Abstract
An anaerobic phthalate isomer-degrading strain (JTT) that we previously isolated was characterized. In addition, a strictly anaerobic, mesophilic, syntrophic phthalate isomer-degrading bacterium, designated strain JIT, was isolated and characterized in this study. Both were non-motile rods that formed spores. In both strains, the optimal growth was observed at temperatures around 37°C and neutral pH. In syntrophic co-culture with the hydrogenotrophic methanogen Methanospirillum hungatei, both strains could utilize two or three phthalate isomers for growth, and produce acetate and methane as end products. Strain JTT was able to grow on isophthalate, terephthalate, and a number of low-molecular weight aromatic compounds, such as benzoate, hydroquinone, 2-hydroxybenzoate, 3-hydroxybenzoate, 2,5-dihydroxybenzoate, 3-phenylpropionate in co-culture with M. hungatei. It could also grow on crotonate, hydroquinone and 2,5-dihydroxybenzoate in pure culture. Strain JIT utilized all of the three phthalate isomers as well as benzoate and 3-hydroxybenzoate for growth in co-culture with M. hungatei. No substrates were, however, found to support the axenic growth of strain JIT. Neither strain JTT nor strain JIT could utilize sulfate, sulfite, thiosulfate, nitrate, fumarate, Fe (III) or 4-hydroxybenzoate as electron acceptor. Phylogenetically, strains JTT and JIT were relatively close to the members of the genera Pelotomaculum and Cryptanaerobacter in ‘Desulfotomaculum lineage I’. Physiological and chemotaxonomic characteristics indicated that the two isolates should be classified into the genus Pelotomaculum, creating two novel species for them. Here, we propose Pelotomaculum terephthalicum sp. nov. and Pelotomaculum isophthalicum sp. nov. for strain JTT and strain JIT, respectively. The type strains are strains JTT (= DSM 16121T = JCM 11824T = NBRC 100523T) and JIT (= JCM 12282T = BAA-1053T) for P. terephthalicum and P. isophthalicum, respectively.




Similar content being viewed by others
References
Brauman A, Müller JA, Garcia JL, Brune A, Schink B (1998) Fermentative degradation of 3-hydroxybenzoate in pure culture by a novel strictly anaerobic bacterium, Sporotomaculum hydroxybenzoicum gen. nov., sp. nov. Int J Syst Bacteriol 48:215–221
de Bok FAM, Harmsen HJM, Plugge CM, de Vries MC, Akkermans ADL, de Vos WM, Stams AJM (2005) The first true obligately syntrophic propionate-oxidizing bacterium, Pelotomaculum schinkii sp. nov., co-cultured with Methanospirillum hungatei, and emended description of the genus Pelotomaculum. Int J Syst Evol Microbiol 55:1697–1703
Felsenstein J (1985) Confidence limits of phylogenies: an approach using the bootstrap. Evolution 39:783–791
Hanada S, Takaichi S, Matsuura K, Nakamura K (2002) Roseiflexus castenholzii gen. nov., sp. nov., a thermophilic, filamentous, photosynthetic bacterium that lacks chlorosomes. Int J Syst Evol Microbiol 52:187–193
Hattori S, Kamagata Y, Hanada S, Shoun H (2000) Thermacetogenium phaeum gen. nov., sp. nov., a strictly anaerobic, thermophilic, syntrophic acetate-oxidizing bacterium. Int J Syst Evol Microbiol 50:1601–1609
Hiraishi A (1992) Direct automated sequencing of 16S rDNA amplified by polymerase chain reaction from bacterial cultures without DNA purification. Lett Appl Microbiol 15:210–213
Hippe H, Hagenauer A, Kroppenstedt M (1997) Menadione requirement for sulfate-reduction in Desulfotomaculum aeronauticum, and emended species description. Syst Appl Microbiol 20:554–558
Imachi H, Sekiguchi Y, Kamagata Y, Ohashi A, Harada H (2000) Cultivation and in situ detection of a thermophilic bacterium capable of oxidizing propionate in syntrophic association with hydrogenotrophic methanogens in a thermophilic methanogenic granular sludge. Appl Environ Microbiol 66:3608–3615
Imachi H, Sekiguchi Y, Kamagata Y, Hanada S, Ohashi A, Harada H (2002a) Pelotomaculum thermopropionicum gen. nov., sp. nov., an anaerobic, thermophilic, syntrophic propionate-oxidizing bacterium. Int J Syst Evol Microbiol 52:1729–1735
Imachi H, Qiu YL, Sekiguchi Y, Kamagata Y, Ohashi A, Harada H (2002b) A novel anaerobic, syntrophic propionate-oxidizing bacterium affiliated with a clone cluster Ih in the genus Desulfotomaculum: taxonomical study of an isolate and polyphasic study on the clone subcluster Ih. Presented at American Society for Microbiology 102nd General Meeting, Salt Lake City, Utha, USA, pp 260–260
Jackson BE, Bhupathiraju VK, Tanner RS, Woese CR, McInerney MJ (1999) Syntrophus aciditrophicus sp. nov., a new anaerobic bacterium that degrades fatty acids and benzoate in syntrophic association with hydrogen-using microorganisms. Arch Microbiol 171:107–114
Juteau P, Côté V, Duckett MF, Beaudet R, Lépine F, Villemur R, Bisaillon JG (2005) Cryptanaerobacter phenolicus gen. nov., sp. nov., an anaerobe that transforms phenol into benzoate via 4-hydroxybenzoate. Int J Syst Evol Microbiol 55:245–250
