Abstract
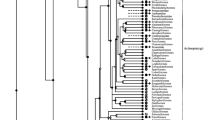
Indoleamine 2,3-dioxygenase (IDO) and tryptophan 2,3-dioxygenase (TDO) are tryptophan-degrading enzymes that catalyze the same reaction, the first step in tryptophan catabolism via the kynurenine pathway. TDO is widely distributed among life-forms, being found not only in eukaryotes but also in bacteria. In contrast, IDO has been found only in mammals and yeast to date. However, recent genome and EST projects have identified IDO homologues in non-mammals and found an IDO paralogue that is expressed in mice. In this study, we cloned the frog and fish IDO homologues and the mouse IDO paralogue, and characterized their enzymatic properties using recombinants. The IDOs of lower vertebrates and the mouse IDO paralogue had IDO activity but had 500–1000 times higher K m values and very low enzyme efficiency compared with mammalian IDOs. It appears that L-Trp is not a true substrate for these enzymes in vivo, although their actual function is unknown. On the phylogenetic tree, these low-activity IDOs, which we have named “proto-IDOs,” formed a cluster that was distinct from the mammalian IDO cluster. The IDO and proto-IDO genes are present tandemly on the chromosomes of mammals, including the marsupial opossum, whereas only the proto-IDO gene is observed in chicken and fish genomes. These results suggest that (mammalian) IDOs arose from proto-IDOs by gene duplication that occurred before the divergence of marsupial and eutherian (placental) mammals in mammalian evolutionary history.





Similar content being viewed by others
References
Ball HJ, Sanchez-Perez A, Weiser S, Austin CJD, Astelbauer F, Miu J, McQuillan JA, Stocker R, Jermiin LS, Hunt NH (2007) Characterization of an indoleamine 2,3-dioxygenase-like protein found in humans and mice. Gene 396:203–213
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
Fabrick JA, Kanost MR, Baker JE (2004) RNAi-induced silencing of embryonic tryptophan oxygenase in the Pyralid moth, Plodia interpunctella. J Insect Sci 4:15
Forouhar F, Anderson JL, Mowat CG, Vorobiev SM, Hussain A, Abashidze M, Bruckmann C, Thackray SJ, Seetharaman J, Tucker T, Xiao R, Ma LC, Zhao L, Acton TB, Montelione GT, Chapman SK, Tong L (2007) Molecular insights into substrate recognition and catalysis by tryptophan 2,3-dioxygenase. Proc Natl Acad Sci USA 104:473–478
Grohmann U, Fallarino F, Puccetti P (2003) Tolerance, DCs and tryptophan: much ado about IDO. Trends Immunol 24:242–248
Hinds LA, Poole WE, Tyndale-Biscoe CH, van Oorschot RAH, Cooper DW (1990) Reproductive biology and the potential for genetic studies in the tammar wallaby, Macropus eugenii. Aust J Zool 37:223–234
Hu X, Bao Z, Hu J, Shao M, Zhang L, Bi K, Zhan A, Huang X (2006) Cloning and characterization of tryptophan 2,3-dioxygenase gene of Zhikong scallop Chlamys farreri (Jones and Preston 1904). Aquat Res 37:1187–1194
Ishimura Y, Nozaki M, Hayaishi O (1970) The oxygenated form of L-tryptophan 2,3-dioxygenase as reaction intermediate. J Biol Chem 245:3593–3602
Iwamoto Y, Lee IS, Tsubaki M, Kido R (1995) Tryptophan 2,3-dioxygenase in Saccharomyces cerevisiae. Can J Microbiol 41:19–26
Kotake Y, Masayama I (1936) The intermediary metabolism of tryptophan. XVIII. The mechanism of formation of kynurenine from tryptophan. Z Physiol Chem 243:237–244
Lake JA (1991) The order of sequence alignment can bias the selection of tree topology. Mol Biol Evol 8:378–385
Littlejohn TK, Takikawa O, Skylas D, Jamie JF, Walker MJ, Truscott RJ (2000) Expression and purification of recombinant human indoleamine 2,3-dioxygenase. Protein Expr Purif 19:22–29
Littlejohn TK, Takikawa O, Truscott RJ, Walker MJ (2003) Asp274 and His346 are essential for heme binding and catalytic function of human indoleamine 2,3-dioxygenase. J Biol Chem 278:29525–29531
