humans and mice. Circ Res 96:27–42
Wang XS, Ishimori N, Korstanje R, Rollins J, Paigen B (2005) Identifying novel genes for atherosclerosis through mouse-human comparative genetics. Am J Hum Genet 77:1–16
Yang GC, Ren J, Li SJ, Mao HR, Guo YM et al (2008) Genome-wide identification of QTL for age at puberty in gilts using a large intercross F2 population between White Duroc × Erhualian. Genet Sel Evol 40:529–539
Acknowledgments
We are very grateful to Xiaofeng Zheng (First Affiliated Hospital of Nanchang University) for his help in measuring phenotypes. This study was supported by the Natural Science Foundation of China (30425045) and the National 973 Program of China (2006CB708213).
Author information
Authors and Affiliations
Corresponding author
Additional information
R. Chen, J. Ren, and W. Li contributed equally to this work.
An erratum to this article can be found at http://dx.doi.org/10.1007/s00335-010-9256-8
Rights and permissions
About this article
Cite this article
Chen, R., Ren, J., Li, W. et al. A genome-wide scan for quantitative trait loci affecting serum glucose and lipids in a White Duroc × Erhualian intercross F2 population. Mamm Genome 20, 386–392 (2009). https://doi.org/10.1007/s00335-009-9190-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-009-9190-9