Kamagata Y, Mikami E (1991) Isolation and characterization of a novel thermophilic Methanosaeta strain. Int J Syst Bacteriol 41:191–196
Kleerebezem R, Hulshoff Pol LW, Lettinga G (1999) Anaerobic degradation of phthalate isomers by methanogenic consortia. Appl Environ Microbiol 65:1152–1160
Kleerebezem R, Lettinga G (2000) High-rate anaerobic treatment of purified terephthalic acid wastewater. Water Sci Technol 42:259–268
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar, Buchner A, Lai T, Steppi S, Jobb G, Förster W, Brettske I, Gerber S, Ginhart AW, Gross O, Grumann S, Hermann S, Jost R, König A, Liss T, Lüßmann R, May M, Nonhoff B, Reichel B, Strehlow R, Stamatakis A, Stuckmann N, Vilbig A, Lenke M, Ludwig T, Bode A, Schleifer KH (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32:1363–1371
Mountfort DO, Bryant MP (1982) Isolation and characterization of an anaerobic syntrophic benzoate-degrading bacterium from sewage sludge. Arch Microbiol 133:249–256
Qiu YL, Sekiguchi Y, Imachi H, Kamagata Y, Tseng IC, Cheng SS, Ohashi A, Harada H (2003) Sporotomaculum syntrophicum sp. nov., a novel anaerobic, syntrophic benzoate-degrading bacterium isolated from methanogenic sludge treating wastewater from terephthalate manufacturing. Arch Microbiol 179:242–249
Qiu YL, Sekiguchi Y, Imachi H, Kamagata Y, Tseng IC, Cheng SS, Ohashi A, Harada H (2004) Identification and isolation of anaerobic, syntrophic phthalate isomer-degrading microbes from methanogenic sludges treating wastewater from terephthalate manufacturing. Appl Environ Microbiol 70:1617–1626
Ribbons DW, Taylor BF, Eaton RW, Anderson BN (1984) Microbial degradation of phthalates. In: Gibson DT (eds) Microbial degradation of organic compounds. Marcel Dekker, New York, NY pp 371–397
Saito N, Nei M (1987) The neighbor-joining method: a new method for constructing phylogenetic trees. Mol Biol Evol 4:406–425
Sekiguchi Y, Kamagata Y, Syutsubo K, Ohashi A, Harada H, Nakamura K (1998) Phylogenetic diversity of mesophilic and thermophilic granular sludges determined by 16S rRNA gene analysis. Microbiology 144:2655–2665
Sekiguchi Y, Kamagata Y, Nakamura K, Ohashi A, Harada H (2000) Syntrophothermus lipocalidus gen. nov., sp. nov., a novel thermophilic, syntrophic, fatty-acid-oxidizing anaerobe which utilizes isobutyrate. Int J Syst Evol Microbiol 50:771–779
Shintani T, Liu WT, Hanada S, Kamagata Y, Miyaoka S, Suzuki T, Nakamura K (2000) Micropruina glycogenica gen. nov., sp. nov., a new Gram-positive glycogen-accumulating bacterium isolated from activated sludge. Int J Syst Evol Microbiol 50:201–207
Stackebrandt E, Sproer C, Rainey FA, Burghardt J, Päuker O, Hippe H (1997) Phylogenetic analysis of the genus Desulfotomaculum: evidence for the misclassification of Desulfotomaculum guttoideum and description of Desulfotomaculum orientis as Desulfosporosinus orientis gen. nov., comb. nov. Int J Syst Bacteriol 47:1134–1139
Swofford DL (2002) PAUP*. Phylogenetic analysis using parsimony (*and Other Methods). Version 4. Sinauer Associates, Sunderland, MA
Szewzyk U, Schink B (1989) Degradation of hydroquinone, gentisate and benzoate by a fermenting bacterium in pure or defined mixed culture. Arch Microbiol 151:541–545
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Zhang H, Hanada S, Shigematsu T, Shibuya K, Kamagata Y, Kanagawa T, Kurane R (2000) Burkholderia kururiensis sp. nov., a trichloroethylene (TCE)-degrading bacterium isolated from an aquifer polluted with TCE. Int J Syst Evol Microbiol 50:743–749
Acknowledgements
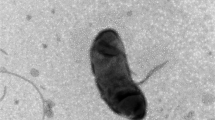
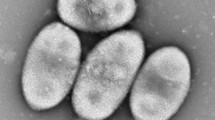
We thank the following researchers in the National Institute of Advanced Industrial Science and Technology (AIST): Xian-Ying Meng for TEM, and Mizuho Muramatsu and Akiko Ohashi for the determinations of quinones and the DNA base composition. This study was financially supported by a research grant on the support of young researchers with a term, subsidized by Ministry of Education, Culture, Sports, Science and Technology, Japan. This study was also supported by grants of the Japan Society for the Promotion of Science (JSPS) Postdoctoral Fellowship for Foreign Researchers.
Author information
Authors and Affiliations
Corresponding author
Additional information
Nucleotide sequence accession number: The GenBank/EMBL/DDBJ accession numbers of the 16S rRNA gene sequences of strains JTT and JIT are AB091323 and AB232785, respectively
Rights and permissions
About this article
Cite this article
Qiu, YL., Sekiguchi, Y., Hanada, S. et al. Pelotomaculum terephthalicum sp. nov. and Pelotomaculum isophthalicum sp. nov.: two anaerobic bacteria that degrade phthalate isomers in syntrophic association with hydrogenotrophic methanogens. Arch Microbiol 185, 172–182 (2006). https://doi.org/10.1007/s00203-005-0081-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-005-0081-5