Lorenzen MD, Brown SJ, Denell RE, Beeman RW (2002) Cloning and characterization of the Tribolium castaneum eye-color genes encoding tryptophan oxygenase and kynurenine 3-monooxygenase. Genetics 160:225–234
Matsumura M, Osada K, Aiba S (1984) L-tryptophan 2,3-dioxygenase of a moderate thermophile, Bacillus brevis. Purification, properties and a substrate-mediated stabilization of the quaternary structure. Biochim Biophys Acta 786:9–17
Mukabayire O, Cornel AJ, Dotson EM, Collins FH, Besansky NJ (1996) The Tryptophan oxygenase gene of Anopheles gambiae. Insect Biochem Mol Biol 26:525–528
Murray MF (2007) The human indoleamine (2,3)-dioxygenase gene and related human genes. Curr Drug Metab 8:91–107
Panozzo C, Nawara M, Suski C, Kucharczyka R, Skoneczny M, Becam AM, Rytka J, Herbert CJ (2002) Aerobic and anaerobic NAD+ metabolism in Saccharomyces cerevisiae. FEBS Lett 517:97–102
Papadopoulou ND, Mewies M, McLean KJ, Seward HE, Svistunenko DA, Munro AW, Raven EL (2005) Redox and spectroscopic properties of human indoleamine 2,3-dioxygenase and a His303Ala variant: implications for catalysis. Biochemistry 44:14318–14328
Reed RD, Nagy LM (2005) Evolutionary redeployment of a biosynthetic module: expression of eye pigment genes vermilion, cinnabar, and white in butterfly wing development. Evol Dev 7:301–311
Schutz G, Feigelson P (1972) Purification and properties of rat liver tryptophan oxygenase. J Biol Chem 247:5327–5332
Searles LL, Voelker RA (1986) Molecular characterization of the Drosophila vermilion locus and its suppressible alleles. Proc Natl Acad Sci USA 83:404–408
Shimizu T, Nomiyama S, Hirata F, Hayaishi O (1978) Indoleamine 2,3-dioxygenase. Purification and some properties. J Biol Chem 253:4700–4706
Sono M, Roach MP, Coulter ED, Dawson JH (1996) Heme-containing oxygenases. Chem Rev 96:2841–2887
Suzuki T, Imai K (1998) Evolution of myoglobin. Cell Mol Life Sci 54:979–1004
Suzuki T, Takagi T (1992) A myoglobin evolved from indoleamine 2,3-dioxygenase. J Mol Biol 228:698–700
Suzuki T, Kawamichi H, Imai K (1998) A myoglobin evolved from indoleamine 2,3-dioxygenase, a tryptophan-degrading enzyme. Comp Biochem Physiol B 121:117–128
Takikawa O (2005) Biochemical and medical aspects of the indoleamine 2,3-dioxygenase-initiated L-tryptophan metabolism. Biochem Biophys Res Commun 338:12–29
Vottero E, Mitchell DA, Page MJ, MacGillivray RT, Sadowski IJ, Roberge M, Mauk AG (2006) Cytochrome b 5 is a major reductant in vivo of human indoleamine 2,3-dioxygenase expressed in yeast. FEBS Lett 580:2265–2268
Yamamoto S, Hayaishi O (1967) Tryptophan pyrrolase of rabbit intestine. D- and L-tryptophan-cleaving enzyme or enzymes. J Biol Chem 242:2560–2566
Yuasa HJ, Suzuki T (2005) Do molluscs possess indoleamine 2,3-dioxygenase? Comp Biochem Physiol B 140:445–454
Yuasa HJ, Hasegawa T, Nakamura T, Suzuki T (2007) Bacterial expression and characterization of molluscan IDO-like myoglobin. Comp Biochem Physiol B 146:461–469
Zhang Y, Kang SA, Mukherjee T, Bale S, Crane BR, Begley TP, Ealick SE (2007) Crystal structure and mechanism of tryptophan 2,3-dioxygenase, a heme enzyme involved in tryptophan catabolism and in quinolinate biosynthesis. Biochemistry 46:145–155
Acknowledgment
This work was partly supported by a Grant-in-Aid for Young Scientists B (to H.J.Y.) from the Japan Society for the Promotion of Science (KAKENHI 17770205).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yuasa, H.J., Takubo, M., Takahashi, A. et al. Evolution of Vertebrate Indoleamine 2,3-Dioxygenases. J Mol Evol 65, 705–714 (2007). https://doi.org/10.1007/s00239-007-9049-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-007-9049-1